Abstract
Introduction
Similarly to many fungal groups, the present taxonomic classification of yeasts still bears the remnants of dual nomenclature and generic concepts that traditionally used a combination of physiological, biochemical and morphological characteristics. Before dual nomenclature was abandoned in 2011 (Hawksworth 2011), yeast genera were divided into two groups representing either sexual and asexual morphs that were referred to as teleomorphic and anamorphic genera, respectively. These two groups of genera were recognized because of differences in reproduction and were treated separately in the last two editions of the compendium The Yeasts, a Taxonomic Study (TYTS, Kurtzman & Fell 1998; Kurtzman et al. 2011). Beyond this nomenclatural classification, it became clear that asexual and sexual species sometimes belong to the same phylogenetic clades in molecular phylogenetic analyses. With the development of molecular taxonomy and its use of DNA-based methods, the classification of yeasts experienced several events of splitting and lumping of genera (reviewed in Boekhout et al. 2021). Phylogenetic analyses using single-gene and later multigene approaches demonstrated that some traditionally delimited ascomycetous yeast genera, i.e., Saccharomyces, Kluyveromyces, Pichia, and Candida, were polyphyletic (Kurtzman & Robnett 1998, 2003). During the last two decades, Kurtzman and colleagues revised most of the teleomorphic ascomycetous yeasts and proposed various newly described genera based on multigene phylogenetic analyses (i.e., Kurtzman & Robnett 2003, 2007, 2010; Kurtzman et al. 2007, 2008; Kurtzman & Suzuki 2010). During this reclassification, the composition of several well-known genera changed following changes in generic concepts and methods for species delimitation. For example, most of the revised teleomorphic genera by Kurtzman (2003) in the Saccharomycetaceae can be recognized by phenotypic, phylogenetic and genomic features (Liu et al. 2024a).
The genus Candida is the largest anamorphic genus of yeasts and includes 314 species in the latest edition of the TYTS (Lachance et al. 2011). The genus Candida was introduced a century ago by C.M. Berkhout (1923) to include asexually reproducing yeasts that form hyphae that may disarticulate and form blastoconidia (or yeast cells) by budding from either hyphae or from other blastoconidia. Candida vulgaris, currently a taxonomic synonym of C. tropicalis (Lachance et al. 2011), was selected by Berkhout as the generic type. Many molecular phylogenetic studies demonstrated that the genus Candida is polyphyletic and consists of 11 to 15 clades spread over large parts of the subphylum Saccharomycotina (Kurtzman & Robnett 1998, 2003; Lachance et al. 2011; Daniel et al. 2014; Takashima & Sugita 2022). Daniel et al. (2014) indicated that the taxonomy of the genus Candida, as well as other yeast genera including asexually reproducing species, needs to be revised to make generic demarcations and membership consistent with their phylogenetic affinities. Following the ‘One fungus = One name’ principle and using results from many molecular phylogenetic sequence analyses, especially those based on multigene-based datasets, approximately half of the Candida species have already been placed into more than 36 existing genera and 14 newly proposed genera (e.g., Kurtzman 2016; Kurtzman et al. 2016; de Vega et al. 2017; Santos et al. 2018; Takashima & Sugita 2022; Liu et al. 2024b). Despite these efforts, the polyphyletic nature of the genus Candida has not been fully resolved, as the phylogenetic positions of many species remain unclear. This is particularly true for under-sampled lineages that occupy basal positions or those that are loosely placed in clades that received weak support (Daniel et al. 2014; Takashima & Sugita 2022). Although the recognition of new monotypic genera is criticized by some members of the yeast taxonomy community (e.g., Lachance 2018), monotypic genera (and higher ranks) are not rare in fungi with yeast states, e.g., see the genera Aciculiconidium, Babjeviella, Cyrenella and Kriegeria (Kurtzman 2011b; Sampaio 2011; Sampaio & Oberwinkler 2011; Smith 2011). Furthermore, Takashima & Sugita (2022) argued that the proposal of monotypic genera might be useful because it reduces the polyphyletic nature of the genera in which the species concerned have been classified before. Whether monotypic lineages are the result of conserved evolution or insufficient sampling is another valid point for discussion. In several instances, new species belonging to these initially monotypic genera have been described shortly after the genus was firstly established. For example, this was true for several originally monotypic genera described within the last decade, Babjevia, Deakozyma, Nematodospora and Yueomyces (Gouliamova et al. 2016; Ren et al. 2016; Zheng et al. 2017; Yamazaki et al. 2020; Yu et al. 2023). While it may still be difficult to decide on whether to introduce new monotypic yeast genera, genome-based statistics can help to objectivate decisions on generic demarcations in the absence of other criteria (Liu et al. 2024a, b). Particularly, the three genomic metrics, average amino acid identity (AAI), percentage of conserved proteins (POCP) and presence-absence patterns of orthologs (PAPO), have been tested for their utility in Saccharomycetaceae and Metschnikowiaceae using genera that were traditionally recognized based on their morphology (including sexual morphs) and physiological traits (Liu et al. 2024a, b). These studies showed that genera delimited using the approach combining sexual reproduction, physiology and multi-gene phylogenies in these two families exhibited a range of 80–92% POCP values and a range of 60–70% AAI values.
In the present study, we generated a robust genome-scale phylogeny, together with the LSU rDNA gene and combined ITS+LSU-based datasets in order to include all Candida species for which no genome data were available. We explored the heterogeneity of currently recognized yeast genera using the same genome metrics, i.e., AAI, POCP and PAPO, as used previously in the studies of Liu et al. (2024a, b), adopted some RED values calculated by Li et al. (2021), and evaluated phenotypic data to resolve the classification of the highly heterogeneous genus Candida. Here, we propose an updated taxonomy of Candida and related genera in the Saccharomycotina, including 175 new combinations, 87 new species, mainly validations of previously invalidly published names including 4 generic names, and 25 new genera.
Materials and methods
Genome assembly and annotation
The nuclear DNA of some yeast strains, i.e., Candida sp. XZY480-2, Candida sp. gmt3-3-4, Candida sp. XZY238F3 was extracted in this study using the method previously described by Wang & Bai (2008) (Table S1). Genomic libraries with 150 bp paired-end reads were constructed using the TruSeq Nano DNA library prep kit (Illumina) according to the manufacturer's instructions and sequenced on an Illumina HiSeq 2000 platform with the TruSeq SBS Kit (Illumina). Low-quality and adapter sequences were removed using Fastp v0.20.1 with default settings (Chen et al. 2018). The genome assembly was performed using SPAdes v3.15.0 (Bankevich et al. 2012) with parameters set to "--memory 800 -k 21,33,55,77,99 --careful --cov-cutoff auto". Gene prediction was carried out using GeneMark-ES (Ter-Hovhannisyan et al. 2008).
Genome quality assessment
We evaluated the quality of genome assemblies using the Benchmarking Universal Single-Copy Orthologs (BUSCO) v5.3.2 (Manni et al. 2021) and the Fungi odb10 database v4. The proportion of single-copy BUSCO genes in each genome indicates its completeness. To reduce missing data and exclude potentially low-quality genomes, we only included genomes with > 40% complete BUSCO genes.
Phylogenomic analysis and comparative genomics
To determine the evolutionary relationships of species classified in the genus Candida and related genera in Saccharomycotina, we analyzed single-copy orthologs from 906 genomes as listed in Table S1. Four genome datasets, namely, the Dipodascomycetes_dataset, the Phaffomycetales_dataset, the Pichiales_dataset and the Serinales_dataset, were used to construct phylogenomic trees. To preserve as much genomic information as possible, we selected 526 single-copy genes for the Dipodascomycetes_dataset, 652 for the Phaffomycetales_dataset, 545 for the Pichiales_dataset, and 583 for the Serinales_dataset, respectively, each of which has more than 95% of each genome dataset occupancy. For strains with a high duplicated BUSCOs rate, we first used the EMBOSS water alignment software (Madeira et al. 2019; Li et al. 2020) to perform intra-copy comparisons for each multicopy gene to select one copy from genes with an identity greater than 95%. This step was used to reduce the bias of our random selection. These orthologues were then aligned using MAFFT v7.475 with the G-INS-i option (Katoh & Standley 2013), concatenated using Perl scripts available at (https://github.com/Liufei0823/Single_Copy_Orthologue/), and a maximum likelihood (ML) gene tree was generated with IQ-TREE 2 (Minh et al. 2020), employing the MFP model and performing 1000 ultrafast bootstrap repeats (-m MFP -B 1000 -redo -mredo -nt AUTO). The alignments were deposited in TreeBASE (www.treebase.org, No. 31986).
Calculation of AAI value
We evaluated the amino acid identity (AAI) of species of Candida and related genera or clades utilizing the CompareM (available at https://github.com/dparks1134/CompareM) v0.1.2 software and applied the default settings as used before (Liu et al. 2024a, b).
Calculation of Presence-absence patterns of orthologs (PAPO)
Orthologous Groups (OGs) were determined by clustering all proteins with OrthoFinder v2.5.4 (Emms & Kelly 2019). The method by Takashima et al. (2019) was used to create presence-absence patterns of orthologs (PAPO). Based on the OrthoFinder results, OGs that were absent were labeled as 0, while those that were present were labeled as 1. Core proteins were defined as OGs that existed in a clade, pan proteins were OGs found in at least one strain of a clade, and unique proteins were OGs found in all strains of a clade but not in any other clades (Liu et al. 2024a, b).
Calculation of the percentage of conserved proteins (POCP) value
Following the method outlined by Qin et al. (2014), we determined the percentage of conserved proteins (POCP) between two strains. We utilized BLASTp (Tatusova & Madden 1999) to compare the proteins of both strains and identified conserved proteins based on criteria such as identity greater than 40%, aligned length of at least 50%, and an e-value of less than 1 × 10−5. The POCP was then calculated as the ratio of conserved proteins in the two proteomes (Liu et al. 2024a, b).
The impact of horizontal transfer genes (HTGs) and reductive evolution on the AAI and POCP analyses
To evaluate the impact of HTGs on the AAI and POCP analyses, the suspected HTGs in the genus Starmerella were identified using a pipeline adapted from Nowell et al. (2018). The protein sequences were aligned against the UniRef90 database using DIAMOND (Buchfink et al. 2021) with an e-value threshold of 1e-⁵, retaining the top 500 hits per query. Alignment results were processed using the diamond_to_HGT_candidates.pl script, which employs a Lowest Common Ancestor (LCA) algorithm. Genes meeting both of the following criteria were designated as horizontal gene transfer (HGT) candidates: 1) HGT Index (hU) ≥ 50: computed as the difference in bitscores between the best outgroup match (e.g., bacteria or plants) and the best ingroup match (fungi); 2) consensus Hit Support (CHS) ≥ 95%: defined as the percentage of all supporting hits consistent with the outgroup classification [(Number of outgroup-supporting hits/total hits) × 100%]. The HGT candidates were removed from the protein sequence files using SeqKit v0.10.0 (Shen et al. 2024). The AAI and POCP values were then calculated using the filtered protein files. Scatter plots were then constructed using the ggplot2 package (Wickham 2016) in R (R Core Team 2025) to visually illustrate the distribution trends of AAI (or POCP) values before and after the removal of HGT candidates. The horizontal axis represented AAI (or POCP) values between any pairwise combination of species calculated based on all proteins (with HTGs retained), while the vertical axis represented AAI (or POCP) values for the corresponding species pairs calculated after removal of HGT candidates. A linear regression model was fitted to the two datasets using the ggpmisc package (Aphalo 2025) in R, and the slopes of the linear regression equations, as well as the coefficients of determination (R²), were calculated. Wherein, the R² value ranges between 0 and 1, and is used to evaluate the goodness of fit of the linear regression model to the data: the closer R² is to 1, the better the model’s fitting effect (Draper & Smith 1998). If the slope of the linear regression equation was close to 1 and the R² value was high, it indicated high consistency in AAI (or POCP) values for any pair of species before and after removal of HTGs, implying that HTGs had no significant interference on AAI (or POCP) analysis. Conversely, a slope noticeably deviating from 1 or a low R² value suggested that HTGs might affect the results of AAI (or POCP) analysis.
The BUSCO analysis showed that Starmerella had a higher rate of missing BUSCOs, ranging from 13.5% to 34.1% (Table S1), which indicates that Starmerella undergoes reductive evolution, which is in agreement with the results from Gonçalves et al. (2020, 2022) and Pontes et al. (2024). To evaluate the impact of reductive evolution on the AAI and POCP analysis, the orthologous groups (OGs) occurring in ≥ 95% and ≥ 90% species of Starmerella were extracted using SeqKit v0.10.0, respectively, and assigned to two datasets, namely the Starmerella_95_dataset and Starmerella_90_dataset, which were used to calculate the AAI and POCP values. Two sets of data were then used to generate scatter plots using the ggplot2 package in R: 1) AAI values (serving as the horizontal axis) between any pairwise species calculated based on all proteins, and 2) AAI values (serving as the vertical axis) for the corresponding species pairs calculated based on Starmerella_90_dataset or Starmerella_95_dataset. Linear regression analyses were performed using the ggpmisc package in R to assess the similarity between these sets of data, determining whether reductive evolution affects AAI analysis. The same approach was applied to evaluate the impact of reductive evolution on POCP analysis using the ggpmisc and ggplot2 packages in R.
The impact of introgressions, hybridization and alloaneuploidy on the AAI and POCP analyses
To assess the impact of hybridization (or alloaneuploidy) on the AAI and POCP analyses, the Dipodascus/Galactomyces/Geotrichum lineage was used as an example in this study. Three distinct datasets with/without duplicated genes were constructed: 1) All_genome_dataset includes all proteins (genes) for each species of Dipodascus/Galactomyces/Geotrichum lineage; 2) Subgenome_1_dataset contains orthologous groups (OGs) without paralogous genes; 3) Subgenome_2_dataset is composed of OGs without paralogous genes and one copy of paralogous genes that was randomly selected from the paralog-containing OGs. All conserved proteins of species in the Dipodascus/Galactomyces/Geotrichum lineage were extracted with SeqKit v.0.10.0. The pairwise AAI and POCP values between species were then calculated based on these three datasets. Then, scatter plots were generated and linear regression analyses were performed to determine whether hybridization (or alloaneuploidy) affected the results of the AAI analysis. Wherein, the horizontal axis represented AAI values between any pairwise species calculated based on the All_genome_dataset, while the vertical axis represented AAI values for the corresponding species pairs calculated based on Subgenome_1_dataset or Subgenome_2_dataset. Following the same approach, the impact of hybridization (or alloaneuploidy) on POCP analysis was assessed.
Currently, phylogenetic evidence indicates that Saccharomyces cerevisiae DBVPG 6765 carries a large amount of introgressed material (D'Angiolo et al. 2020). Saccharomyces pastorianus has been identified as an interspecies hybrid between S. cerevisiae and Saccharomyces eubayanus (Monerawela & Bond 2018). Similarly, Saccharomyces bayanus has been identified as a hybrid derivative of Saccharomyces uvarum and S. eubayanus (Pérez-Través et al. 2014). Saccharomyces cerevisiae × Saccharomyces kudriavzevii strains have been isolated from beer fermentation environments (Peris et al. 2012). Therefore, we used Saccharomyces as an example to address the reliability of AAI and POCO analyses affected by introgressions and hybridization (or alloaneuploidy). Three genome datasets were generated: 1) Saccharomyces_9_dataset includes only nine natural species, namely Saccharomyces arboricola, S. cerevisiae, Saccharomyces chiloensis, S. eubayanus, Saccharomyces jurei, S. kudriavzevii, Saccharomyces mikatae, Saccharomyces paradoxus and S. uvarum; 2) Saccharomyces_10_dataset is composed of nine natural species and S. cerevisiae DBVPG 6765 with introgression; 3) Saccharomyces_12_dataset contains nine natural species and three hybrid species (or strains), namely S. bayanus, S. pastorianus and S. cerevisiae × S. kudriavzevii. The AAI and POCP analyses were conducted separately for each dataset.
Genome-wide analysis of multidrug resistance in the revised Candida species
To evaluate the antifungal resistance profiles of Candida species assigned to the newly created genera in this study, we retrieved known antifungal resistance-associated genes of C. albicans from NCBI, including: 1) the ERG11 gene (with mutation sites F126L, Y132F, K143R, F145L, G448E, F449V, G450E, and G464S) associated with azole resistance; 2) the ERG2 gene (with mutation site F105SfsX23) associated with polyene resistance; 3) the FKS1 gene (with mutation sites S645P/Y/F and F641Y) associated with echinocandin resistance; 4) the FUR1 gene (with mutation site F211I) associated with nucleoside analog resistance (Katiyar et al. 2006; Jensen et al. 2015; Wu & Ying 2016). The blastdb tool within the BLAST software was then used to construct a non-redundant (NR) database for these antifungal resistance-associated gene sequences. The software Blastp v2.11.0+ (Camacho et al. 2009) was used to perform sequence alignment between the protein sequence files of the studied strains and the previously constructed NR database, as well as the extraction of homologous antifungal resistance-associated gene sequences to obtain the corresponding gene sequences of Candida species in the newly created genera. Subsequently, we used Mafft v7.475 (Katoh & Standley 2013) to conduct sequence alignment on all the obtained homologous genes. The alignment results were analyzed to identify the similarities and differences in antifungal resistance-associated genes between resistant and susceptible strains.
Ribosomal DNA (rDNA) phylogenetic analysis
The DNA sequences of ITS (including 5.8S) and LSU D1/D2 domains of ribosomal DNA (rDNA) (Table S2) were acquired from NCBI and aligned using the MAFFT G-INS-i program (Katoh & Standley 2013). A Maximum Likelihood (ML) tree was constructed using RAxML v8.2.12 (Stamatakis 2014) with the GRT+I+G model. The reliability of the phylogenetic branches was determined through 1,000 bootstrap analyses (Felsenstein 1985).
Generic delineation
Takashima et al. (2019) proposed the PAPO analysis to delineate genera for basidiomycetous yeasts in the Trichosporonales. Recently, a range of 80–92% POCP values and a range of 60–70% AAI values were recommended as indicative values to delimitate genera in Saccharomycetaceae (Liu et al. 2024a), which were also used in the reclassification of Candida species in Metschnikowiaceae (Liu et al. 2024b). The RED approach has been applied to yeasts and fungal taxonomy (Li et al. 2021; Groenewald et al. 2023). The RED ± 0.1 intervals have been calculated for different taxonomic levels, i.e., 0.29 ± 0.1 for phylum, 0.695 ± 0.1 for class, 0.79 ± 0.1 for order, 0.889 ± 0.1 for family, 0.96 ± 0.1 for genus. Notably, all clades are assigned in this study based on the combined analyses of the AAI, POCP, PAPO approaches, and the RED index was used if RED values were available.
Results and discussion
Genome-scale and rDNA phylogenetic analyses
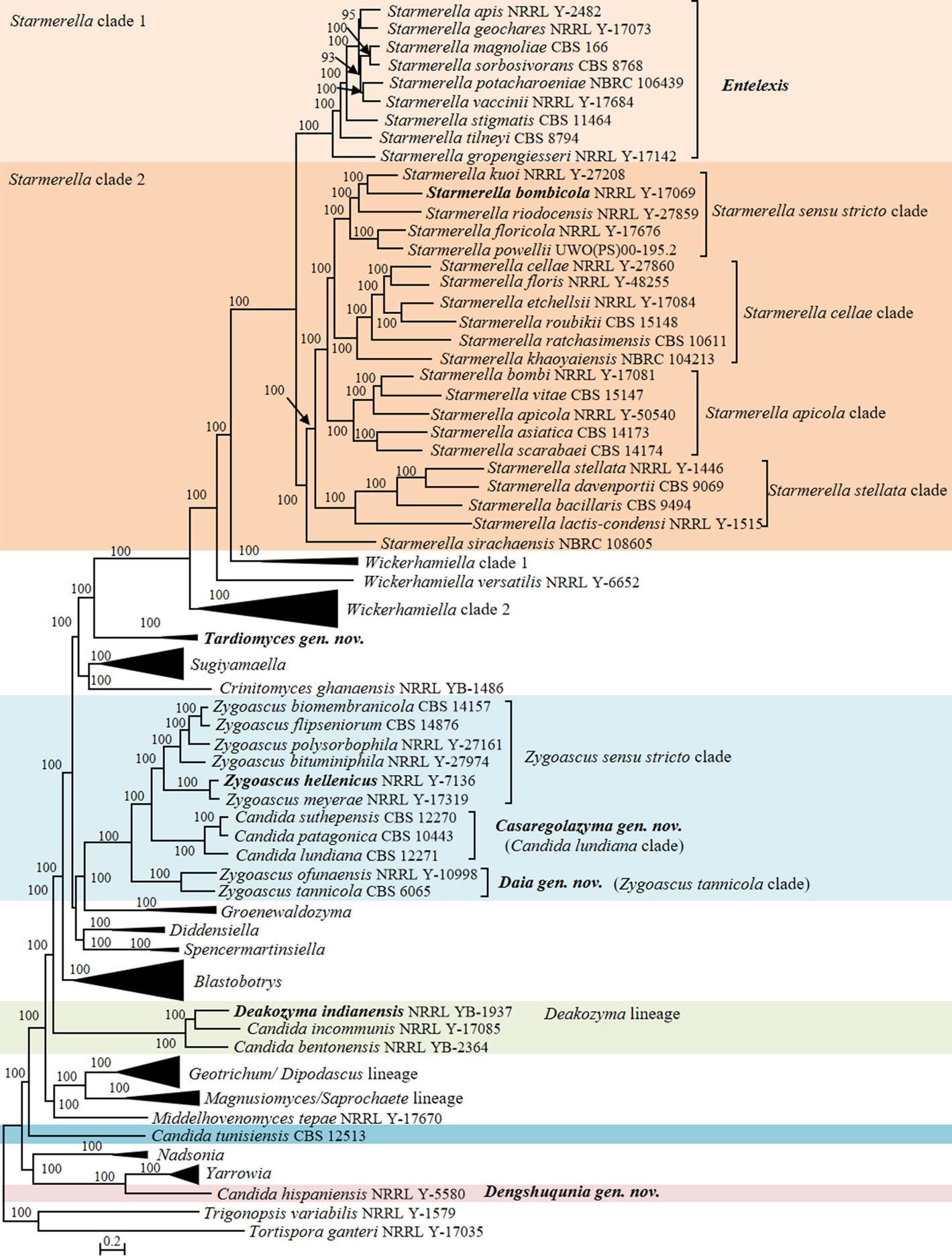
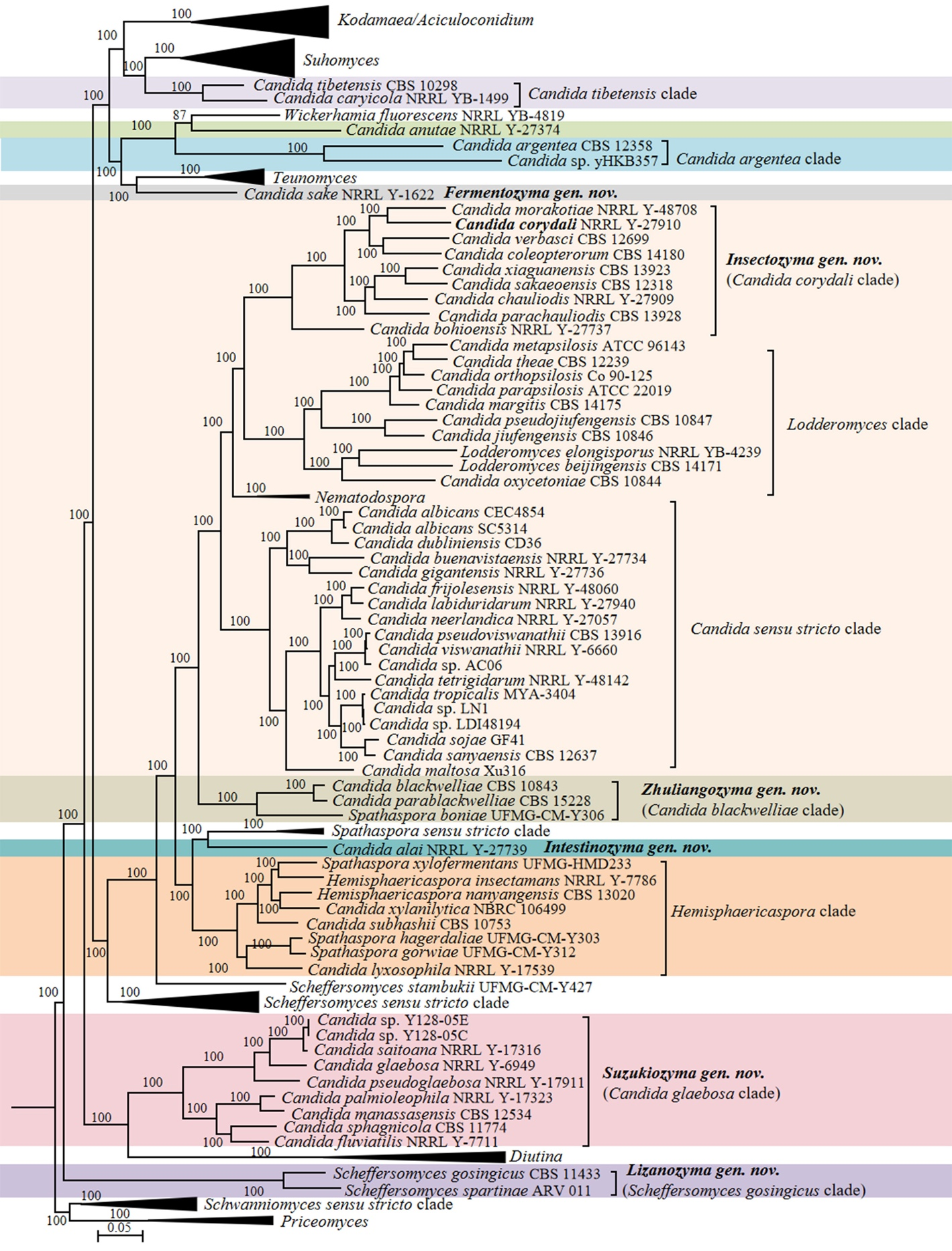
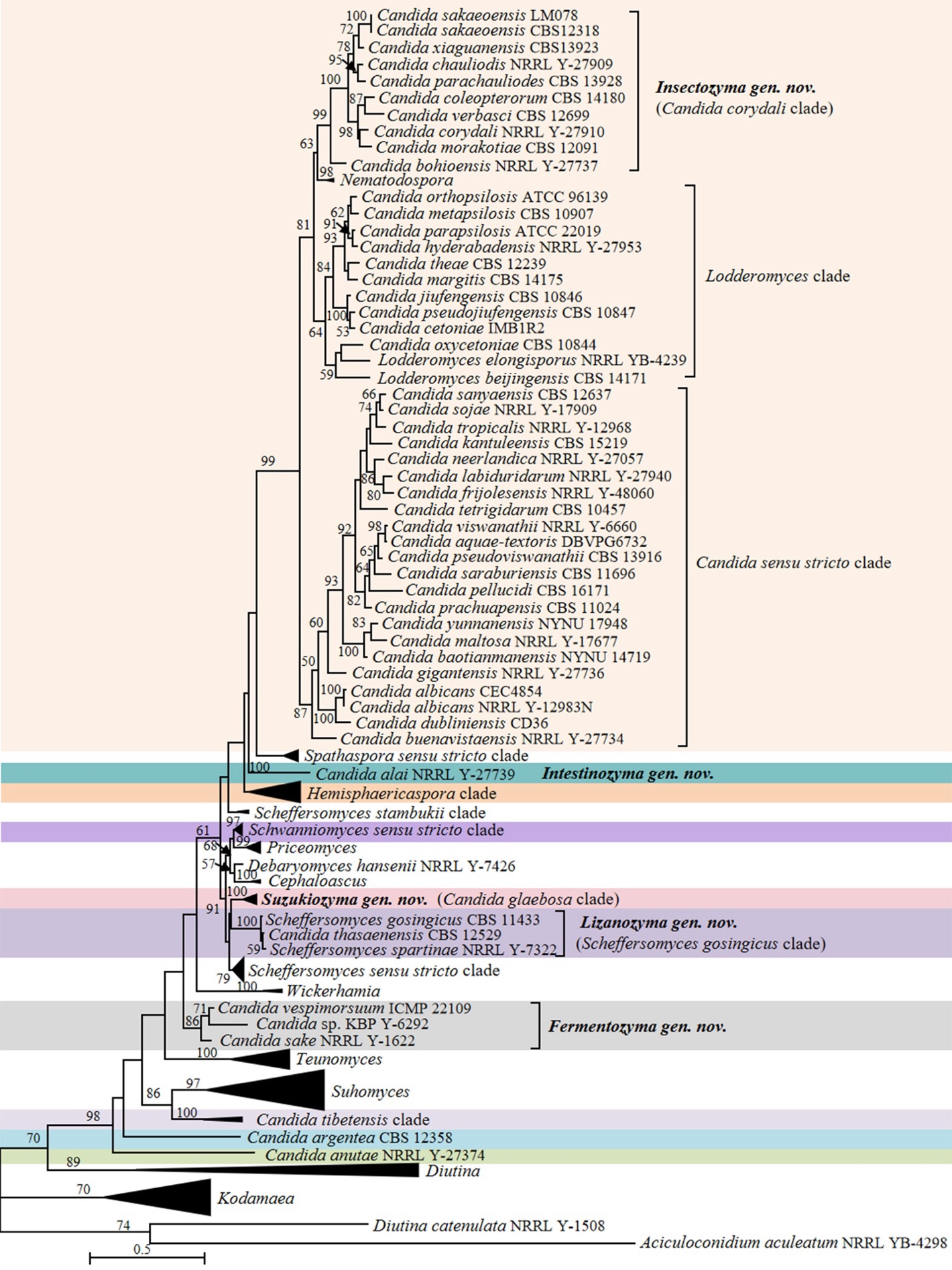
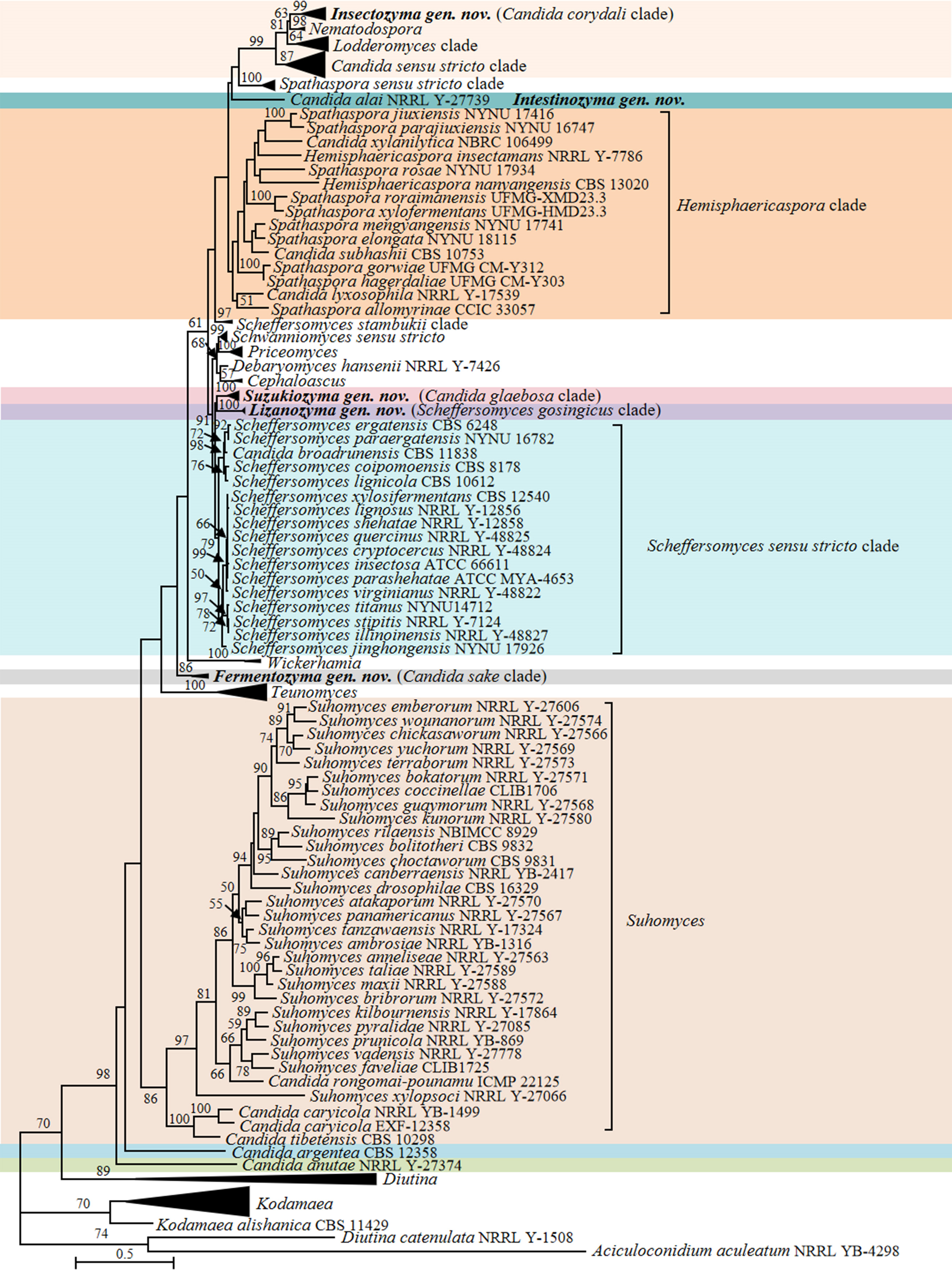
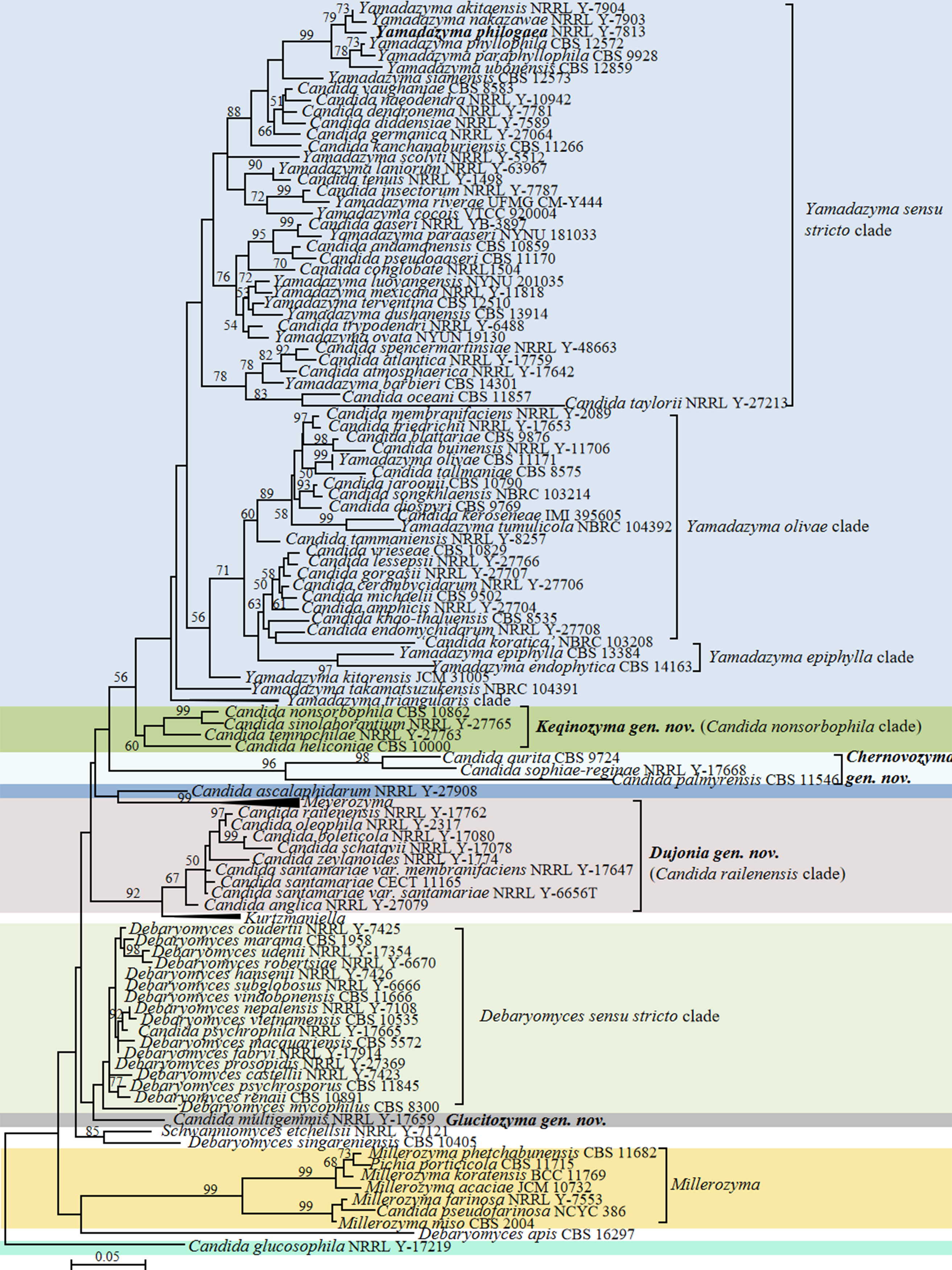
Candida vulgaris, the type species of the genus Candida, and currently interpreted as a synonym under C. tropicalis (Lachance et al. 2011), is phylogenetically placed in the class Pichiomycetes, order Serinales, family Debaryomycetaceae (Groenewald et al. 2023). Although the heterogenic and polyphyletic features of the genus Candida have been reduced by several molecular phylogenetic studies and recent taxonomic reclassifications (e.g., Kurtzman 2016; Kurtzman et al. 2016; de Vega et al. 2017; Santos et al. 2018; Takashima & Sugita 2022; Avesani et al. 2024, Liu et al. 2024b), this genus still includes 189 species scattered across Saccharomycotina (Table S2) with 10, 148 and 31 species belonging to Dipodascomycetes, Pichiomycetes and Saccharomycetes, respectively. To provide a well-supported placement of these species in relation to already described genera, 157 genomes of Candida species and 749 genomes of the related species in the Dipodascomycetes, Pichiomycetes and Saccharomycetes were used to generate phylogenomic trees (Figs. 1, 6, 9, 12, 14–15, 17 and Figs. S1–S4) and run three genome-based metrics analyses, i.e., AAI, POCP and PAPO analyses (Table 1). Although most Candida species have been included in the genome-scale analyses, for about 35 Candida species no genome sequences are available yet. To obtain a reliable placement of these latter yeast species, a phylogenetic analysis of nucleotide sequences representing LSU rDNA (D1/D2 domains) and ITS was carried out using all described Candida species published before December 2024, as well as some yet undescribed taxa (Figs. 2–5, 7–8, 10–11, 13, 16, 18 and Figs. S5–S13). In the next sections, we provide a comprehensive analysis of the phylogenomics data and genome statistics of 906 strains, and the phylogenetic trees based on rDNA data of 840 yeast species.
Candida species in the Dipodascomycetes
Ten Candida species were placed in the Dipodascomycetes (Fig. 1, Table S1 and S2). These species are phylogenetically distant from the core of the genus Candida, they belong to a different class, namely Dipodascomycetes rather than Pichiomycetes. Thus, there is no objective reason to classify these yeasts in the genus Candida. Our phylogenomic analysis showed that eight Candida species were located in the Trichomonascaceae (Dipodascales), where the other Candida species were placed as incertae sedis in the Dipodascomycetes (Fig. 1, Table S1 and S2).
Candida species and related genera Deakozyma and Limtongella (Trichomonascaceae, Dipodascales, Dipodascomycetes)
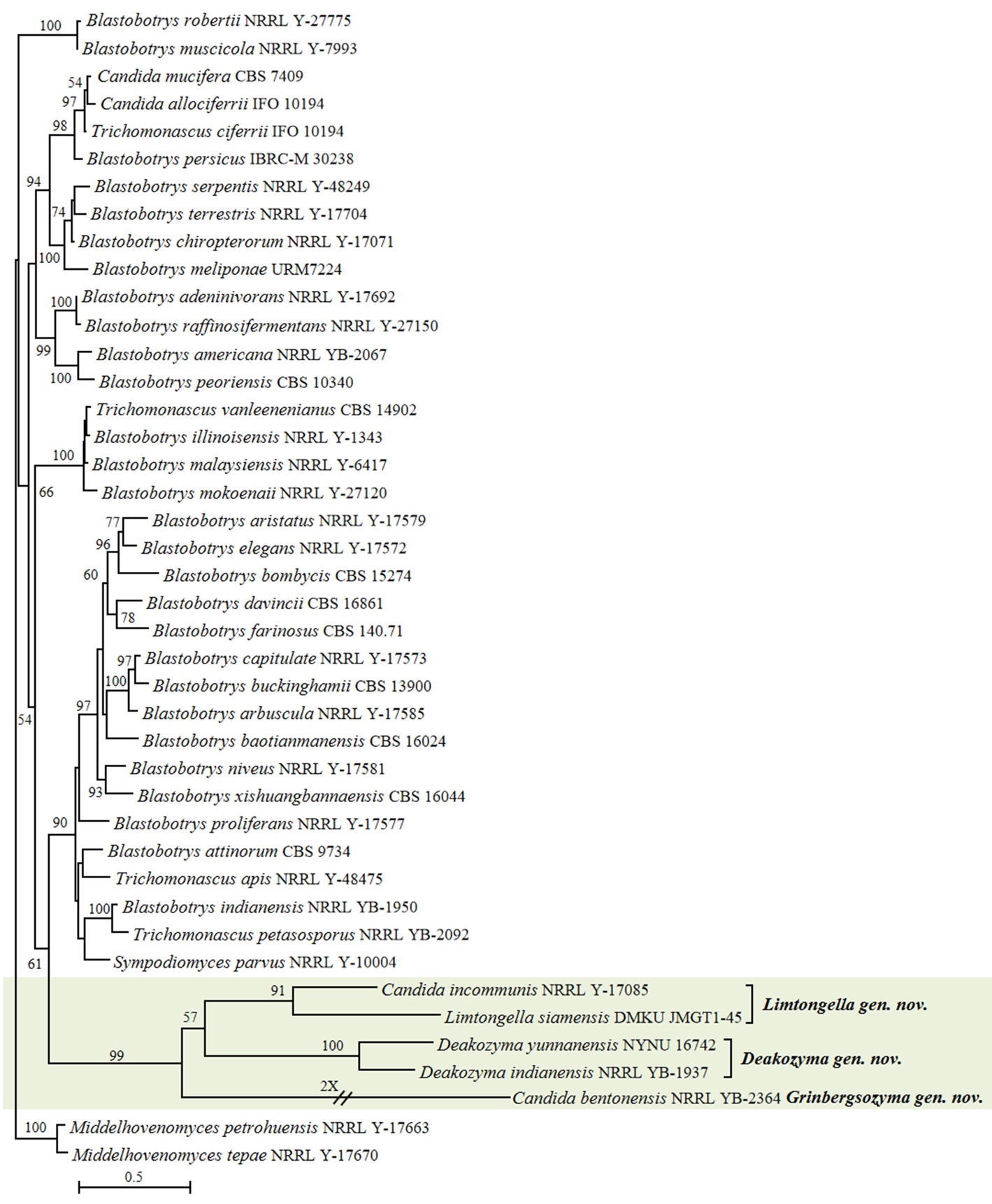
Our analyses showed that Candida bentonensis, Candida incommunis and Deakozyma indianensis were placed within the Deakozyma lineage (Fig. 1). Recently, the new genus Limtongella was proposed by Sakpuntoon et al. (2020) for Limtongella siamensis, which was found to be closely related to D. indianensis and C. incommunis. However, unfortunately, C. bentonensis was not included in the multigene phylogenetic analysis made by Sakpuntoon et al. (2020). Our phylogenetic analysis showed that D. indianensis and Deakozyma yunnanensis (Zheng et al. 2017) formed a well-supported clade in the ITS+D1/D2 tree (Fig. 2). The genus Limtongella and C. incommunis clustered together with 91% bootstrap support, while C. bentonensis formed a separate and long branch, clearly distinct from both Deakozyma and Limtongella (Fig. 2). These findings suggest that C. incommunis belongs to the genus Limtongella, whereas C. bentonensis represents another genus. To accommodate C. bentonensis, we propose the erection of a new genus, named Grinbergsozyma gen. nov. (Fig. 2, Table S2). The D1/D2 LSU rDNA phylogenetic analysis revealed that the newly identified C. bentonensis lineage (Grinbergsozyma gen. nov.) contains at least three potential new species isolated from various substrates, including the flux of Quercus rubra in Canada, the flux of Quercus oleoides in Costa Rica, as well as soil and rotting wood in Brazil (Fig. S5, Table S2), all of which differ from C. bentonensis by 2.16–15% in the D1/D2 domains. Additionally, the genus Limtongella includes three potential new species labelled in GenBank as Candida cf. incommunis UWO(PS)01-669.2 (GenBank AF530616) isolated from the flux of Hymenaea courbaril in Costa Rica, Candida sp. YWW5-1 (GenBank LC387304) isolated from mangrove forests in Thailand, and Candida sp. DMKU-FW29-11 (GenBank OL679539) obtained from food waste in Thailand (Fig. S5, Table S2).
Candida species and related genera Diddensiella and Sugiyamaella (Trichomonascaceae, Dipodascales, Dipodascomycetes)
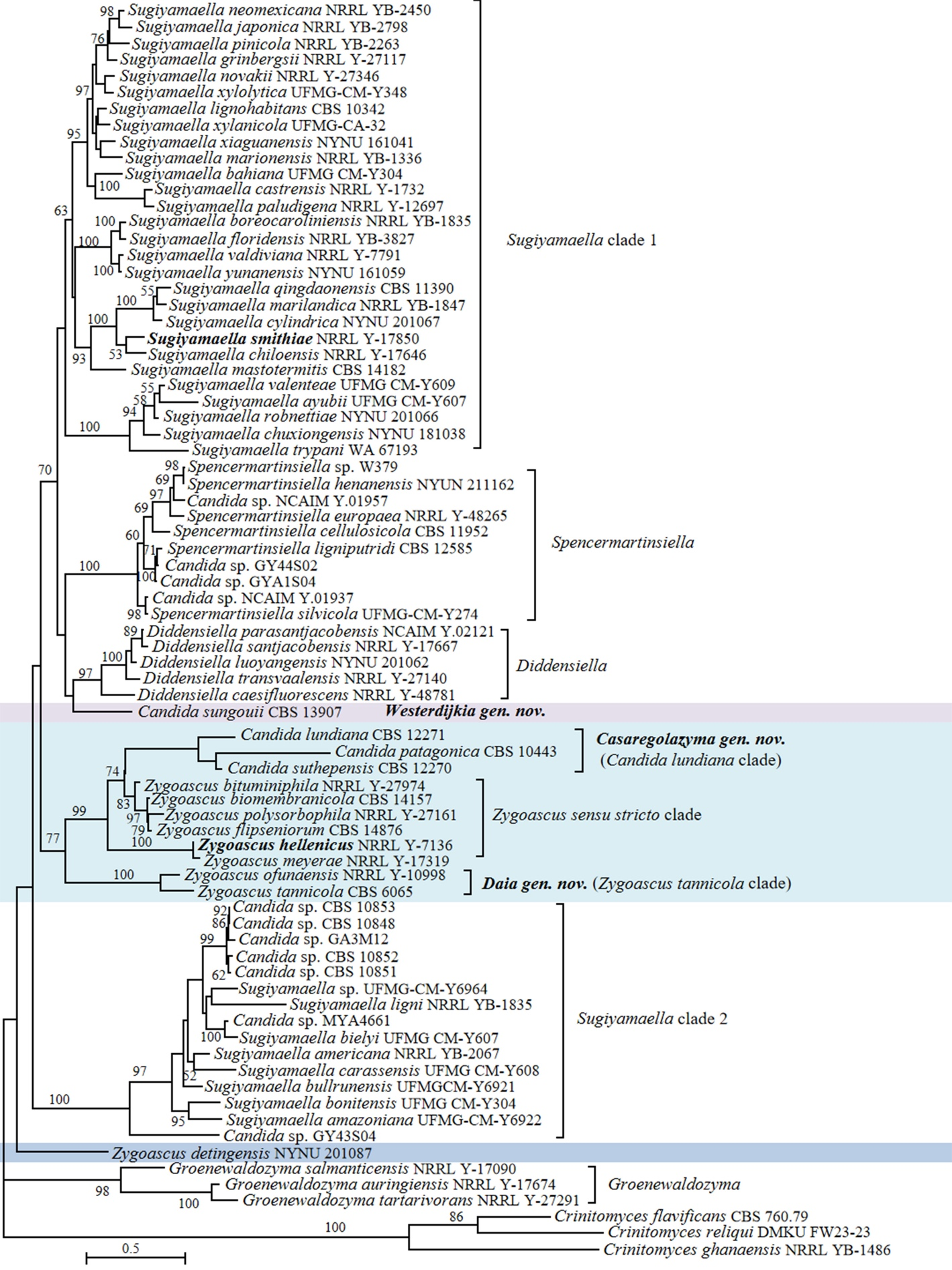
Sylvester et al. (2015) described Candida sungouii and concluded that this species did not belong to any known genus, but indicated its affinity to the genera Diddensiella and Sugiyamaella in Trichomonascaceae. The same authors also indicated close relationships of C. sungouii with two potential new species labelled as Candida sp. BG02-5-30-009A-1 (GenBank AY520421) and Candida sp. BG02-7-18-018A-2-2 (GenBank AY520408), both isolated from basidiocarp-feeding beetles (Suh et al. 2005). Our ITS+D1/D2 LSU phylogenetic analysis showed that C. sungouii clustered with Diddensiella and Spencermartinsiella with low bootstrap support (Fig. 3). Candida sp. BG02-7-18-018A-2-2 (GenBank AY520408) isolated from the gut of a tenebrionid beetle in the USA showed a 4.3% D1/D2 sequence difference from the sequence of C. sungouii, thus suggesting that strain BG02-7-18-018A-2-2 likely represents a new member, closely related to C. sungouii, of this putative new genus (Fig. S6). Hence, we propose a new genus, Westerdijkia gen. nov., to accommodate C. sungouii and Candida sp. BG02-7-18-018A-2-2 (Fig. 3, Fig. S6, Table S2).
Candida species and related genus Starmerella (Trichomonascaceae, Dipodascales, Dipodascomycetes)
Candida spenceri is phylogenetically related to the genus Starmerella (Daniel et al. 2014). Starmerella was introduced by Rosa & Lachance (1998) with a single species, namely Starmerella bombicola. Later, a second species, Starmerella meliponinorum, was described by Teixeira et al. (2003). Based on a rDNA phylogenetic analysis, Lachance et al. (2011) demonstrated that various Candida species were phylogenetically related to the Starmerella lineage and these formed two separate groups. Daniel et al. (2014) argued that there was insufficient support to justify splitting the genus Starmerella into two different genera at that time. Subsequently, Santos et al. (2018) transferred 25 related Candida species into Starmerella based on a D1/D2 LSU rDNA sequence analysis. More recently, Sipiczki & Baghela (2025) proposed a new species Starmerella aleppica in the sensu stricto subclade (referred to as Starmerella clade 2 in this study) and argued that the sensu lato subclade (referred to as Starmerella clade 1 in this study) and the Starmerella sensu stricto subclade may represent two genera. This hypothesis was based on the ITS+D1/D2 LSU rDNA sequence analysis, as well as taxon-specific markers or signatures (InDel markers) in the D1/D2 and ITS regions (Sipiczki & Baghela 2025).
In agreement with the above studies and a recent phylogenomic study (Opulente et al. 2024), our phylogenomic analysis revealed a well-supported bifurcation within Starmerella. Specifically, the Starmerella clade 1 (previously referred to as the Candida magnoliae subclade by Daniel et al.) diverges from the other species within the genus (Fig. 1).
The genus Starmerella appears to be heterogeneous based on our analyses using genomic metrics. Starmerella exhibited a lower than expected range of both, AAI values (55.13–88.10%) and POCP values (49.07–93.24%) (Table 1). The RED value estimated for Starmerella by Li et al. (2021) was 0.91, suggesting that this genus may represent a family-level taxon (family RED interval 0.889 ± 0.1). Considering the taxonomic heterogeneity suggested above, we calculated the genome metrics of Starmerella clade 1. According to our results (AAI: 71.55–88.10%; POCP: 80.33–93.54%; PAPO: 19), the values fell within the range observed for genera in Saccharomycetaceae (Liu et al. 2024a), suggesting that this clade would represent a taxonomic unit at the genus level with genomic heterogeneity within the expected range. However, genomic indices of the Starmerella clade 2 (AAI: 55.13–76.61%; POCP: 49.07–91.12%; PAPO: 0) indicated this clade remained too genetically heterogeneous when compared to the Starmerella clade 1. Our phylogenomic analysis identified four distinct clades within Starmerella clade 2 (Fig. 1). Genomic metrics were applied to assess the genetic heterogeneity within these smaller clades: Starmerella sensu stricto clade (AAI: 65.96–76.61%; POCP: 78.62–91.12%; PAPO: 3), the Starmerella apicola clade (AAI: 62.44–69.07%; POCP: 77.02–85.86%; PAPO: 4), the Starmerella cellae clade (61.78–70.71%; POCP: 60.03–85.26%; PAPO: 1), the Starmerella stellata clade (AAI: 58.08–68.13%; POCP: 60.00–78.43%; PAPO: 2), and the single-species lineage Starmerella sirachaensis (Fig. 1).
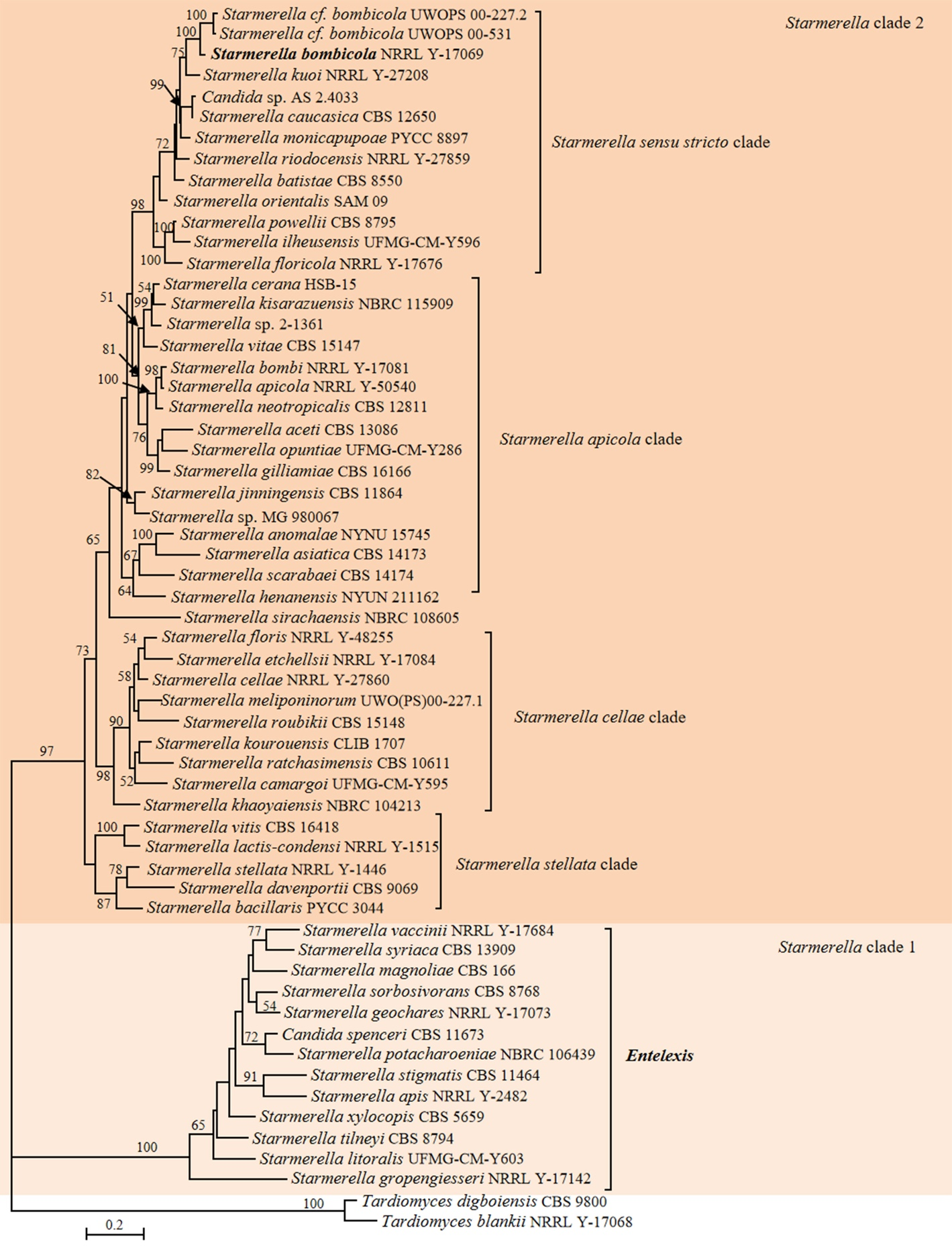
The genus Entelexis was proposed to accommodate the perfect state of Torulopsis magnoliae by van der Walt & Johannsen (1973). Yarrow & Meyer (1978) amended the genus Candida to include non-hyphal species and transferred Torulopsis species into Candida, consequently, T. magnoliae was assigned to Candida as a new combination. Later, Santos et al. (2018) transferred C. magnoliae (as known as T. magnoliae) to the genus Starmerella. The type of Entelexis magnoliae CBS 2798 (AY521568) differed from the type of C. magnoliae (also known as T. magnoliae and Starmerella magnoliae) CBS 166 (NG_060814) by 31 nucleotides in the D1/D2 domains of LSU rDNA, which indicated that E. magnoliae and C. magnoliae belong to different species. Our D1/D2 sequence analysis showed that both E. magnoliae and C. magnoliae were located in the Starmerella clade 1 (Fig. S7). Considering the results of phylogenomic analyses (Fig. 1 in this study, Opulente et al. 2024) and the arguments provided by Sipiczki & Baghela (2025), we propose the reclassification of Candida spenceri and the species within the Starmerella clade 1 (Fig. 4) into Entelexis. Moreover, a new name will be proposed in the Taxonomy section for C. magnoliae to avoid the synonymy of those two species. Although our genome-based metric analyses showed that Starmerella clade 2 is heterogeneous (Table 1), its reclassification will be addressed in the future after consultation with relevant yeast taxonomy experts.
| Taxa | AAI (%) | POCP (%) | PAPO |
| Dipodascomycetes | |||
| Crinitomyces | N/A | N/A | 446 |
| Diddensiella | 65.80–74.65 | 83.30–90.99 | 38 |
| Groenewaldozyma | 62.14–74.45 | 73.70–90.00 | 111 |
| Nadsonia | 67.44–91.67 | 81.87–93.64 | 197 |
| Spencermartinsiella | 74.34–74.34 | 89.58–89.58 | 109 |
| Starmerella | 55.13–88.10 | 49.07–93.24 | 2 |
| Starmerella clade 1 | 71.55–88.10 | 80.33–93.24 | 19 |
| Starmerella clade 2 | 55.13–76.61 | 49.07–91.12 | 0 |
| Starmerella sensu stricto clade | 65.96–76.61 | 78.62–91.12 | 3 |
| Starmerella apicola clade | 62.44–69.07 | 77.02–85.86 | 4 |
| Starmerella cellae clade | 61.78–70.71 | 60.03–85.26 | 1 |
| Starmerella stellata clade | 58.08–68.13 | 60.00–78.43 | 2 |
| Starmerella sensu stricto clade+Starmerella cellae clade | 60.16–76.61 | 60.03–91.12 | 0 |
| Starmerella apicola clade+Starmerella sensu stricto clade+Starmerella cellae clade | 58.62–76.61 | 60.03–91.12 | 1 |
| Starmerella sirachaensis single–species lineage | N/A | N/A | 122 |
| Sugiyamaella | 61.27–89.30 | 65.88–97.11 | 1 |
| Tardiomyces | 70.92–84.63 | 88.82–95.14 | 134 |
| Zygoascus lineage | 59.76–88.95 | 71.75–98.27 | 6 |
| Zygoascus sensu stricto clade | 67.17–88.95 | 87.31–98.27 | 21 |
| Candida lundiana clade | 77.81–89.01 | 88.85–95.34 | 74 |
| Zygoascus tannicola clade | 70.14–70.14 | 90.94–90.94 | 67 |
| Zygoascus sensu stricto clade+Candida lundiana clade | 61.98–89.01 | 74.47–98.27 | 9 |
| Candida tunisiensis single–species lineage | N/A | N/A | 382 |
| Yarrowia | 74.08–93.66 | 83.66–95.78 | 165 |
| Yarrowia+Candida hispaniensis | 60.80–93.66 | 68.29–95.78 | 196 |
| Candida hispaniensis single–species lineage | N/A | N/A | 262 |
| Pichiomycetes | |||
| Pichiales | |||
| Ambrosiozyma | 60.29–100.0 | 62.38-99.93 | 9 |
| Citeromyces | 71.28–85.77 | 87.41–91.71 | 134 |
| Brettanomyces | 61.89–96.77 | 72.69–94.27 | 5 |
| Candida insectalens clade | 62.99–62.99 | 82.72–82.72 | 9 |
| Komagataella | 85.54–97.90 | 94.43–98.89 | 141 |
| Kregervanrija | 88.94–89.70 | 95.45–96.48 | 79 |
| Ogataea | 56.34–99.54 | 55.36–97.74 | 0 |
| Ogataea clade 1 | 58.82–99.54 | 61.91–97.74 | 0 |
| Ogataea sensu stricto subclade | 64.30–99.54 | 76.83–97.74 | 2 |
| Ogataea pilisensis subclade | 61.27–90.18 | 75.32–97.46 | 0 |
| Ogataea saltuana subclade | 66.24–85.70 | 86.12–97.29 | 1 |
| Ogataea wickerhamii subclade | 65.49–86.36 | 87.03–94.58 | 5 |
| Ogataea saltuana subclade+Ogataea sensu stricto subclade | 61.77–99.54 | 71.15–97.74 | 0 |
| Ogataea saltuana subclade+Ogataea wickerhamii subclade+Ogataea sensu stricto subclade | 59.79–99.54 | 67.84–97.74 | 2 |
| Ogataea clade 2 | 58.46–90.74 | 68.35–97.84 | 6 |
| Ogataea naganishii subclade | 58.98–58.98 | 70.68–70.68 | 0 |
| Ogataea ramenticola subclade | 66.21–90.74 | 82.94–97.84 | 32 |
| Candida methanosorbosa subclade | 63.02–63.02 | 78.80–78.80 | 0 |
| Ogataea naganishii subclade+Candida methanosorbosa subclade | 58.98–63.02 | 70.68–78.80 | 0 |
| Ogataea ramenticola subclade+Ogataea naganishii subclade | 58.46–90.74 | 68.35–97.84 | 0 |
| Ogataea methylovora single–species lineage | N/A | N/A | 100 |
| Candida boidinii single–species lineage | N/A | N/A | 284 |
| Pichia | 61.31–94.88 | 62.33–97.35 | 0 |
| Saturnispora | 61.90–97.97 | 77.35–98.62 | 14 |
| Serinales | |||
| Cephaloascaceae | |||
| Cephaloascus | 66.77–66.77 | 84.06–84.06 | 110 |
| Cephaloascus+Candida chilensis | 60.62–66.77 | 74.92–84.06 | 61 |
| Debaryomycetaceae | |||
| Aciculoconidium | N/A | N/A | 380 |
| Debaryomyces | 58.87–91.79 | 73.93–97.16 | 0 |
| Candida/Lodderomyces lineage | 60.91–99.45 | 65.88–98.28 | 1 |
| Candida sensu stricto clade | 69.03–99.45 | 75.64–98.28 | 7 |
| Candida corydali clade | 67.27–81.01 | 81.71–93.33 | 3 |
| Lodderomyces clade | 64.25–83.42 | 73.73–96.02 | 4 |
| Lodderomyces clade+Candida corydali clade | 60.91–83.42 | 68.47–96.02 | 0 |
| Nematodospora | 76.99–76.99 | 94.83–94.83 | 5 |
| Candida aurita clade | 67.97–93.93 | 79.72–98.15 | 11 |
| Candida railenensis clade | 63.07–99.43 | 81.11–97.82 | 4 |
| Candida aurita clade+Candida railenensis clade | 61.51–99.43 | 74.04–98.15 | 2 |
| Candida aurita clade+Candida railenensis clade+Kurtzmaniella | 58.77–99.43 | 67.61–98.15 | 3 |
| Candida blackwelliae clade | 70.58–90.56 | 89.31–96.41 | 8 |
| Candida glaebosa clade | 65.65–86.35 | 82.55–96.03 | 23 |
| Candida nonsorbophila clade | 73.43–73.43 | 90.52–90.52 | 19 |
| Candida tibetensis clade | 74.16–74.16 | 94.09–94.09 | 10 |
| Candida alai single–species lineage | N/A | N/A | 132 |
| Candida anutae single–species lineage | N/A | N/A | 153 |
| Candida argentea single–species lineage | N/A | N/A | 180 |
| Candida ascalaphidarum single–species lineage | N/A | N/A | 68 |
| Candida glucosophila single–species lineage | N/A | N/A | 124 |
| Candida multigemmis single–species lineage | N/A | N/A | 95 |
| Candida sake single–species lineage | N/A | N/A | 84 |
| Diutina | 63.98–96.29 | 70.47–93.52 | 69 |
| Candida glaebosa clade+Diutina | 56.87–100.0 | 58.58–99.90 | 2 |
| Kodamaea | 60.35–91.83 | 60.41–95.76 | 0 |
| Kurtzmaniella | 64.66–85.58 | 84.41–96.46 | 16 |
| Meyerozyma | 67.31–94.33 | 90.39–98.21 | 18 |
| Millerozyma | 62.43–90.68 | 85.42–98.78 | 6 |
| Priceomyces | 63.86–84.38 | 85.48–96.33 | 4 |
| Scheffersomyces | 57.69–98.69 | 64.12–98.73 | 0 |
| Spathaspora | 62.35–90.50 | 70.35–93.21 | 0 |
| Spathaspora sensu stricto clade | 77.82–81.60 | 85.56–93.20 | 32 |
| Hemisphaericaspora clade | 71.91–90.50 | 77.12–93.21 | 11 |
| Spathaspora sensu stricto clade+Hemisphaericaspora clade | 65.17–90.50 | 71.32–93.21 | 2 |
| Spathaspora sensu stricto clade+Hemisphaericaspora clade+Candida alai | 64.79–90.50 | 71.32–93.21 | 1 |
| Spathaspora sensu stricto clade+Candida alai | 65.10–81.60 | 75.10–93.20 | 4 |
| Suhomyces | 66.35–96.44 | 86.82–98.34 | 0 |
| Suhomyces+Candida tibetensis clade | 63.89–96.44 | 84.33–98.34 | 1 |
| Schwanniomyces | 57.79–95.46 | 68.69–96.20 | 0 |
| Teunomyces | 74.25–96.37 | 92.42–98.43 | 12 |
| Wickerhamia fluorescens single–species lineage | N/A | N/A | 329 |
| Yamadazyma | 59.65–90.55 | 73.35–97.21 | 0 |
| Yamadazyma olivae clade | 62.06–97.75 | 77.72–98.83 | 0 |
| Yamadazyma epiphylla clade | 69.16–69.16 | 89.77–89.77 | 5 |
| Yamadazyma triangularis clade | 69.45–69.45 | 87.92–87.92 | 14 |
| incertae sedis in Serinales | |||
| Babjeviella | N/A | N/A | 231 |
| Limtongozyma | 81.76–81.76 | 93.60–93.60 | 299 |
| Candida chilensis single–species lineage | N/A | N/A | 196 |
| Saccharomycetes | |||
| Phaffomycetales | |||
| Barnettozyma | 64.52–86.96 | 81.56–95.37 | 0 |
| Barnettozyma sensu stricto clade | 70.01–86.96 | 88.56–95.37 | 15 |
| Barnettozyma wickerhamii clade | 69.91–95.36 | 88.46–96.27 | 0 |
| Barnettozyma wickerhamii clade+Barnettozyma salicaria | 68.17–95.36 | 87.09–96.27 | 3 |
| Barnettozyma siamensis clade | 79.96–79.96 | 91.36–91.36 | 17 |
| Barnettozyma siamensis clade+Barnettozyma botsteinii | 72.73–79.96 | 87.80–91.36 | 0 |
| Barnettozyma salicaria single–species lineage | N/A | N/A | 46 |
| Millerago | 89.67–89.67 | 92.25–92.25 | 30 |
| Millerago+Candida ficus | 74.47–89.67 | 85.02–92.25 | 5 |
| Phaffomyces | 80.33–89.98 | 92.21–94.30 | 0 |
| Phaffomyces clade | 80.33–94.80 | 91.59–96.74 | 20 |
| Cyberlindnera | 61.78–93.06 | 69.12–96.40 | 0 |
| Cyberlindnera sensu stricto clade | 67.15–90.85 | 76.63–94.72 | 2 |
| Williopsis clade | 63.63–99.97 | 78.24–99.67 | 4 |
| Candida freyschussii single–species lineage | N/A | N/A | 70 |
| Starmera | 61.57–92.68 | 67.37–97.98 | 4 |
| Starmera sensu stricto clade | 67.88–92.68 | 81.76–97.98 | 30 |
| Starmera dryadoides clade | 65.26–83.33 | 81.67–89.88 | 16 |
| Wickerhamomyces | 56.06–100.0 | 44.43–99.82 | 0 |
| Wickerhamomyces sensu stricto clade | 67.21–100.0 | 83.51–99.82 | 4 |
| Hansenula clade | 68.07–84.27 | 80.37–94.72 | 11 |
| Wickerhamomyces bovis clade | 65.53–98.19 | 82.92–96.62 | 2 |
| Wickerhamomyces pijperi clade | 71.66–81.75 | 63.80–92.13 | 87 |
| Wickerhamomyces mucosus single–species lineage | N/A | N/A | 525 |
| Wickerhamomyces mucosus+Wickerhamomyces pijperi lineage | 62.78–81.75 | 54.89–92.13 | 29 |
| Wickerhamomyces hampshirensis clade | 71.33–71.33 | 89.78–89.78 | 27 |
| Wickerhamomyces chambardii clade | 67.51–90.85 | 79.62–91.90 | 91 |
| Wickerhamomyces silvicola single–species lineage | N/A | N/A | 126 |
| Wickerhamomyces kurtzmanii single–species lineage | N/A | N/A | 280 |
Note: N/A referring to data not available
Candida species and related genus Zygoascus (Trichomonascaceae, Dipodascales, Dipodascomycetes)
Three species, Candida lundiana, Candida patagonica and Candida suthepensis, formed a well-supported clade, referred to as the Candida lundiana clade, which is nested within the genus Zygoascus (Fig. 1). This placement agrees with a previous study (Opulente et al. 2024). The genus Zygoascus was originally proposed by Smith (1986) for only the yeast species Zygoascus hellenicus, characterized by septate hyphae and hemispherical to galeate ascospores (referred to as the Zygoascus sensu stricto clade). Later, Kurtzman & Robnett (2007) transferred Pichia ofunaensis and Pichia tannicola into Zygoascus based on a multigene sequence analysis, recognizing their close relatedness to other Zygoascus species. However, unlike species comprising Zygoascus sensu stricto clade, Z. ofunaensis and Z. tannicola do not produce septate hyphae. Similarly, members of the Candida lundiana clade can be distinguished from Zygoascus sensu stricto by some physiological characteristics, such as the lack of fermentation and septate hyphae by the former. Given these distinctions, we explored the possibility of separating the Candida lundiana clade from the Zygoascus sensu stricto clade. The RED analysis conducted by Li et al. (2021) revealed substantial heterogeneity within Zygoascus and suggested that this genus is under-classified, potentially representing an order-level taxon. This conclusion was particularly based on the RED value (0.813) of this genus, which falls in the range of the order-level RED interval (viz., 0.79 ± 0.1). Our genome-based metric analyses showed that Zygoascus exhibited lower AAI (59.76–88.95%) and POCP (71.75–98.27%) values (Table 1) compared to several genera in Saccharomycetaceae studied by Liu et al. (2024a). Next to it, we calculated genomic metrics of the combined Zygoascus sensu stricto clade+Candida lundiana clade. Our genomic metrics showed that the POCP value of the two clades combined was 74.47–98.27% (Table 1), indicating that even this combined group remains too heterogeneous compared to values expected for genera based on the previous experience from Saccharomycetaceae and Metschnikowiaceae (Liu et al. 2024a, b).
Considering the evolving concept of the genus Zygoascus, the phenotypically distinguished characteristics between the Candida lundiana clade, the Zygoascus sensu stricto clade and the Zygoascus tannicola clade (Table S3) and the genome-based metric analyses, we suggest to resolve this taxonomic complexity and reduce the heterogeneity within the genus. We propose two new genera: Casaregolazyma gen. nov. to accommodate species in the Candida lundiana clade and Daia gen. nov. for members of the Zygoascus tannicola clade.
One newly published Zygoascus species, namely Zygoascus detingensis, was not included in the phylogenomic analysis because of the unavailability of its genome. Phylogenetically, this species is positioned distantly from the other Zygoascus members in our ITS+D1/D2 LSU tree (Fig. 3). Chai et al. (2022) showed that Z. detingensis occurred in a long basal to other Zygoascus species based on the combined ITS and D1/D2 LSU sequences analysis. However, our phylogenetic analysis demonstrated that Z. detingensis does not belong to any of the described clades within the Dipodascomycetes and represents a candidate monotypic genus, which can be formally proposed after more related new species are described in the future.
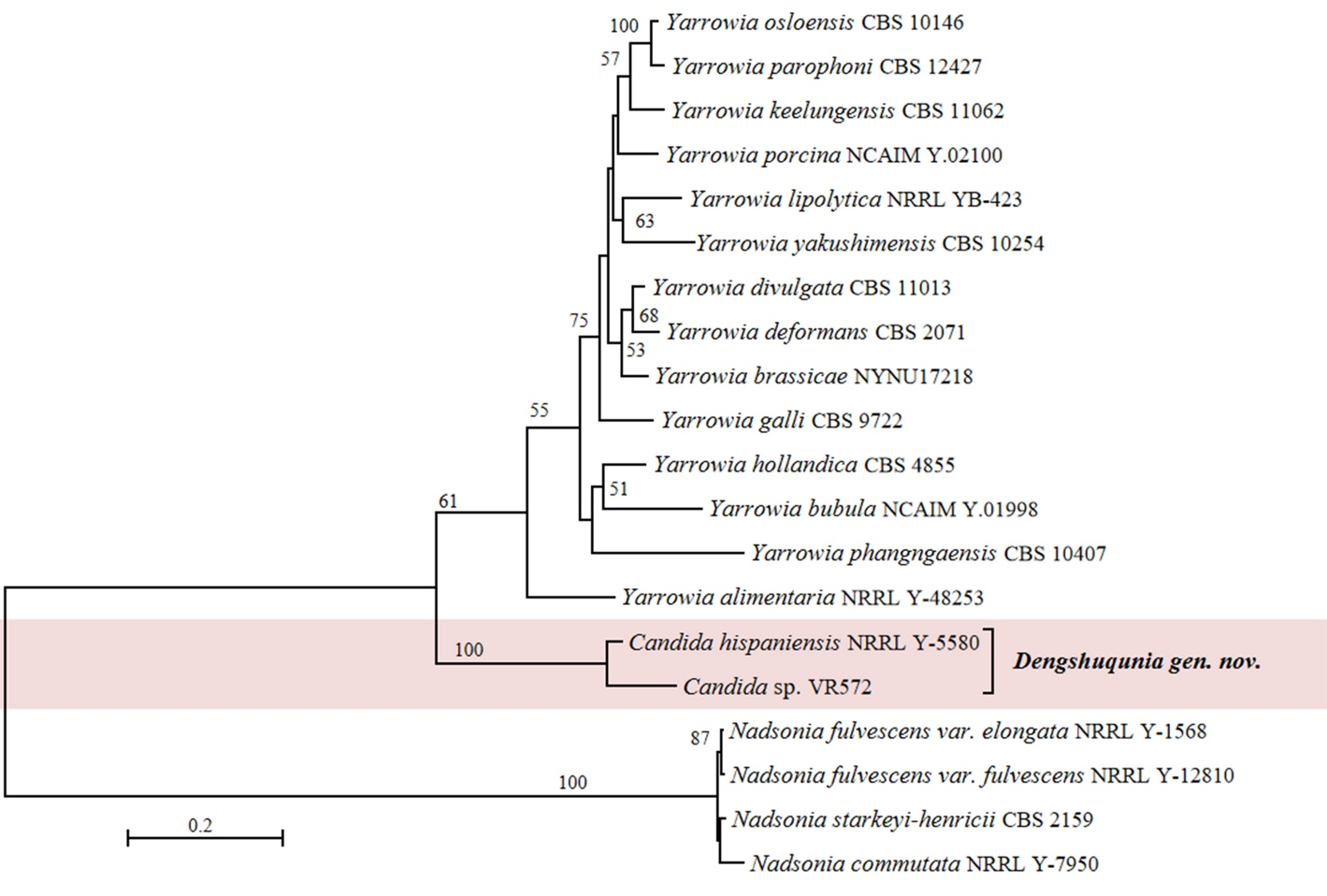
Candida species and related genera Nadsonia and Yarrowia (incertae sedis, Dipodascomycetes)
Phylogenetic analyses conducted in this study, along with previous research data (Shen et al. 2018; Opulente et al. 2024), have identified two distinct Candida lineages, namely Candida hispaniensis and Candida tunisiensis single-species lineages, within this class. The Candida hispaniensis lineage formed a long sister branch to the genera Yarrowia and Nadsonia in the phylogenomic tree (Fig. 1). While it is evident that this Candida species requires reclassification, its phylogenetic placement raises the possibility of assigning it to the genus Yarrowia. To assess the validity of this classification, we first evaluated its genome content and size. C. hispaniensis has about 11 Mb genome size and 41.69% GC content, whereas Yarrowia species have double the genome size (about 20 Mb) and 43.46–50.89% GC content compared to C. hispaniensis (Table S3). Our genome-based metric analysis showed that the POCP value of Yarrowia is 83.66–95.78% that fell in the range of the generic values (80–92%) suggested by Liu et al. (2024a), whereas the POCP value (68.29–95.78%) of Yarrowia+C. hispaniensis is lower than the recommended generic values, which suggests that it is better to place Yarrowia and C. hispaniensis in different genera, rather than to combine them in one genus. The Candida tunisiensis lineage is placed in the phylogenomic analysis as a sister group to the other known genera within Dipodascomycetes, but without a well-supported association with any specific genus (Fig. 1). In our opinion, the above analyses showed that C. hispaniensis and C. tunisiensis do not belong to any known genera in Dipodascomycetes and should be classified in two new genera instead of merging them with already existing genera. Therefore, Dengshuqunia gen. nov. is proposed for C. hispaniensis lineage. Although only one species is included in the Dengshuqunia so far, our ITS+D1/D2 LSU and D1/D2 LSU analyses showed that four potential new species belong to Dengshuqunia (Fig. 5 and Fig. S8), including organisms labelled as Yarrowia sp. VR546, Yarrowia sp. VR547, Yarrowia sp. VR571 and Yarrowia sp. VR572, which were all isolated from soils in Brazil (Table S2). The Candida tunisiensis lineage will remain a candidate genus until more closely related species are discovered in the future.
Candida species in the Pichiomycetes
One hundred forty-eight Candida species occur in the orders Pichiales and Serinales of Pichiomycetes (Table S2). The core of the genus Candida is also located in Pichiomycetes, namely in the order Serinales, family Debaryomycetaceae. Despite previous attempts to reclassify Candida, numerous species still require reclassification, the vast majority of which are placed in Pichiomycetes.
Candida species and related genera Allodekkera, Ambrosiozyma, Brettanomyces and Pichia (Pichiaceae, Pichiales, Pichiomycetes)
Candida insectalens and Candida silvatica formed a clade labelled as Candida insectalens clade, which was placed close to the genera Brettanomyces and Allodekkera (Fig. 6). Candida sorboxylosa was related to Pichia terricola in the phylogenomic tree (Fig. 6). The position of the three species was different in the ITS+D1/D2 LSU analysis. Specifically, C. insectalens and C. sorboxylosa formed two distinct branches with an affinity to the genus Komagataella, whereas C. silvatica was found to be phylogenetically more closely related to Brettanomyces, but with low bootstrap support for this placement (Fig. 7). The LSU rDNA analysis showed that these three Candida species were located in distinct branches (Fig. S9). C. silvatica and two sequences, strain Candida sp. JCM 16747 (GenBank AB552927) and environmental sequence Fungal sp. QmPlPB-1-59 (GenBank AB291684), isolated from galleries of Platypus quercivorus, the oak ambrosia beetle, in Japan, formed a C. silvatica clade without bootstrap support. C. sorboxylosa formed a very long branch near the C. silvatica clade. C. insectalens and Fungal sp. QmPlEG-2-8 (GenBank AB291677) isolated from the gallery of P. quercivorus in Japan formed a long branch with low bootstrap support (Fig. S9). The phylogenetic position of these yeasts remained unresolved in previous studies that utilized solely rDNA sequences in their phylogenetic analyses. Kurtzman & Robnett (1998) placed C. sorboxylosa as a sister species to C. silvatica, indicating its affinity with Dekkera species. They also positioned C. insectalens and C. incommunis together based on a D1/D2 LSU sequence analysis. In contrast, Sugita & Nakase (1999) demonstrated that C. insectalens was located in a basal position next to the Starmerella clade based on a phylogenetic analysis using SSU rDNA sequences. Suzuki & Nakase (2002) indicated that C. sorboxylosa formed a long branch near Saturnispora species and Pichia membranifaciens. More recently, Lachance et al. (2011) showed that C. insectalens occurred on a long branch next to C. sorboxylosa using a phylogenetic analysis of D1/D2 LSU rDNA data. In the multi-locus analysis based on the combined rDNA and TEF1 sequence, C. sorboxylosa was more closely related to Komagataella (Kurtzman et al. 2008). Considering that the phylogenomic analysis is usually more reliable than the rDNA and multigene-based phylogenetic analyses, we assign C. sorboxylosa to the genus Pichia and propose a new genus Xiuguozyma gen. nov. to accommodate C. insectalens and C. silvatica.
Candida boidinii was located at a branch positioned basal to Allodekkera, Ambrosiozyma, Brettanomyces, Kregervanrija, Martiniozyma, Ogataea, Pichia, and Saturnispora (Fig. 6), which suggests that this single-species lineage represents a genus. This species has been resolved as a basal single-species lineage to Ogataea in a previous study (Kurtzman & Robnett 2010). Therefore, Ramirezia gen. nov. is proposed to accommodate C. boidinii. In the search for potential members of the novel genus, more than 92 ITS sequences of C. boidinii, such as CBS 6056 (GenBank KY101981), and four nucleotide sequences (GenBank EF060568, EF060866, EF060905 and EF060925) were retrieved from public databases, showing 97–100% sequence similarity with the type strain of C. boidinii. Sequences with lower similarity values may correspond to species different from C. boidinii. Our ITS phylogenetic analysis showed that at least two groups, namely group 1 and group 2, represent two potential new species closely related to C. boidinii (Fig. S10).
Candida awuaii was located within the genus Pichia (Fig. 7) and is therefore transferred to Pichia. Similarly, Candida wuzhishanensis was positioned within the genus Ambrosiozyma, showing a close relationship to Ambrosiozyma pseudovanderkliftii with 99% and 100% sequence similarity in the ITS and D1/D2 LSU regions, respectively. This high similarity strongly suggests that C. wuzhishanensis is a synonym of A. pseudovanderkliftii (Fig. 8).
Candida species and related genus Ogataea (Pichiaceae, Pichiales, Pichiomycetes)
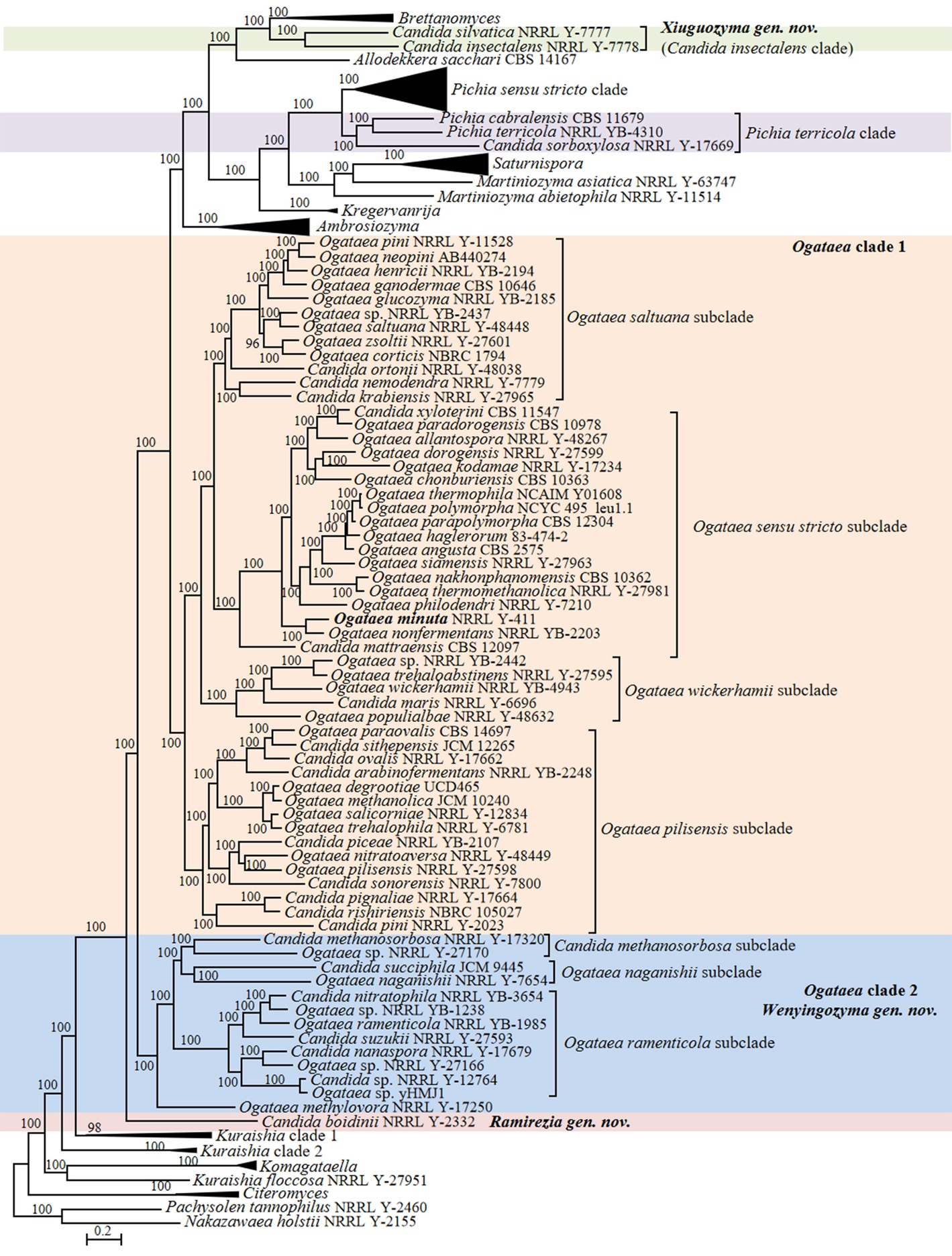
Our phylogenetic analyses showed that 20 Candida species were placed in the genus Ogataea in Pichiales (Figs. 6, 8, Table S2). The genus Ogataea is mainly comprised of methanol-assimilating yeasts. Since the erection of the genus to accommodate hat-shaped ascospore-forming, nitrate-assimilating Pichia species (Yamada et al. 1994a), the circumscription and size of the genus varied with more species being transferred to Ogataea or newly discovered in nature. Kurtzman & Robnett (2010) demonstrated a distant relatedness between Ogataea and Pichia, and transferred eight species to Ogataea and the newly erected genus Peterozyma. The authors employed a multi-locus phylogeny to resolve subclades of Ogataea and distinguish them from Ambrosiozyma. Previous studies that used rDNA sequences only, failed to resolve Ogataea and Ambrosiozyma, but Glushakova et al. (2010) used a combined rDNA phylogeny and advanced maximum-likelihood-based statistics, confirming the observations by Kurtzman & Robnett (2010) regarding the monophyly of Ogataea and the clade represented by Pichia methanolica, Pichia trehalophila, and Williopsis salicorniae. Later, Kurtzman (2011a) argued that the clade represented by Ogataea naganishii may be separated from Ogataea as a sister genus when a more robust dataset becomes available. One of the molecular features of yeasts comprising this phylogenetic group that complicates the phylogenetic analyses and species demarcation is the low variability of rDNA sequences. Therefore, whole-genome sequencing seems a good opportunity to reassess the boundaries of the genus and the relatedness of the species presently accommodated in Ogataea. More recently, a large phylogenomic analysis revealed complex relationships between Ogataea and Ambrosiozyma, suggesting that Ogataea is polyphyletic (Shen et al. 2018). The clade that is comprised by Candida succiphila, O. naganishii, Ogataea ramenticola, and Ogataea methylivora occupied a basal position to the core of Ogataea and Ambrosiozyma. The same phylogenetic relationship has been observed in the subsequent study conducted by Opulente et al. (2024), who additionally identified Candida methanosorbosa, Candida nanaspora, Candida nitratophila, and Candida suzuki as the members of that basal clade. The phylogenomic analysis performed in our study confirmed the previous observations regarding the polyphyly of the genus Ogataea that was split into two large clades labelled here as Ogataea clade 1 and Ogataea clade 2 (Fig. 6).
Further examination of the genus Ogataea as currently accepted, using the genome-based metric analyses, demonstrated that the genus is heterogeneous (Table 1). Specifically, the ranges of AAI values (56.34–99.54%) and POCP values (55.36–97.74%) were substantially lower than that of the genera recognized in the Saccharomycetaceae (Liu et al. 2024a). The RED value calculated by Li et al. (2021) was 0.829, which suggests that this genus is likely under-classified and may represent a family-level or order-level taxon. The PAPO value is 0 and no common gene (viz., unique gene) has been found across the clade, indicating that this genus is too heterogeneous and likely under-classified.
Based on the above arguments, we propose to split the genus Ogataea and propose a new genus Wenyingozyma gen. nov. to accommodate the Ogataea clade 2 that contains six Candida species. A total of fourteen Candida species located in the Ogataea clade 1 are transferred to the genus Ogataea (Fig. 6 and Table S2). To test for the degree of genomic heterogeneity for this proposed reclassification, we determined indices values for that Ogataea clade 1 (AAI: 58.82–99.54%; POCP: 61.91–97.74%; PAPO: 0) and Ogataea clade 2 (AAI: 58.46–90.74%; POCP: 68.35–97.84%; PAPO: 6). Both clades are characterized by higher genetic diversity compared to the genera accepted in Saccharomycetaceae (Liu et al. 2024a) and Pichiales (Table 1), e.g., Citeromyces, Komagataella and Kregervanrija. Our phylogenetic analysis identified four subclades in the Ogataea clade 1, namely the Ogataea sensu stricto subclade, the Ogataea pilisensis subclade, the Ogataea saltuana subclade, and the Ogataea wickerhamii subclade (Fig. 6). The four subclades were characterized by the following genomic metrices, the Ogataea sensu stricto subclade (AAI: 64.30–99.54%; POCP: 76.83–97.74%; PAPO: 2) including two Candida species; the Ogataea pilisensis subclade (AAI: 61.27–90.18%; POCP: 75.32–97.46%; PAPO: 0) including eight Candida species; the Ogataea saltuana subclade (AAI: 66.24–85.70%; POCP: 86.12–97.29%; PAPO: 1) including three Candida species; and the Ogataea wickerhamii subclade (AAI: 65.49–86.36%; POCP: 87.03–94.58%; PAPO: 5) including one Candida species, namely Candida maris. In the Ogataea clade 2, several well-supported subclades were detected in the phylogenomic analysis (Fig. 6), namely the Ogataea naganishii subclade (AAI: 58.98%; POCP: 70.68%; PAPO: 0) including two Candida species, the Candida methanosorbosa subclade (AAI: 63.02%; POCP: 78.80%; PAPO: 0) including one Candida species, the Ogataea ramenticola subclade (AAI: 66.21–90.74%; POCP: 82.94–97.84%; PAPO: 32) including three described Candida species, and one single-species lineage represented by Ogataea methylovora. The multigene analysis, including LSU rDNA, SSU rDNA, TEF1 and mitochondrial SSU rDNA genes, positioned C. methanosorbosa distantly related from C. succiphila and O. naganishii (Kurtzman & Robnett 2010). The above genome metric analyses showed that the Ogataea clade 1 and the Ogataea clade 2 are likely too heterogeneous and should be considered separate genera. However, more robust analyses and careful evaluation by community experts are needed to confirm their taxonomic conclusions in the future.
Candida species and related genera Hemisphaericaspora, Lodderomyces, Nematodospora and Spathaspora (Debaryomycetaceae, Serinales, Pichiomycetes)
The order Serinales currently includes families Cephaloascaceae, Debaryomycetaceae and Metschnikowiaceae. The family Debaryomycetaceae contains more than 109 Candida species, including the type species, C. vulgaris, which is now considered a synonym of C. tropicalis (Lachance et al. 2011). The phylogenetic lineage bearing the type species is often referred to as the Candida/Lodderomyces lineage (or clade), which currently contains 42 Candida species, including most clinically important ones (Figs. 9–10 and Table S2). Our phylogenomic analysis revealed four clades in the lineage, namely the Candida sensu stricto clade (AAI: 69.03–99.45%; POCP: 75.64–98.28%; PAPO: 7), the Candida corydali clade (AAI: 67.27–81.01%; POCP: 81.71–93.33%; PAPO: 3), the Lodderomyces clade (AAI: 64.25–83.42%; POCP: 73.73–96.02%; PAPO: 4), and the Nematodospora clade (AAI: 76.99%; POCP: 94.83%; PAPO: 5) (Fig. 9, Table 1). For a long time, the lineage included only one known sexual species, Lodderomyces elongisporus. The genus was proposed by van der Walt (1966), who reclassified the species Saccharomyces elongisporus, placing it in a lineage closely related to Candida parapsilosis. The genus Nematodospora was later established in the Candida/Lodderomyces lineage to accommodate a distinct yeast species characterized by a unique ascospore morphology, distinguishing it from Lodderomyces (Gouliamova et al. 2016). Although the genus Nematodospora was placed close to Lodderomyces, both currently known species exhibit an ascospore morphology distinct from that of Lodderomyces (Gouliamova et al. 2016; Ren et al. 2016). In the phylogenomic analysis, the Lodderomyces and the Candida corydali clades clustered together closely to Nematodospora, whereas the Candida sensu stricto clade was positioned as a sister group to Nematodospora, the Candida corydali clade, and the Lodderomyces clade (Fig. 9). This topology is consistent with previous phylogenomic analyses (Shen et al. 2018; Opulente et al. 2024). As the result, the phylogenomic analysis reveals four clades, which are harbouring nomenclature types of three genera, namely Candida, Lodderomyces and Nematodospora. This Candida corydali clade puts it in the position of a sister taxon to the Candida sensu stricto clade and also makes the genus polyphyletic.
The phylogenetic lineage Candida/Lodderomyces showed a rather low range of POCP values (65.88–98.28%), indicating its greater genetic heterogeneity compared to most genera in Saccharomycetacea (Liu et al. 2024a). Consequently, we further examined the genetic metrics in the Candida/Lodderomyces lineage to explore available options for a meaningful and statistically supported reclassification. Based on the phylogenomic tree, one potential option is to create a large clade that would comprise the Candida corydali and the Lodderomyces clades. However, the POCP (68.47–96.02%) and PAPO (0, without unique genes) values of the combined Candida corydali clade+Lodderomyces clade (Table 1) suggest that it is preferable to recognize those two clades as distinct genera, rather than combine them into a single genetically heterogeneous genus. At the moment, the Candida corydali clade contains 10 Candida species, six of which were isolated from insects (Nguyen et al. 2007; Lachance et al. 2011; Liu et al. 2016), while the remaining three species were isolated from leaves of Pterocarpus indicus, flowers of Verbascum, and a mushroom (Nakase et al. 2009; Limtong et al. 2012; Sipiczki 2013). Considering the origin of species in the Candida corydali clade, we propose the new genus Insectozyma gen. nov. to accommodate these yeasts and transfer 10 Candida species to this genus. Additionally, eight Candida species, including the clinically-relevant C. parapsilosis and two Lodderomyces species, clustered in the Lodderomyces clade (Fig. 9, Table S2). Consequently, we transfer them into the Lodderomyces genus. The genus Nematodospora, comprising two species, forms an isolated clade in Candida/Lodderomyces lineage. The Candida sensu stricto clade, which includes 22 species, including the clinically relevant species C. albicans, C. dubliniensis, and C. tropicalis, forms a well-supported clade in the phylogenomic tree. Furthermore, the analysis revealed a clade comprising Candida parablackwelliae, Candida blackwelliae and Spathaspora boniae (i.e., the Candida blackwelliae clade), which is positioned basally to the Candida/Lodderomyces lineage (Fig. 9).
At the time of description, phylogenetic and phylogenomic analyses placed Spathaspora boniae outside the Spathaspora clade in a basal position to the Candida/Lodderomyces lineage (Morais et al. 2017). The same authors concluded that the genus Spathaspora as currently defined is paraphyletic, but nonetheless opted to describe the new species as Spathaspora boniae in that genus. This Candida blackwelliae clade is well-defined based on the phylogenomic analysis and genome-based metric analyses with AAI, POCP and PAPO values of 70.58–90.56%, 89.31–96.41% and 8, respectively. Both, phylogenetic position of the clade and the range of genomic metrics suggest this is a good candidate genus. Hence, the new genus Zhuliangozyma gen. nov. is proposed for this clade.
Candida alai was located in a long branch close to the Spathaspora sensu stricto clade (Fig. 9). Candida alai differs from species of the Spathaspora sensu stricto clade by the lack of assimilation of erythritol and N-acetyl-D-glucosamine (Table S3). The above physiological and phylogenetic analyses indicated that C. alai do not belong either to the genus Candida, nor to Spathaspora. Therefore, the new genus Intestinozyma gen. nov. was proposed to accommodate C. alai. The sequence of a yeast labelled as Candida sp. B53C (GenBank MW165503), isolated from an insect gut in Brazil, differs by 11 nt ITS sequence differences from the sequence of C. alai (Fig. S11). This placement suggests that Candida sp. B53C may represent another new member of the genus Intestinozyma, which is currently only represented by C. alai.
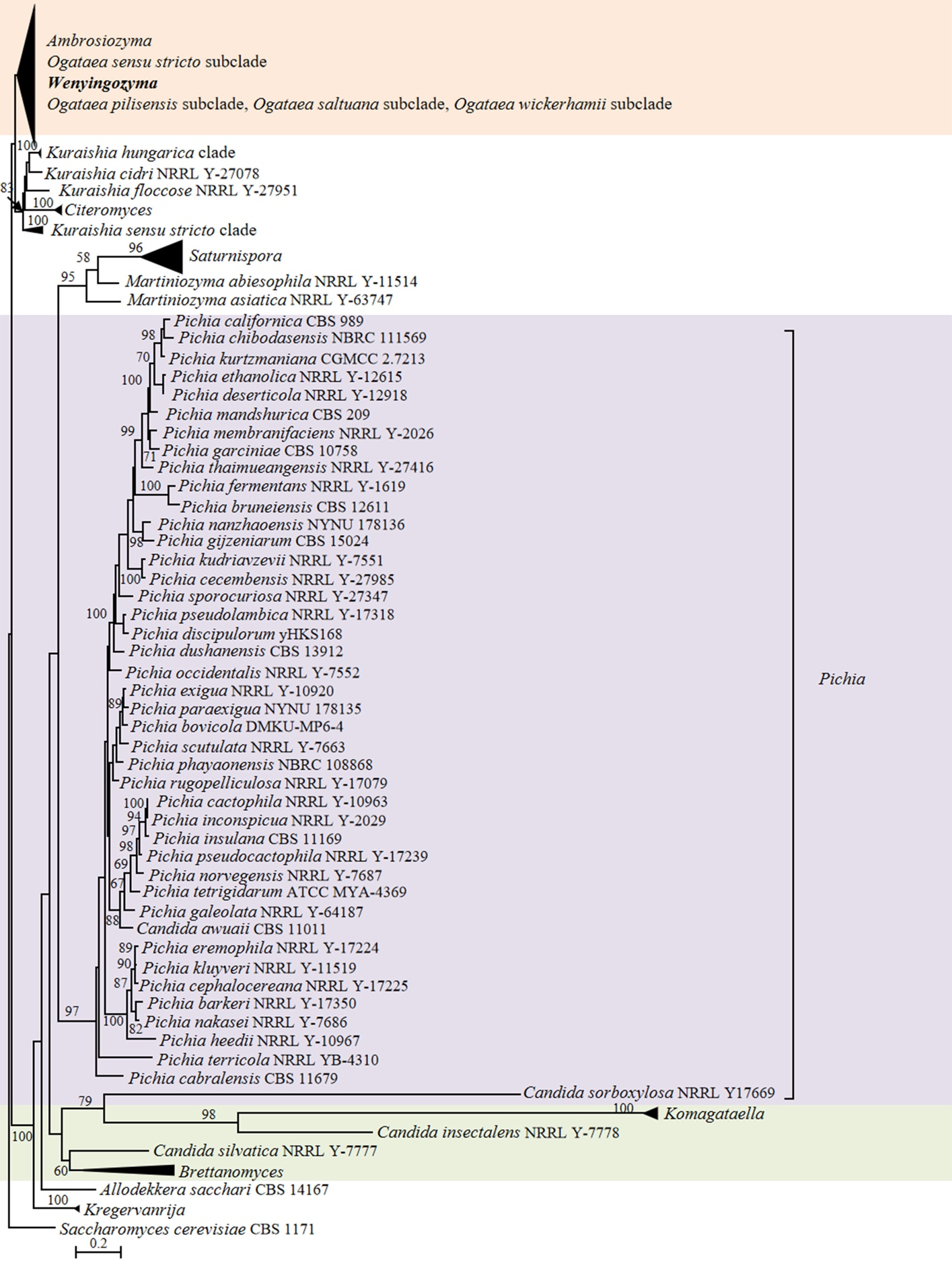
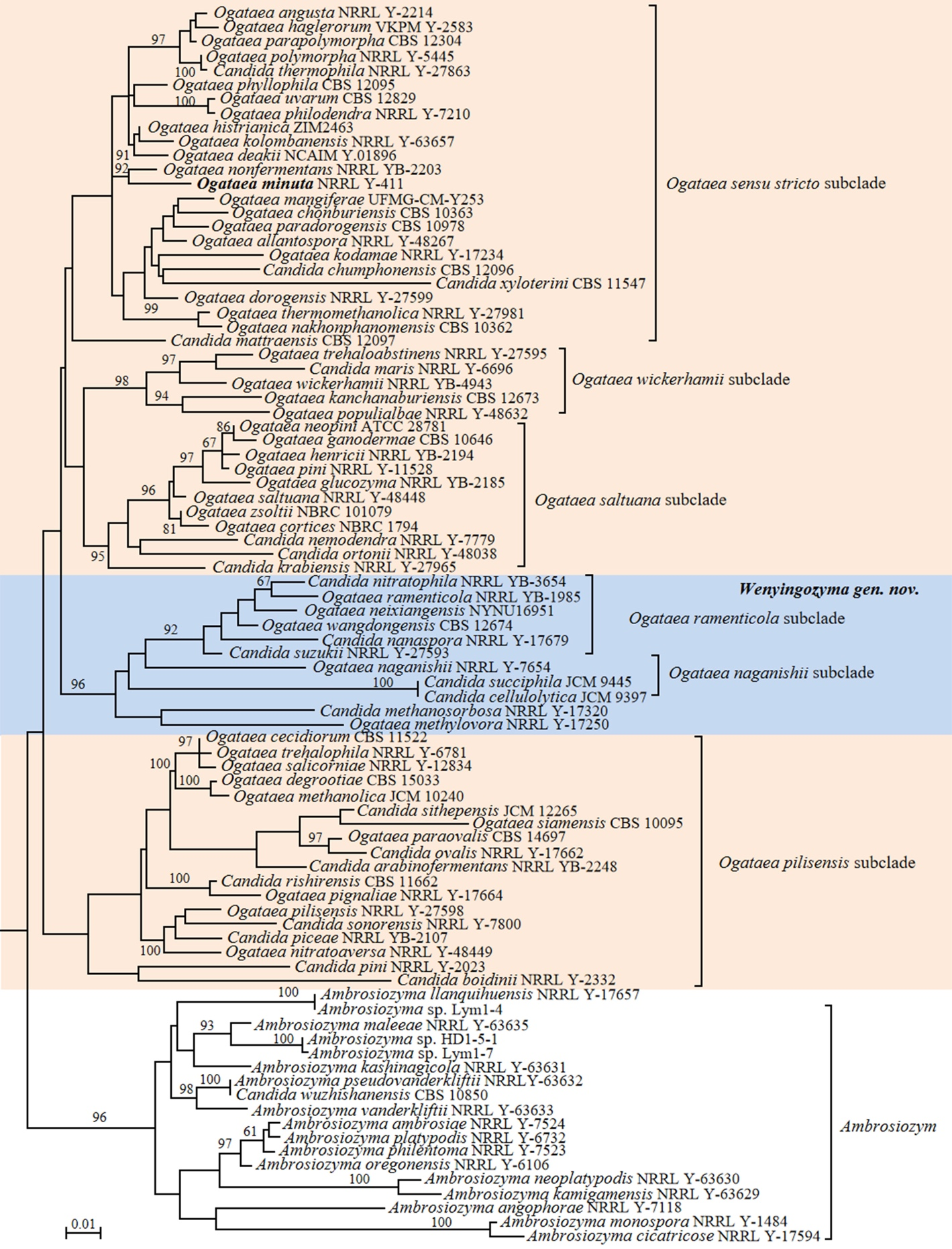
In agreement with previous observations (Opulente et al. 2024), Candida lyxosophila, Candida subhashii and Candida xylanilytica, and three Spathaspora species were located in the Hemisphaericaspora clade (Figs. 9–11). Consequently, we suggest transferring those species into the genus Hemisphaericaspora.
Candida species and related genus Diutina (Debaryomycetaceae, Serinales, Pichiomycetes)
The Candida glaebosa clade includes seven Candida species that are closely related to the genus Diutina (Fig. 9). Members of the clade share the following features with a lower GC content (31.16–41.24%), whereas the genus Diutina has a higher GC content (41.23–53.05%) (Table S3). The genome metrics analyses showed that the Candida glaebosa clade (AAI: 65.65–86.35%; POCP: 82.55–96.03%; PAPO: 23) is a good candidate genus in terms of its genomic heterogeneity, as demonstrated by its AAI and POCP values that are within the ranges of those previously observed for Saccharomycetaceae and Metschnikowiaceae (Liu et al. 2024a, b). Therefore, Suzukiozyma gen. nov. is proposed to accommodate members of the Candida glaebosa clade
Candida species and related genera Suhomyces, Teunomyces and Wickerhamia (Debaryomycetaceae, Serinales, Pichiomycetes)
Candida caryicola and Candida tibetensis formed the Candida tibetensis clade, which is positioned basal to the genus Suhomyces (Fig. 9). Our genome-based metric analyses showed that the AAI and POCP values of Suhomyces+Candida tibetensis clade were 63.89–96.44% and 84.33–98.34%, respectively, which fell in the range of generic AAI and POCP values previously reported for Saccharomycetaceae by Liu et al. (2024a) and supported of the transfer of C. caryicola and C. tibetensis to Suhomyces. New combinations for these two species are provided in the Taxonomy section.
Candida sake was placed in a long branch closely related to Teunomyces (Fig. 9). The species C. sake has a 14 Mb genome size with GC 38.76%, whereas Teunomyces has a low genome size (10–13Mb) and higher GC content (41.03–46.16%) (Table S3). The above analyses show C. sake do not belong either to the genus Candida, nor to any presently known genera. Therefore, the most pragmatic solution is to accommodate C. sake in the genus Fermentozyma gen. nov. Crous et al. (2017) showed that Candida vespimorsuum was closely related to C. sake with which it formed a well-supported clade, suggesting that they might belong to the same genus. Our ITS+D1/D2 LSU rDNA phylogenetic analysis supports the observation that C. vespimorsuum and C. sake belong to the same new genus Fermentozyma (Fig. 10). Another potential new species in that genus is represented by the strain Candida sp. KBP Y-6292 (GenBank OP941477), isolated from ants in Vietnam, is placed in Fermentozyma with good statistical support (Fig. 10).
Candida anutae and Candida argentea formed two distinct long branches closely related to Wickerhamia fluorescens (Fig. 9). In the absence of other closely related species, C. anutae and C. argentea were retained as Candida pro tempore at present. Because of their distant placement from the core of the genus Candida, they will be reclassified into new genera after more closely related taxa are found.
Candida species and related genera Cephaloascus and Kurtzmaniella (Debaryomycetaceae, Serinales, Pichiomycetes)
The phylogenomic analysis revealed that Candida chilensis was placed in a long branch closely related to the genus Cephaloascus (Fig. 12). Given this distant placement from the type clade of Candida and other hitherto described genera, we propose Nothofagozyma gen. nov. to accommodate C. chilensis. Our D1/D2 LSU rDNA phylogenetic analysis showed that strain Candida cf. chilensis CBS 11766 (GenBank FN824503) isolated from soil in Germany (Yurkov et al. 2012), is closely related to, but distinct from C. chilensis, and appears to represent a new species of Nothofagozyma, currently only represented by C. chilensis (Fig. S9). The rDNA blast against the NCBI nucleotide database showed that CBS 11766 differed from C. chilensis by 10 nucleotides (1.8%) in the D1/D2 LSU sequences.
According to our phylogenomic analyses, six Candida species and three Candida species were placed in the Candida railenensis clade and the Candida aurita clade, respectively. Both clades were found to be related to the genus Kurtzmaniella (Fig. 12). Lachance et al. (2011) and Daniel et al. (2014) suggested that Candida anglica, Candida boleticola, Candida fragi, Candida oleophila, C. railenensis, Candida santamariae, Candida schatavii and Candida zeylanoides were members of the Kurtzmaniella clade. Lopes et al. (2019) transferred C. fragi, C. quercitrusa and C. natalensis to the genus Kurtzmaniella, and argued that reclassification of C. anglica, C. boleticola, C. oleophila, C. railenensis, C. santamariae, C. schatavii and C. zeylanoides into Kurtzmaniella needed more robust data, because C. schatavii was found to be highly divergent from Kurtzmaniella and placed on a long branch to the clade (Shen et al. 2018). In order to evaluate the relative heterogeneity of the Kurtzmaniella clade, genomic metrics were calculated. The AAI and POCP values of the large clade, Kurtzmaniella+the Candida railenensis clade+Candida aurita clade, are 58.77–99.43% and 67.61–98.15%, respectively, which are lower than the values observed in well-defined genera in Saccharomycetaceae and Metschnikowiaceae (Liu et al. 2024a, b) and in other genera in Debaryomycetaceae (Table 1). Considering a larger genus size and heterogeneity compared to other genera in the family, we prefer not to merge the Candida railenensis clade and the Candida aurita clade with Kurtzmaniella, but keep them as separate genera. This option is supported by our phylogenomic and rDNA phylogenetic analyses that showed that the Candida railenensis clade (AAI: 63.07–99.43%; POCP: 81.11–97.82%; PAPO: 4) and the Candida aurita clade (AAI: 67.97–93.93 %; POCP: 79.72–98.15 %; PAPO: 11) are distinct from Kurtzmaniella (Figs. 12–13). We explored the option of accommodating these Candida species in a single larger genus. The members of the two Candida clades are solely asexual morphs, but differ from each other by growth on 0.1% cycloheximide (Table S3). The POCP value of the Candida railenensis clade+Candida aurita clade was 74.04–98.15%, which is lower than the values observed in well-defined and generally accepted genera in Saccharomycetaceae (Liu et al. 2024a), thus suggesting that the Candida aurita clade and the Candida railenensis clade are better to accommodate in two genera. Therefore, Chernovozyma gen. nov. and Dujonia gen. nov. are proposed for the Candida aurita clade and the Candida railenensis clade, respectively. The single-species lineage C. anglica was closely related to Kurtzmaniella in our phylogenomic analysis (Fig. 12), but this species was placed on a basal branch related to the Candida railenensis clade and the Candida aurita clade (Opulente et al. 2024). Therefore, C. anglica is assigned as Candida pro tempore at present, which has to be resolved in the future.
Candida species and related genera Debaryomyces, Millerozyma and Schwanniomyces (Debaryomycetaceae, Serinales, Pichiomycetes)
Candida glucosophila formed a long branch related to Debaryomyces singareniensis and Schwanniomyces etchellsii (Fig. 12). Candida multigemmis was located at a long branch without any genera as close relatives (Fig. 12). As the above two Candida species were characterized as separated long branches or clades, they do not seem to belong to any known genera or assigned clades in the Serinales. Therefore, it is pragmatic to accommodate them in new genera. The name Glucitozyma gen. nov. is proposed for C. multigemmis. A search for sequences of potential new species in GenBank resulted in Candida sp. CPD-35-1 (GenBank MZ701688), which is different from C. multigemmis by 14 nt (2%) in the ITS region and represents a potential new member of Glucitozyma gen. nov. (Fig. S12). In contrast, single species-lineage C. glucosophila will be assigned as Candida pro tempore due to the current lack of close relatives.
Candida thasaenensis, Scheffersomyces gosingicus and Scheffersomyces spartinae formed a clade in the phylogenomic tree (i.e., the Scheffersomyces gosingicus clade), which is more closely related to Priceomyces and Schwanniomyces than to the Scheffersomyces sensu stricto clade (Fig. 10). An observation that is consistent with the analysis by Opulente et al. (2024). Our ITS+LSU rDNA analysis showed that C. thasaenensis, clustered with S. gosingicus and S. spartinae with 100% bootstrap support, forming a clade closely that is related to the Candida glaebosa clade, and yet distinct from the other Scheffersomyces species (Fig. 11). The above rDNA-based and phylogenomic analyses suggested that the Scheffersomyces gosingicus clade should be accommodated in a distinct genus for which we propose the name Lizanozyma gen. nov. Consequently, C. thasaenensis, S. gosingicus and S. spartinae will be transferred to this newly created genus in the Taxonomy section below.
Four more Candida species were located in known genera in the Serinales. Specifically, Candida broadrunensis, Candida pseudofarinosa, Candida psychrophila and Candida rongomai-pounamu were placed in the genera Scheffersomyces, Millerozyma and Debaryomyces, respectively (Figs. 12–13).
Candida species and related genus Yamadazyma (Debaryomycetaceae, Serinales, Pichiomycetes)
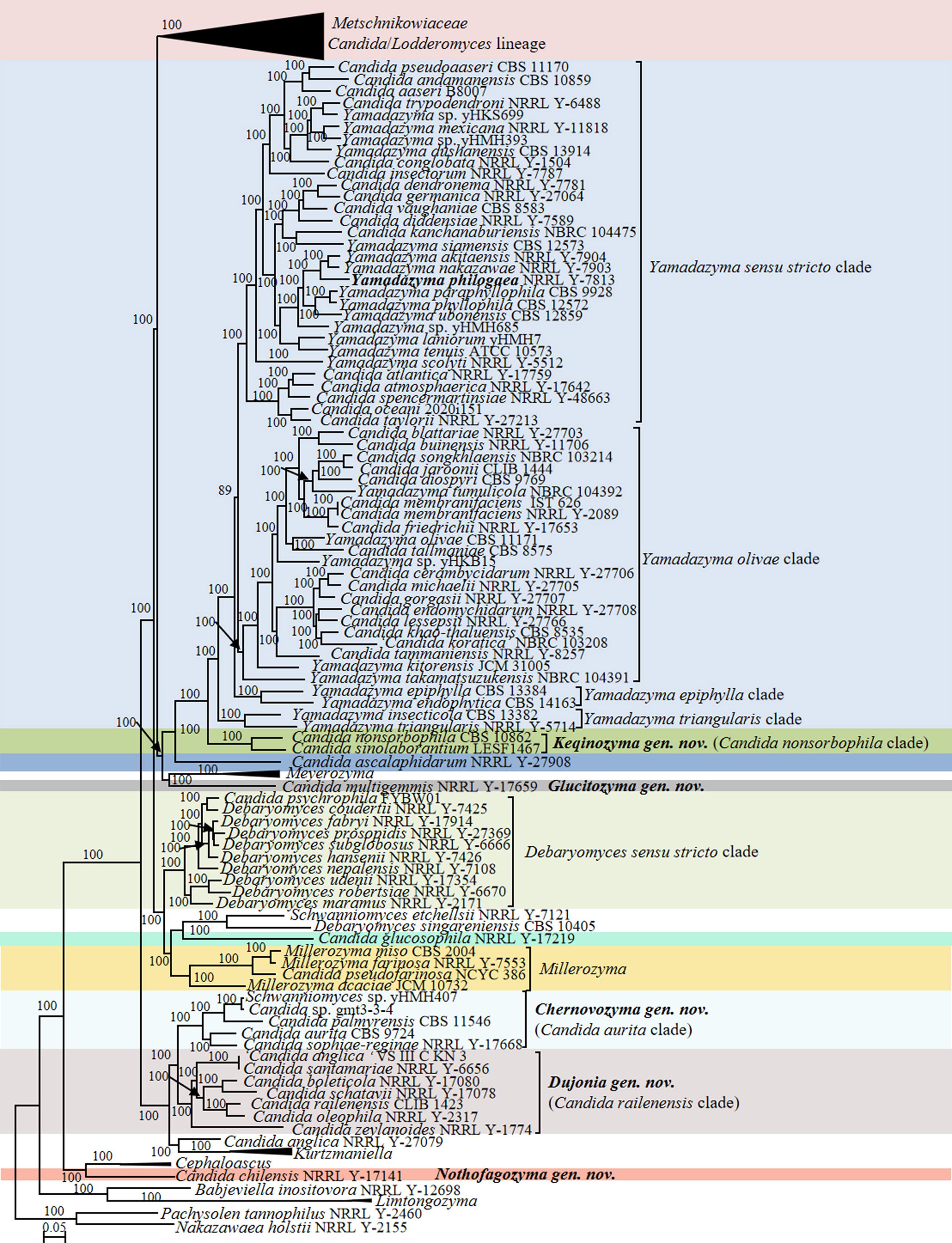
Twenty-eight Candida species were distributed across the genus Yamadazyma in the phylogenomic analysis (Figs. 12–13, Table S2). The RED value (0.814) of Yamadazyma calculated by Li et al. (2021) suggested that the genus Yamadazyma is likely under-classified and may represent an order-level taxon. We further assessed the heterogeneity of this genus using the genome-based metrics. Our analyses confirmed that the genus Yamadazyma is heterogeneous, as indicated by the lower than expected ranges of metrics (AAI: 59.65–90.55%; POCP: 73.35–97.21%; PAPO: 0). The phylogenomic analysis revealed four distinct clades, namely the Yamadazyma sensu stricto clade, the Yamadazyma epiphylla clade, the Yamadazyma olivae clade, and the Yamadazyma triangularis clade. Our analyses suggest that the four clades may correspond to genus-level taxa, as suggested by the ranges of genomic metrics such as Yamadazyma olivae clade (AAI: 62.06–97.75%; POCP: 77.72–98.83%; PAPO: 0), the Yamadazyma epiphylla clade (AAI: 69.16%; POCP: 89.77%; PAPO: 5), and the Yamadazyma triangularis clade (AAI: 69.45%; POCP: 87.92%; PAPO: 14). The delimitation and composition of the genus Yamadazyma, including potential competing names in the genus, will be carefully discussed in the community of yeast taxonomists before undertaking a large-scale reclassification.
Eight Candida species were located in the Yamadazyma sensu stricto clade and should therefore be transferred to the genus Yamadazyma. Recently, Avesani et al. (2024) described two new Yamadazyma species and transferred 11 Candida species into Yamadazyma. These two new species and 11 new combinations are all placed in the Yamadazyma sensu stricto clade, which is supported by this study. However, five new combinations proposed by Avesani et al. (2024) in the Yamadazyma sensu stricto clade were invalid (https://www.indexfungorum.org/). Therefore, we will validate these five species in the genus Yamadazyma in the Taxonomy section. Eighteen Candida species were placed in the Yamadazyma olivae clade (Fig. 13). Among them, more recently, 15 species have been transferred to Yamadazyma (Qiu et al. 2025), but three ones are still in the genus Candida. Considering the consistency with the above study, we will transfer those three species into Yamadazyma in the Taxonomy section.
Candida ascalaphidarum, Candida nonsorbophila and Candida sinolaborantium were found to be closely related to Yamadazyma (Fig. 12). Candida ascalaphidarum was located in a separate branch, whereas C. nonsorbophila and C. sinolaborantium formed a well-supported Candida nonsorbophila clade, which indicated that those two lineages may represent two new genera. Here, we just propose Keqinozyma gen. nov. for Candida nonsorbophila clade, but assigned C. ascalaphidarum as Candida pro tempore due to the current lack of close relatives. Considering species for which no genome data is available, Candida heliconiae and Candida temnochilae are closely related to Candida nonsorbophila clade in an ITS+D1/D2 LSU rDNA phylogenetic analysis (Fig. 13) and will be recombined in the genus Keqinozyma (see Taxonomy below).
Candida species and related genus Metschnikowia (Metschnikowiaceae, Serinales, Pichiomycetes)
The re-classification of Candida species and related genera in the family Metschnikowiaceae has been done recently by Liu et al. (2024b) and resulted in 13 new genera to accommodate species that are distantly related to the core Candida-clade in Debaryomycetaceae. However, for pragmatic reasons, the circumscription of the large genus Metschnikowia was maintained in that study and will be reconsidered in a future study. This decision particularly concerns the classification of Candida wancherniae in the M. agaves clade, Candida hawaiiana in the M. arizonensis clade, Candida golubevii and Candida magnifica in the M. bicuspidata clade, Candida hainanensis in the M. caudata clade and the single-species Candida danieliae in the Metschnikowia lineage (Fig. 14, Fig. S1 in Liu et al. 2024b). These species will be kept pro tempore in Candida. The position of Candida citri and Candida xylosifermentans was also not resolved by Liu et al. (2024b) and we refrain also here from any taxonomic changes until more closely related species are described.
Candida species in the Saccharomycetes
In the phylogenomic analysis, a total of 31 Candida species in the Phaffomycetales (Saccharomycetes) have been placed close to the genera Barnettozyma, Cyberlindnera, Phaffomyces, Starmera and Wickerhamomyces (Fig. 15). The results of the analysis revealed several taxonomic conflicts and showed that several large genera in the order Phaffomycetales are polyphyletic. The genera Barnettozyma, Millerago, and Phaffomyces formed a well-supported clade, in which Millerago was nested inside a polyphyletic genus Barnettozyma (Fig. 15). Phaffomyces formed a well-supported clade, but members of the genus Barnettozyma were placed in three different clades. Two Candida species, i.e., Candida coquimbonensis and Candida orba, clustered within the genus Phaffomyces, suggesting they should be transferred to this genus.
Candida species and related genera Barnettozyma, Millerago and Phaffomyces (Phaffomycetaceae, Phaffomycetales, Saccharomycetes)
The Barnettozyma sensu stricto clade includes the type of the genus, Barnettozyma populi, and the type of the former genus Zygowilliopsis (Kudryavtsev 1960), Zygowilliopsis californica (current name Barnettozyma californica), which is also the type species of the genus Zygohansenula. Candida sanyiensis clustered in the Barnettozyma sensu stricto clade, suggesting this species should be transferred to this clade. Whether the names Barnettozyma, Zygohansenula or Zygowilliopsis will be selected for this clade will be decided after discussions with the yeast taxonomic community. Consequently, C. sanyiensis is kept as pro tempore in the genus Candida at present. Other species were placed in three subclades that formed a well-supported clade with Phaffomyces. The subclades included (i) Barnettozyma siamensis, Barnettozyma botsteinii, and Candida montana; (ii) Millerago and Candida ficus; (iii) Barnettozyma pratensis, Barnettozyma salicaria, Barnettozyma wickerhamii, Candida norvegiva and Candida qinlingensis. The phylogenetic analyses convincingly demonstrated the taxonomic conflict between Barnettozyma, Millerago, and Phaffomyces, and additionally showed that the genus Barnettozyma in its present circumscription is not monophyletic. When restricted to its core, the composition of Barnettozyma (= Zygowilliopsis = Zygohansenula) must be revised, addressing the classification of the above subclades. This is also supported by the genomic metrics for the Barnettozyma clade showing AAI values of 64.52–86.96% and POCP values of 81.56–95.37%, both falling within the range of the generic boundaries as observed by Liu et al. (2024a). However, the absence of unique genes (PAPO: 0) suggests that Barnettozyma might be phylogenetically heterogeneous. Here, we refrain from further taxonomic decisions on the matter and prefer to discuss this with the broader yeast taxonomy community.
Candida ficus and the genus Millerago clustered together in the phylogenomic tree (Fig. 15). Recently, García-Acero et al. (2024) proposed the new genus Millerago to accommodate Candida galis and the newly described species Millerago phaffii based on the ITS+LSU rDNA sequences and phylogenomic analyses. García-Acero et al. (2024) did not place C. ficus in the genus Millerago because this species formed a long branch near Millerago in the D1/D2 LSU rDNA-based tree and suggested leaving C. ficus in its current taxonomic position and resolving this in the future. A recent phylogenomic analysis demonstrated that C. ficus and C. galis form a well-supported clade, with genetic distances visually comparable to those in neighboring clades (Opulente et al. 2024). To determine whether these two species could be grouped into a single clade, we estimated genomic metrics for this lineage (Table 1). The metrics for Millerago+Candida ficus showed the following ranges: AAI (74.47–89.67%), POCP (85.02–92.25%) and PAPO (5), suggesting that they fall within the values for genera previously observed in Saccharomycetaceae (Liu et al. 2024a). These results support a close relationship between C. ficus and the genus Millerago and further highlight the superior resolution of phylogenomic trees compared to LSU rDNA-based trees. Therefore, we suggest to merge C. ficus to the genus Millerago.
Candida montana and B. siamensis formed a well-supported clade in our phylogenomic analysis (Fig. 15) that was distantly positioned to the Barnettozyma sensu stricto clade, suggesting that the two clades might represent distinct genera. This observation agrees with previous phylogenomic analyses (Shen et al. 2018; Opulente et al. 2024). Barnettozyma botsteinii was not included in earlier studies. This species occupied a basal position to the clade formed by C. montana and B. siamensis (Fig. 15). Similarly to B. siamensis, this species requires to be reclassified due to its distant position from the Barnettozyma sensu stricto clade. We explored the genetic heterogeneity in Barnettozyma siamensis clade+B. botsteinii and observed AAI values of 72.73–79.96% and POPC values of 87.80–91.36%, falling within the range of the generic values previously observed in Saccharomycetaceae by Liu et al. (2024a). Therefore, we suggest to recombine these species into a new genus, for which the name Gotozyma gen. nov. (see Taxonomy).
Candida norvegica, C. qinlingensis, B. pratensis, and B. wickerhamii were placed in the Barnettozyma wickerhamii clade (Fig. 16). This clade included B. pratensis, and B. wickerhamii, B. salicaria, C. qinlingensis, and C. norvegica and was previously resolved in a recent phylogenomic analysis by Opulente et al. (2024), where it received strong (100%) support. Before Kurtzman et al. (2008) reclassified yeast species with saturn-shaped spores based on multi-gene phylogeny and introduced the genus Barnettozyma, the genus Komagataea was described by Yamada et al. (1994b) based on an LSU and SSU rDNA phylogenetic analysis with Komagataea pratensis (basionym: Williopsis pratensis Babeva & Reshetova) as the type species. Although Kurtzman et al. (2008) considered the genus Komagataea congeneric to Barnettozyma, several earlier observations suggested that these clades may represent distinct genera. For example, Naumova et al. (2004) observed that species of Komagataea and the Williopsis sensu stricto complex have different karyotypes. We propose to reinstate the genus Komagataea to accommodate B. pratensis, B. salicaria, B. wickerhamii, C. norvegica and C. qinlingensis because of the distant relationships between the B. wickerhamii and Barnettozyma sensu stricto clades, as well as the topology of the phylogenomic tree that showed that the Barnettozyma sensu stricto clade branched first within the clade that included the Phaffomyces, Gotozyma, Millerago, and Komagataea clades. Our LSU rDNA-based analysis revealed another potential new species in Komagataea (Fig. S13), namely the sequence labelled as B. salicaria isolate 0H4 (KM103057) and B. salicaria isolate 0H7 (KM103057), from surface water in South Africa that differed from the type strain CBS 5456 of B. salicaria by 7–39 nt (1.3–6.5%) in the D1/D2 domain of LSU rDNA.
Candida species and related genus Cyberlindnera (Phaffomycetaceae, Phaffomycetales, Saccharomycetes)
In the reclassification of the genus Pichia, Kurtzman et al. (2008) introduced the genus Lindnera for a monophyletic lineage typified by Lindnera americana (basionym: Hansenula bimundalis var. americana). However, since the name Lindnera was a homonym of a plant genus, Minter (2009) renamed and reclassified it into Cyberlindnera. In our phylogenomic analysis, Cyberlindnera has been resolved as a monophyletic lineage consisting of two large clades: Cyberlindnera sensu stricto and Williopsis (Fig. 17), this topology agreeing with results from earlier phylogenomic studies by Shen et al. (2018) and Opulente et al. (2024). We examined the genetic heterogeneity within this lineage using the genomic metrics. The entire Cyberlindnera lineage was characterized by slightly higher heterogeneity (AAI: 61.78–93.06%; POCP: 69.12–96.40%; PAPO: 0) compared to that of genera in the Saccharomycetaceae (Liu et al. 2024a). The two clades showed values in the following ranges: Cyberlindnera sensu stricto (AAI: 67.15–90.85%, POCP: 76.63–94.72%, PAPO: 2) and Williopsis clade (AAI: 63.63–99.97%, POCP: 78.24–99.67%, PAPO: 4) (Table 1). The lower RED score (0.867) calculated for the genus Cyberlindnera by Li et al. (2021) suggests that this lineage is likely under-classified and may correspond to the family- or order-level taxon.
Candida takata and Candida vartiovaarae occurred in the Williopsis clade (Fig. 17) that contains Williopsis saturnus (presently Cyberlindnera saturnus), the type species of the genus Williopsis and some other Cyberlindnera species. Like the genus Hansenula described below, the genus name Williopsis will be reintroduced instead of creating a new genus to accommodate the Williopsis clade. As a result, C. takata and C. vartiovaarae will be transferred into the genus Williopsis.
Candida adriatica, Candida easanensis, Candida hungchunana, Candida maesa, Candida pattaniensis, Candida stauntonica and Candida taoyuanica belong to the Cyberlindnera sensu stricto clade, which also contains the type species of Cyberlindnera, Cyberlindnera americana (Fig. 17). As discussed below, most of the sexual species in the clade were previously classified in the genera Hansenula and Pichia. The Cyberlindnera sensu stricto clade also contains Cyberlindnera rhodanensis, the type species of the genus Petasospora (Boidin & Abadie 1954). Although Kurtman et al. (2008) has considered Saccharomyces rhodanensis Ramírez & Boidin (1953), the basionym of Cyberlindnera rhodanensis, in their reclassification, neither these authors nor Minter (2009) acknowledged the name Petasospora as a potentially competing name for the clade. However, Petasospora is a validly published name and predates Cyberlindnera, thereby holding taxonomic priority. Therefore, those seven Candida species, along with 17 Cyberlindnera species, will be transferred to the reinstated and emended genus Petasospora as new combinations.
In our phylogenomic analysis, Candida freyschussii was placed on a long branch closely related to Cyberlindnera (Figs. 15 and 17), which is in agreement with the observations by Opulente et al. (2024). Additionally, in the LSU rDNA-based sequence analysis, this species clustered with the sequence labelled as Candida sp. NIAH-01 (GenBank AB703242) from a clinical bovine mastitic milk sample in Japan (Fig. S13). Candida sp. NIAH-01 differs from C. freyschussii by 38 nucleotides (6%) in the D1/D2 LSU sequence dataset, which indicates that this strain likely represents another species closely related to C. freyschussii. Considering its distant relationship to Cyberlindnera and the clade containing the type of the genus Candida, Buckleya gen. nov. is proposed to accommodate C. freyschussii and Candida sp. NIAH-01.
Candida species and related genus Starmera (Phaffomycetaceae, Phaffomycetales, Saccharomycetes)
The genus Starmera received good support in the phylogenomics analysis that revealed two clades in the genus, namely the Starmera sensu stricto clade and the Starmera dryadoides clade (Fig. 17). The same topology and composition were observed in a recent phylogenomic analysis by Opulente et al. (2024). The genus Starmera was proposed by Yamada et al. (1998) based on LSU and SSU rDNA sequence analysis to accommodate Pichia amethionina and its variety Pichia amethionina var. pachycereana. Pichia amethionina exhibited several unique morphological and physiological characteristics, which correlated with its phylogenetic divergence from other Pichia species. Somewhat later, Pichia caribaea was transferred to Starmera (Yamada et al. 1999). The genus was further expanded by Kurtzman et al. (2008) to include Pichia dryadoides and Pichia quercuum, based on their multi-gene phylogenetic analysis of the genus Pichia. Two species, Candida berthetii and Candida dendrica, clustered with S. dryadoides and Starmera quercuum in the Starmera dryadoides clade (Fig. 17).
Our genome-based analyses showed that the genus Starmera is genetically heterogeneous, as shown by the genomic metrics AAI (61.57–92.68%), POCP (67.37–97.98%), and PAPO (4) (Table 1). Particularly, the POCP values of Starmera are lower than those observed in genera of the Saccharomycetaceae (Liu et al. 2024a) and some members of the Metschnikowiaceae (Liu et al. 2024b). Additionally, the lower RED score (0.823) for Starmera calculated by Li et al. (2021) suggests that this lineage is likely under-classified and may correspond to a family- or order-level taxon. Genomic metrics calculated for the two Starmera clades (Table 1) indicate a higher consistency of their respective ranges: Starmera sensu stricto clade (AAI: 67.88–92.68% and POCP: 81.76–97.98%), and Starmera dryadoides clade (AAI: 65.26–83.33% and POCP: 81.67–89.88%).
Kurtzman (2011c) showed that S. dryadoides and S. quercuum do not require an exogenous source of L-methionine or L-cysteine, whereas the species of Starmera sensu stricto clade depend on these two amino acids. This metabolic distinction suggests that S. dryadoides and S. quercuum may represent a new sister genus to Starmera. Thus, we propose the establishment of a new genus, Liangdongia gen. nov., for the Starmera dryadoides clade. In agreement with their phylogenetic position in this clade, C. berthetii and C. dendrica will be transferred to Liangdongia. Additionally, Candida laemsonensis was placed in the Starmera dryadoides clade in our ITS+LSU rDNA-based analysis (Fig. 16), indicating that this species should also be reassigned to Liangdongia.
Candida species and related genus Wickerhamomyces (Wickerhamomycetaceae, Phaffomycetales, Saccharomycetes)
Our study, along with the previous phylogenomic analysis by Opulente et al. (2024), resolved the genus Wickerhamomyces is polyphyletic. Similar to Barnettozyma, the genus Wickerhamomyces comprises multiple clades that do not form a monophyletic group, with some clades occurring interspersed among other genera (Figs. 15–18, Table S2). Two major clades, i.e., the Wickerhamomyces sensu stricto and the Wickerhamomyces bovis clades, have been resolved in the phylogenomic analysis. Wickerhamomyces silvicola occupied a basal position to the two clades, as well as to the Phaffomyces-Millerago-Barnettozyma lineage (Fig. 15). Other clades containing Wickerhamomyces species were placed closer to the Cyberlindnera, Starmera, and Williopsis clade. Consequently, the genus Wickerhamomyces showed genetic heterogeneity in the analysis of genomic metrics (AAI: 56.06–100.00%; POCP: 44.43–99.82%; PAPO: 0). The genus Wickerhamomyces contained six clades and three single-species lineages that might represent genera based on the phylogenomic analysis and the genome-based indexes (Figs. 15 and 17, Table 1). Candida jianshihensis, Candida quercuum and Candida ulmi were located in the Wickerhamomyces sensu stricto clade (Figs. 15–16), and will therefore be transferred to Wickerhamomyces as new combinations in the Taxonomy section. The single-species lineage Wickerhamomyces silvicola is placed as pro tempore at present, although it occupied a basal position to the two clades, as well as to the Phaffomyces-Millerago-Barnettozyma lineage (Fig. 15).
Candida dajiaensis, Candida odintsovae, Candida peoriensis and Candida yuanshanica belonged to the Wickerhamomyces bovis clade (Fig. 16), which is phylogenetically located near the lineage of Barnettozyma-Millerago-Phaffomyces. The clade received good support in both the present phylogenetic analysis and the study by Opulente et al (2024). The clade is characterized by moderate genomic heterogeneity, as indicated by its genomic metrics (AAI: 65.53–98.19%, POCP 82.92–96.62% and PAPO: 2), which are comparable with those observed within genera of Saccharomycetaceae (Liu et al. 2024a). Species in the clade have been previously described in the genera Candida and Pichia. The oldest species, Wickerhamomyces (Pichia) bovis, was also assigned to the genus Zymopichia by Novák & Zsolt (1961). However, since the authors did not designate a type species for the genus, this name is nomenclaturally invalid and unavailable for reinstatement. A new generic name, Taiozyma gen.nov., will be proposed for members of this clade in the Taxonomy section.
Candida solani is located in the Wickerhamomyces pijperi clade, which is close to the single-species lineage Wickerhamomyces mucosus (Fig. 17). The clade has also been resolved in the study by Opulente et al. (2024) and was phylogenetically placed closely to the Cyberlindnera-Williopsis lineage. The genetic heterogeneity examined within the clade (AAI 71.66–81.75%, POCP 63.80–92.13%, PAPO: 87) is comparable to that observed in the accepted genera of Saccharomycetaceae (Liu et al. 2024a). Although the POCP values appear to be somewhat lower, the number of 87 shared orthologs was high (Table 1). The species Wickerhamomyces mucosus was originally described as Pichia mucosa by Wickerham & Kurtzman (1971). Later, Muller & Kock (1986) reassigned this species to the newly established genus Waltiozyma. However, Kurtzman et al. (2008) transferred this species once again, moving it from Waltiozyma to Wickerhamomyces based on a multigene analysis. Based on our analyses, we propose to reinstate the genus Waltiozyma for the Wickerhamomyces mucosus+ Wickerhamomyces pijperi lineage. Another clade, closely positioned to the Cyberlindnera-Williopsis lineage and comprising Wickerhamomyces hampshirensis and Wickerhamomyces strasburgensis, will be reclassified due to its distant relationship with the Wickerhamomyces sensu stricto clade. While Wickerhamomyces hampshirensis has been originally described in the genus Pichia, Wickerhamomyces strasburgensis has been previously classified in the genera Saccharomyces (basionym: Saccharomyces strasburgensis), Petasospora and Zymopichia. Because the generic name Zymopichia is invalid, and Saccharomyces and Petasospora phylogenetically belong to different phylogenetic lineages, we propose to accommodate these two species in a new genus Xingzhongia gen. nov. (see Taxonomy section below).
Candida namnaoensis and Candida ponderosae are positioned within the Wickerhamomyces chambardii clade, which is phylogenetically related to the genus Starmera (Fig. 17). This observation is in agreement with the analysis by Opulente et al. (2024). Because this clade is distant from the type lineage of Wickerhamomyces, we propose to accommodate these yeasts in a new genus, Ruyongia gen. nov. Another species, Wickerhamomyces kurtzmanii, occupied a basal position in Wickerhamomycetaceae, which seems to represent a new genus. But we place it as Wickerhamomyces pro tempore at present, which will be resolved in the future.
Candida silvicultrix, Wickerhamomyces anomalus (syn. Hansenula anomala, the type species of the genus Hansenula) and four other Wickerhamomyces species occurred in the Hansenula clade (Fig. 17). This clade is positioned closer to the Cyberlindnera-Williopsis lineage and the genus Starmera, as indicated in the phylogenomic tree (Fig. 17) and the recent study by Opulente et al. (2024). The genus Hansenula was distinguished from Pichia based on phenotypic characteristics, particularly in the assimilation of nitrate – Hansenula species assimilating nitrate, whereas Pichia species not utilizing this compound (Stelling-Dekker 1931; Kurtzman 1984a, b; Bhunjun et al. 2024). Kurtzman (1984c) argued that nitrate assimilation could not serve as a genus defining character, as DNA-DNA reassociation values among phenotypically similar species of Pichia and Hansenula species indicated a close taxonomic relationship. Consequently, Hansenula was synonymized under Pichia, leading to an expanded concept of the genus Pichia that included 91 species (Kurtzman 1998). Subsequent phylogenetic analyses based on rDNA and multigene sequences revealed that this broadened concept of Pichia was highly polyphyletic (Billon-Grand 1989; Liu & Kurtzman 1991; Yamada et al. 1994a, b, 1995a, b, 1998; Kurtzman & Robnett 1998; Kurtzman 2006; Kurtzman et al. 2008). As a result, several genera were split from Pichia (as adopted by Kurtzman 1998), including Barnettozyma, Cyberlindnera (originally Lindnera), Kregervanrija, Kodamaea, Komagataella, Kuraishia, Nakazawaea, Phaffomyces, Ogataea, Saturnispora, Starmera, Yamadazyma, and Wickerhamomyces (Bhunjun et al. 2024). Pichia anomala (formerly classified as Hansenula anomala) was reassigned to the genus Wickerhamomyces based on a multigene sequence analysis (Kurtzman et al. 2008).
Our phylogenomic analysis showed that H. anomala (currently named Wickerhamomyces anomalus) and Pichia myanmarensis, C. silvicultrix, and 10 other Wickerhamomyces species formed a well-supported Hansenula clade. This clade exhibited moderate genetic heterogeneity (AAI: 68.07–84.27%; POCP: 80.37–94.72%; PAPO: 11), with genomic metrics in the range of those reported for genera in Saccharomycetaceae (Liu et al. 2024a). This clade appeared distinct from the other Wickerhamomyces species, as well as from Barnettozyma, Cyberlindnera and Phaffomyces (Figs. 17 –18), suggesting that the Hansenula clade does not belong to the genus Wickerhamomyces.
Kurtzman et al. (2008) and Kurtzman (2011c) argued against the reintroduction of Hansenula and discussed the reclassification of Pichia anomala. While considering the relatedness of P. anomala to C. silvicultrix, Pichia ciferrii, Pichia subpelliculosa, Pichia sydowiorum and Pichia lynferdii (i.e., the Hansenula clade in Fig. 17), Kurtzman (2011c) suggested that these species belonged to the large Wickerhamomyces clade, as indicated by the multi-gene phylogenetic analysis performed by Kurtzman et al. (2008). The taxonomic history of Saccharomyces anomalus (the original name of P. anomala) is complex, having undergone transitions through Willia (homonym of a moss) and Hansenula. The situation was further complicated by an older heterotypic synonym, Saccharomyces sphaericus, later synonymized with H. anomala, though no type material was preserved. The lack of type material for reliable authentication rendered these names impractical, even if not always formally invalid. As Kurtzman (2011c) indicated, the oldest valid name associated with available type material for a heterotypic synonym of H. anomala was Endoblastoderma pulverulentum. Although a neotype was proposed for H. anomala, two different strains were independently selected around the same time by Lodder & Kreger-van Rij and Wickerham, respectively. However, no formal proposal was made to conserve Hansenula using a neotype of either S. sphaericus or S. anomalus, or by adopting E. pulverulentum as a conserved name (Kurtzman 2011c). Consequently, Daniel et al. (2012) proposed conserving the name Wickerhamomyces against Hansenula and rejecting the questionable name Saccharomyces sphaericus Sacc. 1877. They designated strain NRRL Y-366 as an epitype to the newly designated lectotype of S. anomalus. Furthermore, these authors indicated that the concerns expressed by Kurtzman (2011c) about the lack of type material did not render the names S. sphaericus and S. anomalus invalid. As a result, the name Hansenula was validly published, thus holding priority over Wickerhamomyces (Daniel et al. 2012). Additionally, the latter authors suggested that Endoblastoderma was a synonym of Saccharomyces, rather than Hansenula.
The conservation of Wickerhamomyces over Hansenula was preferred because Hansenula was highly polyphyletic, whereas Wickerhamomyces was adopted for a monophyletic group at that time (Daniel et al. 2012). If future taxonomic revisions will indicate that the types of Wickerhamomyces and Hansenula belong to different genera, both names are available from a nomenclatural point of view. Malimas et al. (2023) reintroduced Hansenula and assigned nine species to this genus based on pair-wise sequence similarities of the LSU rDNA. However, C. silvicultrix, Wickerhamomyces queroliae, Wickerhamomyces spegazzinii and Wickerhamomyces sylviae, belonging to the Hansenula clade (Figs. 17–18), were not assigned to Hansenula by Malimas et al. (2023). Therefore, those four species will be transferred to Hansenula in the Taxonomy section below.
The impact of reductive evolution, HTGs, hybridization (or alloaneuploidy) and introgressions on the values of genomic metrics
Genomic metrics, such as AAI and POCP, have been used as valuable approaches to delineate genera for bacteria (Qin et al. 2014; Kuzmanović et al. 2022; Montecillo 2023). Recently, those two approaches were recommended to delimit yeast genera (Liu et al. 2024a, b). However, some evolutionary factors, such as reductive evolution, hybridization, horizontal transfer genes (HTGs), introgressions, and alloaneuploidy, may affect the values of AAI and POCP. To address those issues, we used the Dipodascus/Galactomyces/Geotrichum lineage, Saccharomyces and Starmerella as examples to evaluate those impacts.
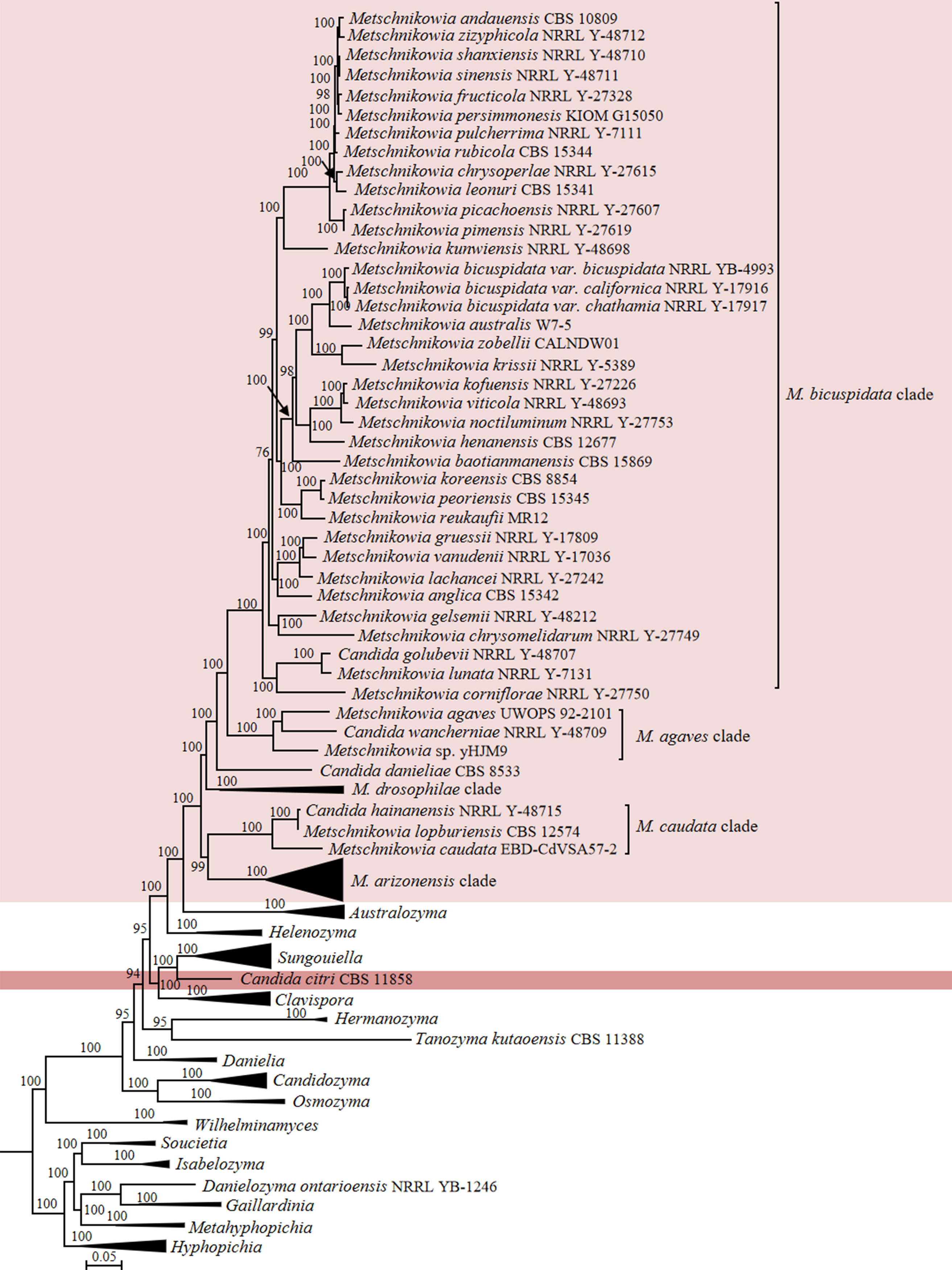
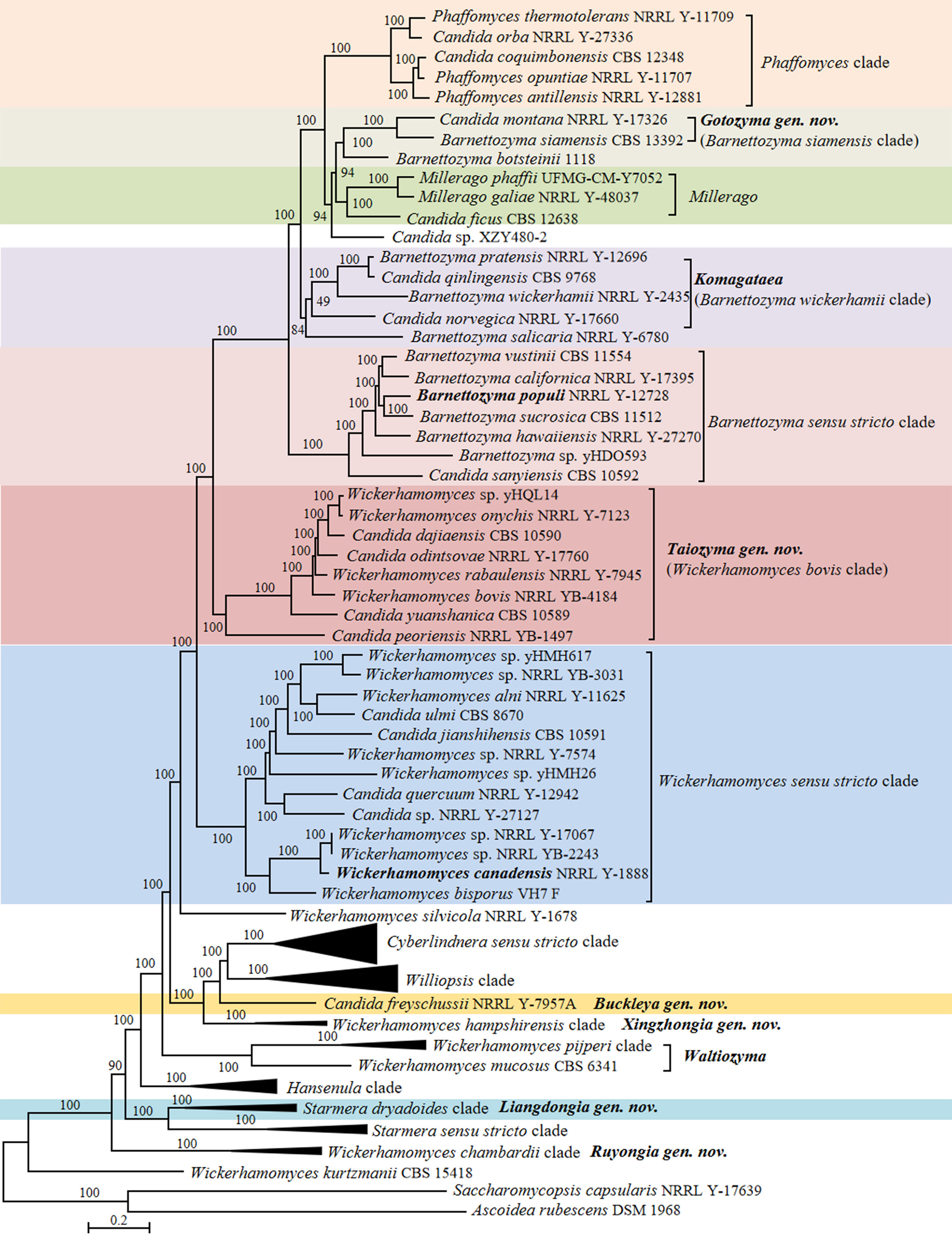
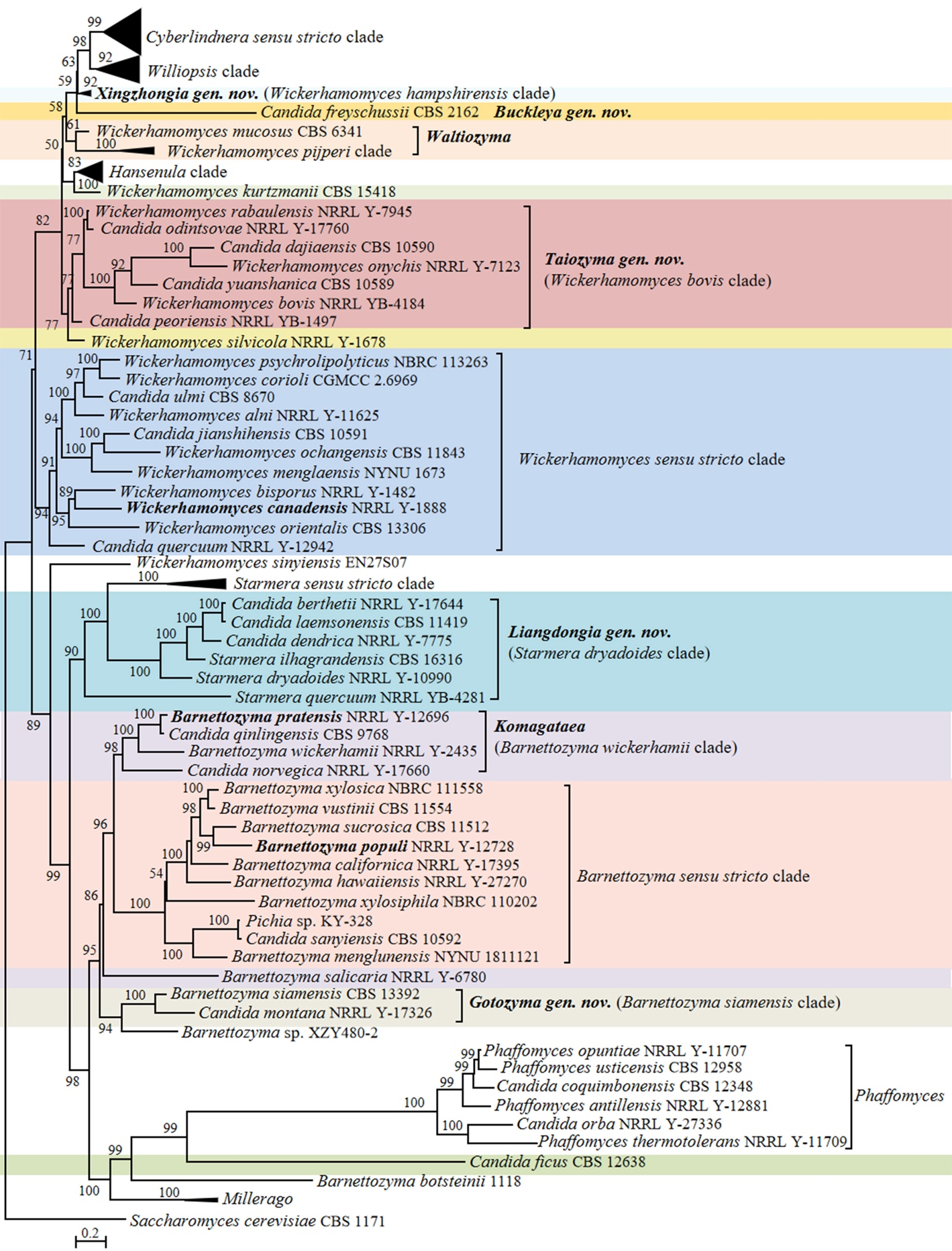
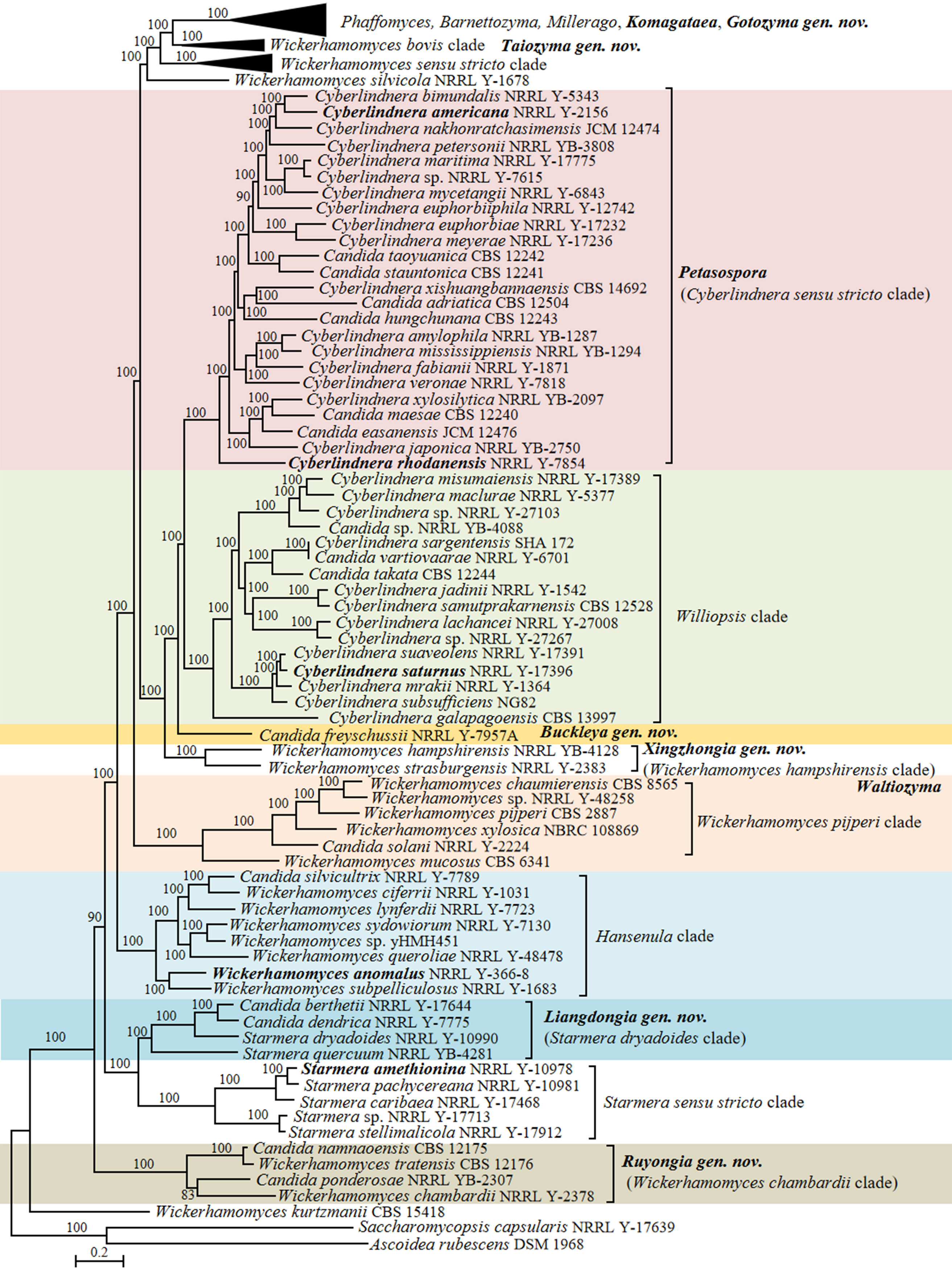
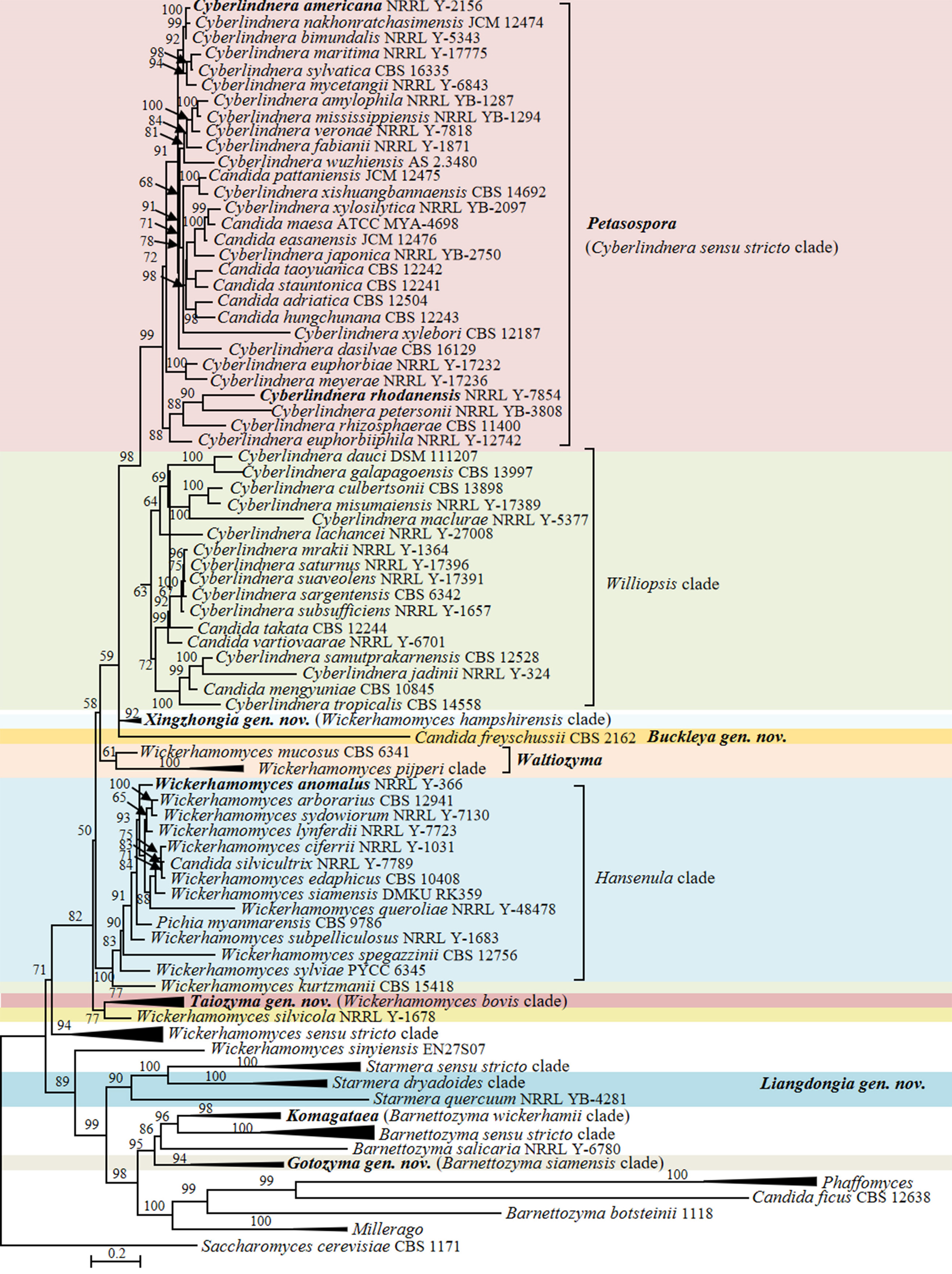
Most species of Dipodascus/Galactomyces/Geotrichum lineage seem to be hybrid (or alloaneuploidy) due to having a higher rate of duplicated BUSCOs (Table S1). Therefore, this lineage was selected to evaluate the impact of hybridization (or alloaneuploidy) on AAI and POCP values. We calculated pairwise AAI and POCP values using the three datasets described in the method section and conducted linear regression analysis. Our results demonstrated that the AAI and POCP values of All_genome_dataset (with paralog-OGs, containing hybrid) and that of Subgenome_1_dataset and Subgenome_2_dataset (without paralog-OGs, omitting hybrid) are remarkably consistent (Table 2). Furthermore, extremely high consistency was observed in the AAI values of the same pairwise species with and without paralog-OGs. When evaluating the impact of hybridization (or alloaneuploidy) on AAI analysis using Subgenome_1_dataset and Subgenome_2_dataset (Fig. 19), the slopes of the linear regression equations were both close to 1 (1.08 or 1), indicating that hybridization (or alloaneuploidy) had no significant impact on the overall level of AAI values within the lineage. And high R² values (0.98 or 1) further confirmed that the linear relationship between the two sets of AAI values was stable with no systematic bias caused by the removal of paralog-OGs. When evaluating the impact of hybridization (or alloaneuploidy) on the POCP analysis (Fig. 20), the POCP values of the same pairwise species combinations also showed high consistency across different datasets. Although local variations in hybridization levels or gene duplication levels resulting from alloaneuploidy among different species led to slight deviations of some points in the scatter plots, the regression equations based on the two subgenome datasets still maintained high slopes (1.03 or 0.912), and the R² values (0.92 or 0.97) were also at a high level. The above analyses indicate that the effect of hybridization (or allopolyploidy) seems not to cause a major impact on AAI and POCP values.
| Datasets | AAI values (%) | POCP values (%) |
| All_genome_dataset | 61.01–91.09 | 60.12–94.63 |
| Subgenome_1_dataset | 59.58–90.80 | 57.10–92.60 |
| Subgenome_2_dataset | 60.53–89.71 | 62.91–93.10 |
Note: All_genome_dataset includes all proteins (genes) for each species of Dipodascus/Galactomyces/Geotrichum lineage; Subgenome_1_dataset contains orthologous groups (OGs) without paralogous genes; Subgenome_2_dataset is composed of OGs without paralogous genes and one copy of paralogous genes that was randomly selected from the paralog-containing OGs.
| Taxa | 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 |
| Starmerella | 56.87–89.06% | 56.92–89.11% | 57.76–90.67% | 58.36–91.08% | 49.07–93.24% | 49.47–93.24% | 60.46–97.89% | 62.05–97.90% |
| Starmerella clade 1 | 72.77–89.06% | 72.85–89.11% | 74.55–90.67% | 75.21–91.08% | 80.33–93.24% | 80.38–93.24% | 87.08–97.89% | 88.59–97.90% |
| Starmerella clade 2 | 56.87–77.34% | 56.92–77.40% | 57.76–79.36% | 58.36–80.05% | 49.07–91.12% | 49.47–90.89% | 60.46–96.58% | 62.05–96.98% |
| Starmerella apicola clade | 63.11–70.44% | 63.17–70.49% | 64.45–71.72% | 65.13–72.36% | 77.02–85.86% | 77.11–85.83% | 87.44–93.86% | 88.48–94.63% |
| Starmerella cellae clade | 62.67–74.03% | 62.67–74.11% | 63.83–75.26% | 64.48–75.76% | 60.03–85.26% | 59.80–85.48% | 73.52–91.79% | 75.25–92.51% |
| Starmerella sensu stricto clade | 66.62–77.34% | 66.70–77.40% | 68.41–79.36% | 69.19–80.05% | 78.62–91.12% | 78.63–90.89% | 87.41–96.58% | 88.21–96.98% |
| Starmerella stellata clade | 58.45–69.19% | 58.50–69.24% | 59.45–70.62% | 59.97–71.18% | 60.00–78.43% | 60.52–78.57% | 69.78–87.14% | 71.41–87.99% |
Note: 1, The AAI values calculated based on all proteins (genes) for each species of the clade (genus); 2, The AAI values of the clade (genus) calculated after removal of HTGs; 3, The AAI values of the clade (genus) calculated based on Starmerella_90_dataset that contains orthologous groups (OGs) occurring in ≥ 90% species of Starmerella; 4, The AAI values of the clade (genus) calculated based on Starmerella_95_dataset that contains orthologous groups (OGs) occurring in ≥ 95% species of Starmerella; 5, The POCP values calculated based on all proteins (genes) for each species of the clade (genus); 6, The POCP values of the clade (genus) calculated after removal of HTGs; 7, The POCP values of the clade (genus) calculated based on Starmerella_90_dataset that contains orthologous groups (OGs) occurring in ≥ 90% species of Starmerella; 8, The POCP values of the clade (genus) calculated based on Starmerella_95_dataset that contains orthologous groups (OGs) occurring in ≥ 95% species of Starmerella.
Starmerella species acquired many HTGs from bacteria or other fungi (Gonçalves et al. 2020, 2022; Pontes et al. 2024). To assess the bias introduced by HTGs on AAI and POCP analyses, we omitted the HTGs from the genome datasets of Starmerella, then we calculated the AAI and POCP values and compared them with the results obtained from the whole genome dataset. After removal of the HTGs, the AAI values of Starmerella (i.e., 56.92–89.11%) exhibited minor changes
compared to that calculated from the all-genome datasets with HTGs (i.e., 56.87–89.06%). The POCP values of Starmerella calculated after removal of HTGs (i.e., 49.47–93.24%) had a slight difference from that calculated based on whole genome datasets (i.e., 49.07–93.24%) (Table 3). Linear regression analysis further confirmed that extremely high consistency was observed in AAI and POCP values of the same pairwise species within this genus before and after removal of HTGs, respectively (Fig. 21). The slopes of the regression equations based on AAI and POCP values were 1 and 0.987, respectively, indicating that the presence or absence of HTGs had no significant impact on the overall levels of these two indicators. High coefficients of determination (R² values: 0.98 for AAI; 1 for POCP) further confirmed that the linear relationships of the two sets of AAI and POCP values were stable, and the same pairwise species showed high consistency for the same indicator. Based on these results, it can be inferred that the presence or removal of HTGs had no significant interference with the AAI and POCP analyses.
To evaluate the impact of the reductive evolution on AAI and POCP calculations, two subgenome datasets were created in the method section. Our results demonstrated that the AAI values calculated based on Starmerella_90_dataset and Starmerella_95_dataset are similar to that calculated based on the whole genome dataset. However, the POCP values calculated based on those two subgenome datasets increased compared to the whole genome dataset without the removal of reductive OGs (Table 3). Linear regression analysis further showed that when evaluating the impact of reductive evolution on AAI analysis (Fig. 22), the slopes of the regression equations and coefficients of determination (R²) based on the two subgenome datasets were high, indicating that the AAI values calculated based on different subgenome datasets are similar to that calculated based on the whole genome dataset. So reductive evolution may have no significant effect on AAI analysis. In contrast, when evaluating its impact on POCP analysis (Fig. 23), the slopes (0.78 or 0.736) of the linear regression equations deviated significantly from 1, but the R² values (0.96) were high, suggesting that the POCP values calculated based on different subgenome datasets had evident differences from that calculated based on the whole genome dataset. Based on the linear regression equations, it can be inferred that reductive evolution may cause an evident increase in POCP results. Thus, it can be concluded that reductive evolution may have a significant impact on POCP analysis (leading to elevated results) but may have no effect on the outcome of the AAI analysis. It is noted that the POCP values of Starmerella calculated based on Starmerella_90_dataset and Starmerella_95_dataset (Table 3) are still much lower than the generic values recommended by Liu et al. (2024a), which indicates that the genus Starmerella is still genetically heterogeneous even when omitting the impact of reductive evolution. The higher POCP values of Starmerella_90_dataset and Starmerella_95_dataset may be caused by the reductive denominator during the POCP analysis.
The genus Saccharomyces includes at least two hybrid species, namely S. bayanus and S. pastorianus, and some hybrid strains, e.g., S. cerevisiae × S. kudriavzevii. D'Angiolo et al. (2020) demonstrated that the S. cerevisiae Alpechin lineage, e.g., strain DBVPG6765, carries abundant S. paradoxus introgressions. We used Saccharomyces with three datasets described in the method as an example to address introgressions and hybridization (or alloaneuploidy) impacts on the reliability of AAI and POCP analyses. The obtained AAI values of Saccharomyces_9_dataset, Saccharomyces_10_dataset and Saccharomyces_12_dataset were 82.30–97.09%, 82.30–99.39% and 80.66–98.13%, respectively (Table 4). The POCP values of the Saccharomyces_9_dataset, Saccharomyces_10_dataset and Saccharomyces_12_dataset were 95.95–98.42%, 95.95–98.97% and 94.38–98.42%, respectively (Table 4). The above analysis results indicate that the hybridization (or alloaneuploidy) does not have an impact on the AAI and POCP values, which is in agreement with the results from the analysis of Dipodascus/Galactomyces/Geotrichum lineage described above. The results from the Saccharomyces analysis also indicate that introgression does not cause any bias for the reliability of AAI and POCO analyses (Table 4), but this issue should be studied further using a more robust data analysis.
Genomic insights into multidrug resistance in revised Candida species assigned to the new genera
Fifty-five Candida species were assigned to 22 newly created genera in this study. Only three Candida species, namely, C. palmioleophila, C. sake and C. zeylanoides, are clinical-related yeasts (https://www.atlasclinicalfungi.org/), while the other Candida species do not seem to be identified in clinical settings based on the data analysis collected from the latest edition of the TYTS, The Yeasts Trust Database (https://theyeasts.org/), the medical database, Atlas of Clinical Fungi (https://www.atlasclinicalfungi.org/), and related references searched from PubMed (https://pubmed.ncbi.nlm.nih.gov/). To evaluate the antifungal resistance profiles of species assigned to the newly created genera in this study based on the genome analysis, we retrieved known antifungal resistance-associated genes of C. albicans from NCBI, including: 1) the ERG11 gene (with mutation sites F126L, Y132F, K143R, F145L, G448E, F449V, G450E, and G464S) associated with azole resistance; 2) the ERG2 gene (with mutation site F105SfsX23) associated with polyene resistance; 3) the FKS1 gene (with mutation sites S645P/Y/F and F641Y) associated with echinocandin resistance; 4) the FUR1 gene (with mutation site F211I) associated with nucleoside analog resistance. Using these genes as reference sequences, we employed the BLASTP software to extract the corresponding genes from the protein sequences of the target species. Subsequently, multiple sequence alignment was performed on the obtained homologous genes to identify differences and similarities in antifungal resistance-associated genes between resistant and susceptible strains. The results (Table S4) showed that no mutations were detected at the sites of the ERG2, FUR1 and ERG11 genes across all revised Candida species assigned to the newly created genera in this study. In the analysis of echinocandin resistance, three Candida species that have been reclassified into novel genera in this study, namely C. sake NRRL Y-1622 (S645A), C. multigemmis NRRL Y-17659 (S645A) and C. tibetensis CBS 10298 (S645A), have a mutation in the S645 locus (Table S4). Only one type of amino acid substitution (A) was observed, but no mutations were detected in the F641 locus. This suggests that there may be a certain locus preference in the echinocandin resistance-associated mutations of these reclassified species. In contrast, the remaining Candida species that have been reclassified into novel genera exhibited no mutations at the antifungal resistance-associated gene mutation loci related to various antifungal drugs.
| Datasets | AAI values (%) | POCP values (%) |
| Saccharomyces_9_dataset | 82.30–97.09 | 95.95–98.42 |
| Saccharomyces_10_dataset | 82.30–99.39 | 95.95–98.97 |
| Saccharomyces_12_dataset | 80.66–98.13 | 94.38–98.42 |
Note: Saccharomyces_9_dataset includes only nine natural species; Saccharomyces_10_dataset is composed of nine natural species and S. cerevisiae DBVPG6765; Saccharomyces_12_dataset contains nine natural species and three hybrid species (or strains).
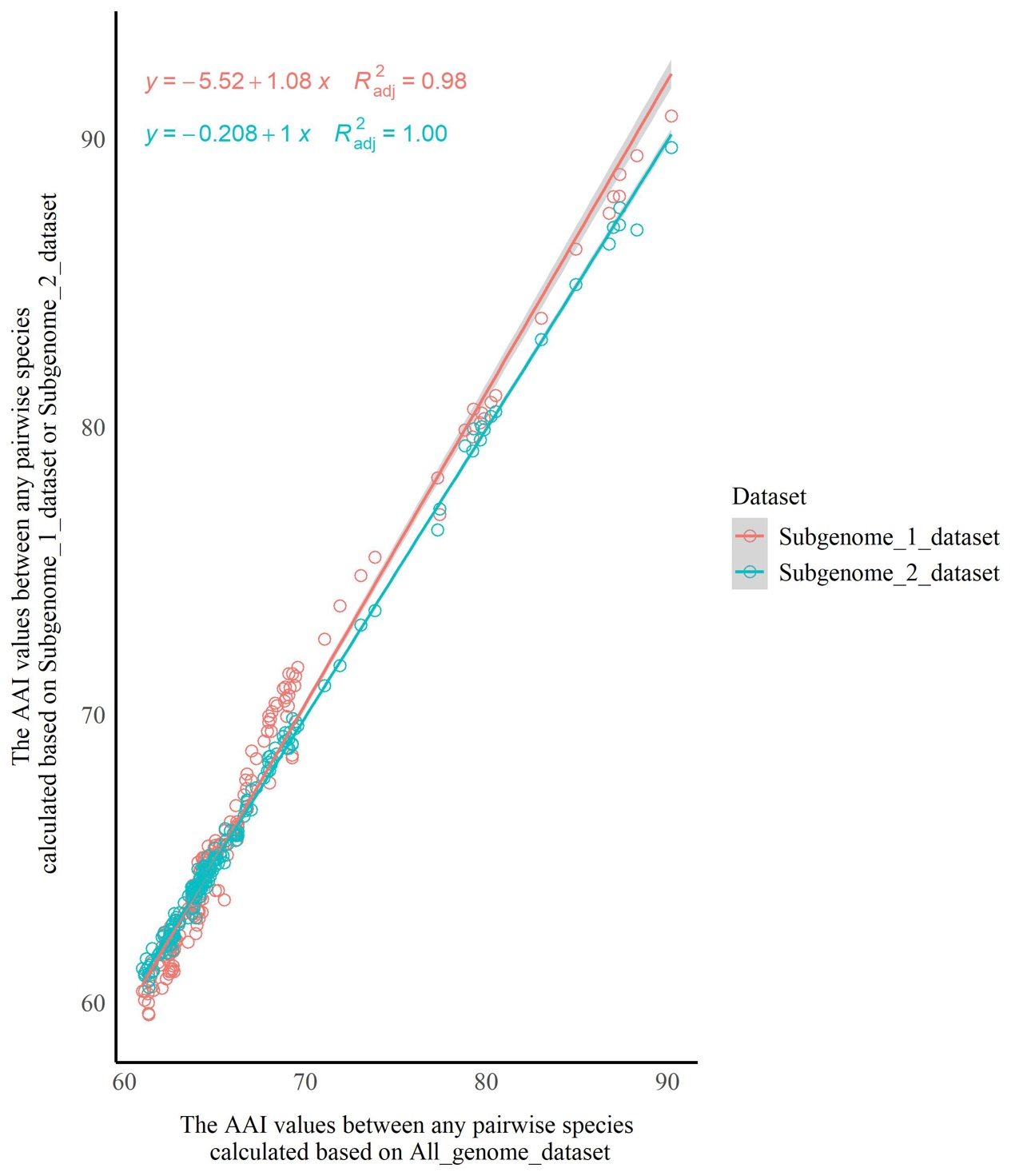
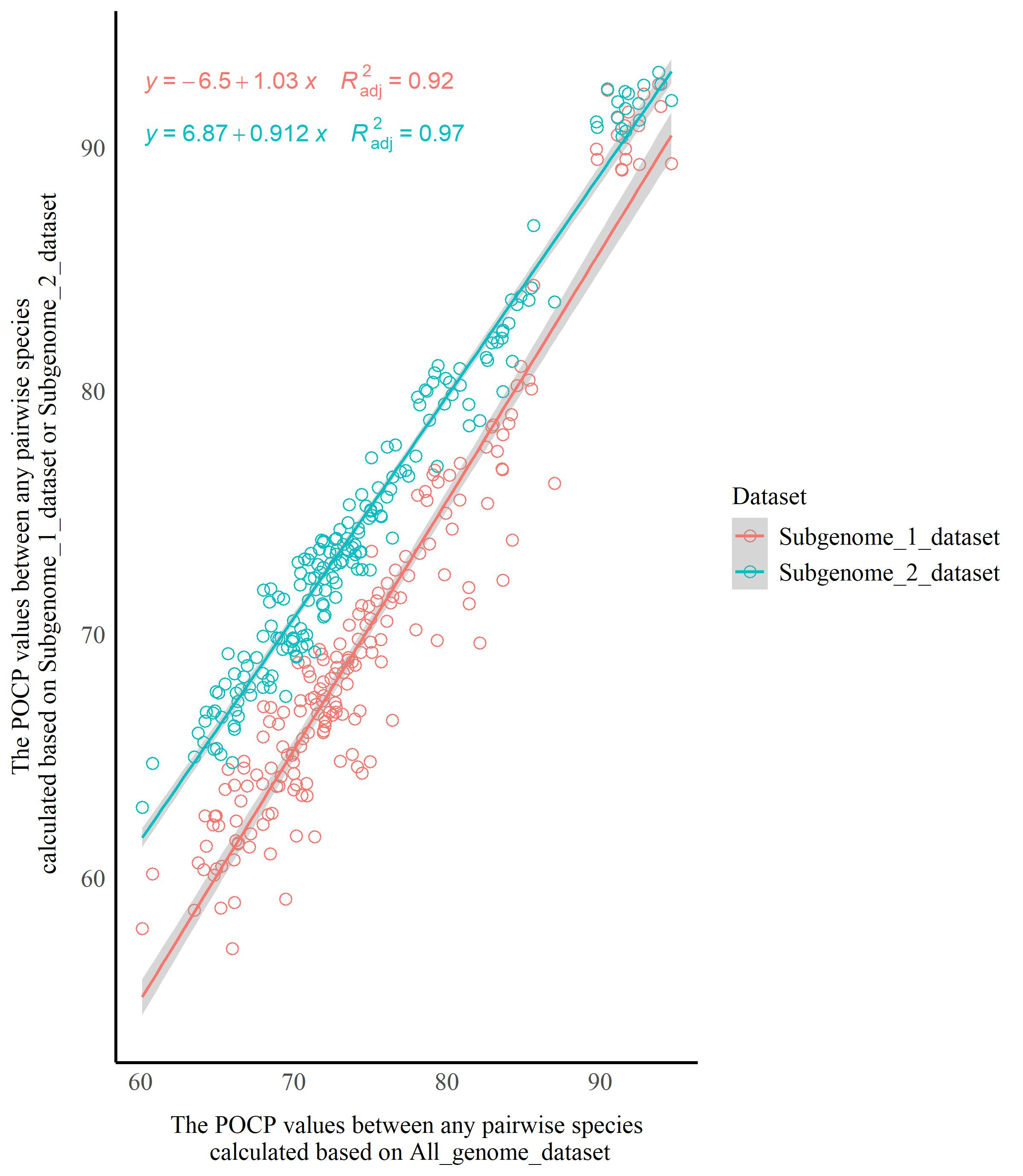
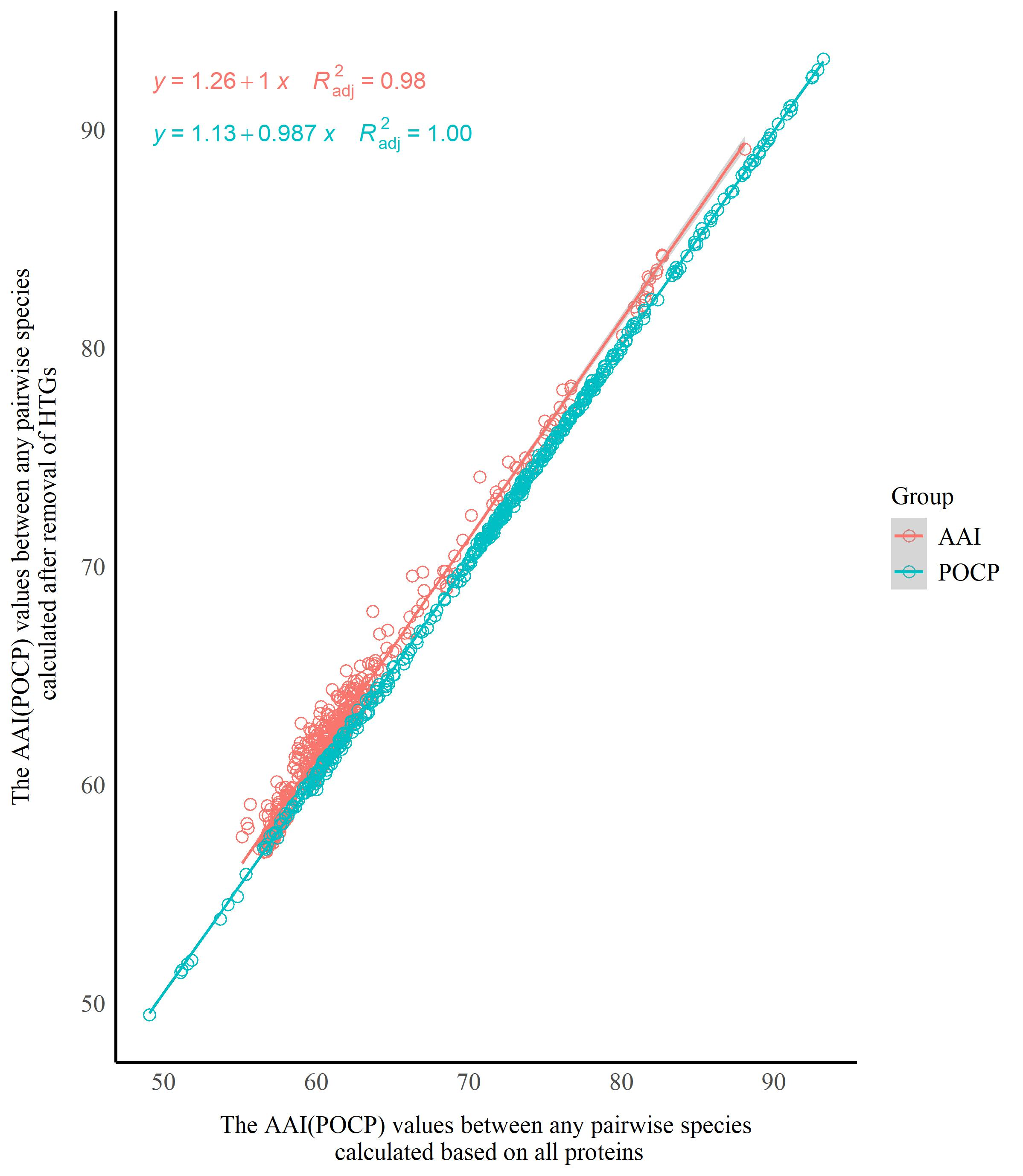
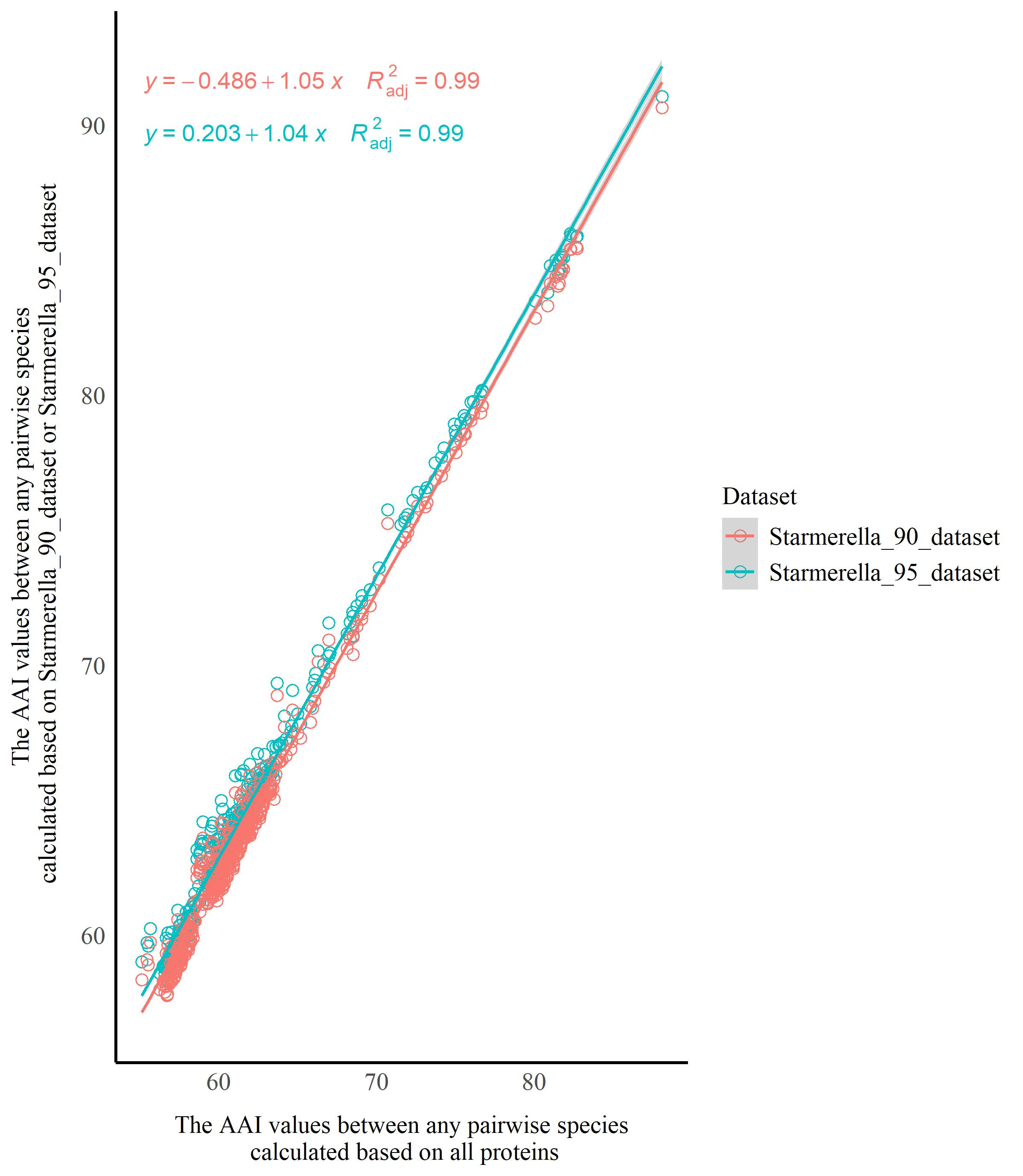
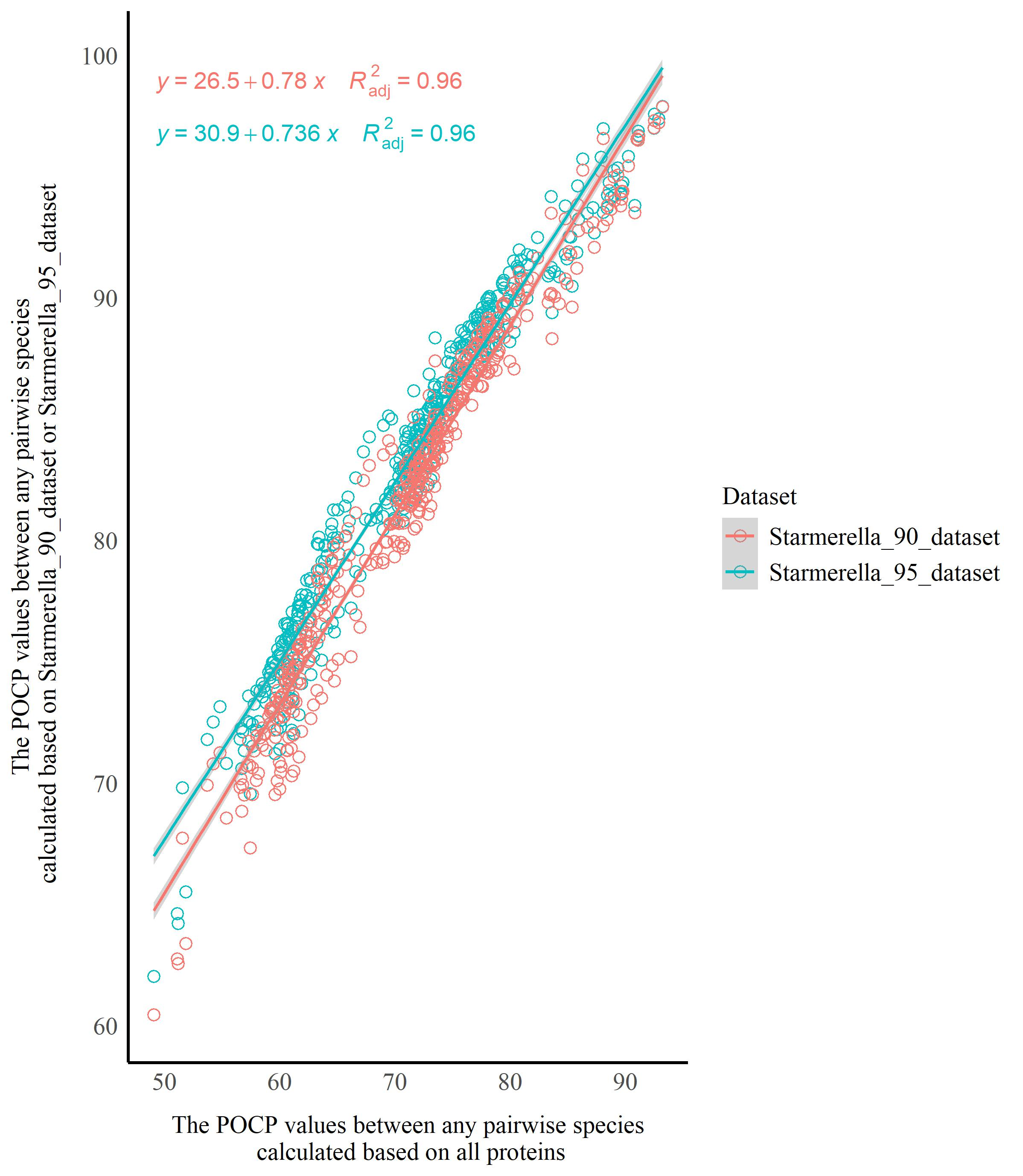
The above results indicate that: 1) these non-clinical Candida species may be generally susceptible to polyene drugs; 2) the three clinically relevant species, namely C. zeylanoides, C. palmioleophila and C. sake, have 8–16 μg/mL fluconazole MIC values (Pereira et al. 2010; Jensen & Arendrup 2011; Kucukates et al. 2016), which indicate a level of resistance distinct from typical C. albicans and may have new resistance mechanisms differing from C. albicans or other core Candida species. Stavrou et al. (2019) and Schmalreck et al. (2014) demonstrated that yeast phylogeny may be a guide to predict antifungal susceptibility profiles (ASPs), thereby improving treatment strategies, the impact of clinical diagnostics, and epidemiological tracking of yeast infections. Therefore, the different ASPs of C. zeylanoides, C. palmioleophila and C. sake and the core Candida species may be correlated with their different phylogenetic position.
Phenotypic features for yeast genus recognition
Yeast genera were traditionally recognized based on their phenotypic characteristics (Boekhout et al. 2021). However, molecular phylogenetic analyses have shown that most phenotypically defined genera are not monophyletic (Kurtzman 2011a; Boekhout et al. 2021). Can phenotypic features still be used to additionally characterize genera in both ascomycetous yeasts and basidiomycetous yeasts? In some cases, this is possible as, several genera can be recognized by their morphological or physiological characteristics. For example, the genus Eremothecium is characterized by its fusiform or acicular (needle-shaped) ascospores. The sister genera Zygosaccharomyces and Zygotorulaspora can be distinguished by physiology: all species of Zygotorulaspora can ferment raffinose, whereas all members of Zygosaccharomyces cannot. Other sister genera in the Saccharomycetaceae can be differentiated by phenotypic synapomorphies; some more distinctly than others (see Liu et al. 2024a). Most genera recognized through the phylogenomic analysis and the genomic metrics AAI, POCP and PAPO approaches in Liu et al. (2024b) can be recognized by phenotypic characters. For example, Australozyma vs Helenozyma vs Hermanozyma, Candidozyma vs Osmozyma, Danielozyma vs Gaillardinia vs Metahyphopichia, Isabelozyma vs Soucietia, and Clavispora vs Sungouiella exhibit notable phenotypic differences between them (for details see Liu et al. 2024b). In this study, we conducted a phenotypic comparison of sister genera identified, seeking morphological or physiological synapomorphies to support their generic circumscriptions. Among 25 newly described genera, 11 ones (44%) have distinct phenotypic features that differ from related genera (for details, see the Taxonomy section below). The new genera with phenotypic synapomorphies are Cariosilvazyma, Casaregolazyma, Chernovozyma, Daia, Grinbergsozyma, Intestinozyma, Liangdongia, Lizanozyma, Nothofagozyma, Westerdijkia and Xiuguozyma. Although the new genera Gotozyma and Xingzhongia can’t be distinguished from their related taxa by phenotypic characteristics, they have distinct GC% content from their relatives (Table S3).
Are the monotypic genera realistic?
About 14 Candida species are placed at basal positions of specific clades in our phylogenomic analysis and are thus separated from known genera. This is in agreement with earlier rDNA-based sequence phylogenetic analyses (Daniel et al. 2014; Takashima & Sugita 2022). The phylogenomic and rDNA phylogenetic analyses suggest that those single-genome (single-species) lineages could be interpreted as new genera. However, the proposal of new monotypic genera has been disputed (Lachance 2018). In fact, monotypic genera are not rare for yeasts, e.g., the genera Aciculiconidium, Babjeviella, Cyrenella and Kriegeria (Kurtzman 2011b; Sampaio 2011; Sampaio & Oberwinkler 2011; Smith 2011). This issue is not only relevant for interpreting phylogenies of yeasts, but is also true for all fungi. Hyde et al. (2024) demonstrated that 39.5% of described fungal genera are monotypic and argued whether other species in these monotypic genera will be found during a worldwide survey. Bhunjun et al. (2022) demonstrated that, likely, new species will be found and described after monotypic genera are published. For various initially monotypic yeast genera, such as Babjevia, Deakozyma, Nematodospora and Yueomyces, new species have been described shortly after the genus was proposed (Gouliamova et al. 2016; Ren et al. 2016; Zheng et al. 2017; Yamazaki et al. 2020; Yu et al. 2023).
To assign single-species Candida lineages as new genera has two benefits: 1) the proposal of monotypic genera might be useful as it will reduce the polyphyletic nature of the genus Candida (Takashima & Sugita 2022); 2) the proposal of monotypic genera will accelerate new species descriptions and assignment to those monotypic genera. Many undescribed Candida species have been deposited in the GenBank database (https://www.ncbi.nlm.nih.gov/nucle-otide/), and some of them are closely related to the single-species Candida lineages that we recognized in this work. For example, more than one potential new species is closely related to C. alai, C. bentonensis, C. boidinii, C. chilensis, C. freyschussii, C. hispaniensis, C. multigemmis, C. sake and C. sungouii (Fig. 5, Figs. S5–S13, and see in the phylogenetic analysis section). Those undescribed species (or potential new species) hamper the study of yeast diversity, taxonomy, and even our understanding of yeasts’ evolution. Thus, those single-species Candida lineages containing potential new species need to be assigned to new genera and making it possible to formally describe those species new to science in the future. For the time being, we decided to consider single-species Candida lineages without known closely related undescribed species, such as Candida anutae, Candida argente, C. ascalaphidarum, C. glucosophila and C. tunisiensi, as lineages in Candida pro tempore.
Strengths and limitations of the genome-based phylogeny of Candida
Since the genus Candida was introduced (Berkhout 1923), the concept and taxonomy of this genus have been changed various times (Lachance et al. 2011). The molecular sequence, including the LSU rDNA and ITS, studies demonstrated that the genus Candida, initially defined based on the phenotypic characteristics, is highly polyphyletic, and species can be found in various families across Saccharomycotina (Kurtzman & Robnett 1998, 2003; Lachance et al. 2011; Daniel et al. 2014; Takashima & Sugita 2022). This genus name is, however, of high importance in the medical field due to some species, such as C. albicans, C. dubliniensis, C. parapsilosis and C. tropicalis, which are frequently isolated from patients and can cause various infections, including candidemia (Kurtzman et al. 2011; Stavrou et al. 2019; Takashima & Sugita 2022). The current knowledge shows that most medically important Candida species occur in the Serinales (Pichiomycetes), whereas few of them, such as Candida glabrata (currently named Nakaseomyces glabratus), are classified in the Saccharomycetales (Saccharomycetes) (Takashima & Sugita 2022; Groenewald et al. 2023), which indicates that most Candida species in other orders and classes are not pathogenic, at least not yet.
In contrast, some Candida species are used in the biotechnological and fermentation industries, i.e., Cyberlindnera jadinii (asexual morph Candida utilis), are used for protein production, biosynthesis of (R)-phenylacetylcarbinol and other biotechnological products (Miura et al. 1998; Lee & Kim 2001; Villas-Bôas et al. 2002; Kurtzman 2011d); and Candida apicola, Candida batistae, Candida kuoi, Candida riodocensis and Candida stellata produce sophorolipids (Kurtzman et al. 2015). To reduce the heterogeneity of Candida from an evolutionary and taxonomic perspective and to distinguish it in the medical and biotechnological fields, it is necessary to reclassify the genus. More than half of the Candida species have been reassigned into existing and newly proposed genera (Takashima & Sugita 2022; Liu et al. 2024b), but the heterogeneous nature of the genus remains, and especially the phylogenetic positions of those Candida species that occur as deep internal branches in the LSU rDNA or multigene-based trees were not resolved (Lachance et al. 2011).
In this study, we mostly used the phylogenomic analysis and the complementary genome-based metrics, including the AAI, POCP and PAPO that have been used to discriminate genera in Saccharomycetaceae and Metschnikowiaceae (Liu et al. 2024a, b) and the RED approach that has been applied to evaluate the evolutionary divergences of different taxonomic ranks across fungi (Li et al. 2021; Groenewald et al. 2023), to reclassify the genus Candida. Our analyses assigned most Candida species into phylogenetically distinct known or newly created genera. The phylogenetic and taxonomic positions of species, which were unresolved based on LSU rDNA or multigene-based datasets, were resolved based on the phylogenomic analysis and the genome-based metric analyses in this study. For example, the phylogenetic position of C. anutae, C. argentea, C. insectalens, C. silvatica, and C. sorboxylosa varied in different phylogenetic studies due to the high divergent LSU rDNA sequence (Bab'eva et al. 2000; Holland et al. 2011), but our phylogenomic analysis showed that 1) C. anutae and C. argentea are closely related to Wickerhamia fluorescens; 2) C. insectalens and C. silvatica form a clade related to the genus Brettanomyces; 3) C. sorboxylosa is located in the genus Pichia, which are all in agreement with the results from Opulente et al. (2024).
Although most issues of Candida taxonomy have been resolved in this study, 15 Candida species were assigned as Candida pro tempore, especially those single-species lineages in the phylogenomic trees (Figs. 1, 6, 9, 12, 14–15, 17 and Figs. S1–S4). As we discussed above, those single-species Candida lineages will be resolved with more related species described. The other limitation of this reclassification is that seven newly created genera, i.e., Buckleya, Keqinozyma, Ramirezia, Ruyongia, Taiozyma and Wenyingozyma, are phylogenetically distinct from other genera, but they are described mostly based on phylogenomic and genome-based metric analyses without distinguishing phenotypic characteristics because species in those genera and their related genera are phenotypically divergent and do not have physiological or biochemical synapomorphic features (Table S3). This problem may be resolved based on the chromosome syntenic analysis or a special metabolic network analysis with complete genome (T2T assembled genome) datasets in the future.
The other limitation of this study is that the generic AAI and POCP values recommended by Liu et al. (2024a) were not suitable for some genera (or clades), such as the genus Yamadazyma, the Ogataea clade 1 and the Ogataea clade 2, which are characterized by high genetic diversity compared to the genera accepted in Saccharomycetaceae (Liu et al. 2024a) and Pichiales (Table 1), e.g. Citeromyces, Komagataella and Kregervanrija. Therefore, a fixed generic boundary of the genome-based metrics should be addressed for different families or higher ranks, as was done in prokaryotes by Riesco & Trujillo (2024), in the future. The combined approaches, phylogenomic analysis, phenotypic synapomorphic comparison and the genome-based metrics, are recommended to provide a conceptual or methodological framework for yeasts, and even for other fungal groups, taxonomic delineation.
Conclusions
We assigned 160 Candida species to 25 new genera, 6 reinstated genera and 15 known genera, including 4 validated genera, in this study. However, for pragmatic reasons, 15 Candida species are listed as pro tempore as we suggest to delay taxonomic conclusions until consensus is reached among yeast taxonomists regarding the classification of those lineages. Among those unresolved Candida species, C. tunisiensis occurs in the Dipodascomycetes; C. argentea, C. anglica, C. anutae, C. ascalaphidarum and C. glucosophila occur in the Debaryomycetaceae (Serinales, Pichiomycetes); C. citri, C. danieliae, C. golubevii, C. hainanensis, C. hawaiiana, C. magnifica, C. wancherniae and C. xylosifermentans locate in the Metschnikowiaceae (Serinales, Pichiomycetes); C. sanyiensis occurs in Barnettozyma (Phaffomycetaceae, Phaffomycetales, Saccharomycetes). Like Liu et al. (2024b), we refrain from making taxonomic conclusions on those lineages as the discovery of more related species is needed to classify them properly.
Taxonomy
In this section, we propose a new classification of 160 Candida species and related taxa in Saccharomycotina presented in Tables S1–S2, including the reinstatement of several genera, i.e., Entelexis, Komagataea, Petasospora, Waltiozyma and Williopsis based on the results of the studies discussed above. Furthermore, 91 invalid names of the taxa indicated in the Index Fungorum, Fungal Names and MycoBank are validated. The four invalid genera, namely Deakozyma, Limtongella, Tardiomyces and Hemisphaericaspora, are validated as new genera; the other 89 invalid species are treated as new species for validation. Note that we list only the most important synonyms for those species with applied and clinical relevance. For the full list of synonyms, we refer to The Yeasts database (https://theyeasts.org) and TYTS, 5th edition (Kurtzman et al. 2011). All proposed new taxa are listed in Table 5. The genus-specific OGs (unique genes) used as diagnostic characters for the newly proposed, emended and reinstated genera are listed in Table 6.
| Taxa | Basionym or important synonyms |
| Dipodascomycetes | |
| Dipodascales | |
| Trichomonascaceae | |
| Casaregolazyma gen. nov. | |
| C. lundiana comb. nov. | Candida lundiana |
| C. patagonica sp. nov. | Candida patagonica |
| C. suthepensis comb. nov. | Candida suthepensis |
| Daia gen. nov. | |
| D. ofunaensis comb. nov. | Hansenula ofunaensis; Zygoascus ofunaensis |
| D. tannicola comb. nov. | Pichia tannicola; Zygoascus tannicola |
| Deakozyma gen. nov. | |
| D. indianensis sp. nov. | Deakozyma indianensis |
| D. yunnanensis sp. nov. | Deakozyma yunnanensis |
| Entelexis | |
| E. apis comb. nov. | Candida apis; Starmerella apis; Torulopsis apis |
| E. geochares comb. nov. | Candida geochares; Starmerella geochares; Torulopsis geochares |
| E. gropengiesseri comb. nov. | Candida gropengiesseri; Starmerella gropengiesseri; Torula gropengiesseri |
| E. litoralis comb. nov. | Starmerella litoralis |
| E. magnoliae | |
| E. paramagnoliae nom. nov. | Candida magnoliae; Starmerella magnoliae; Torulopsis magnoliae |
| E. potacharoeniae comb. nov. | Candida potacharoeniae; Starmerella potacharoeniae |
| E. sorbosivorans comb. nov. | Candida sorbosivorans; Starmerella sorbosivorans |
| E. spenceri comb. nov. | Candida spenceri |
| E. syriaca sp. nov. | Starmerella syriaca |
| E. tilneyi sp. nov. | Candida tilneyi; Starmerella tilneyi |
| E. vaccinii comb. nov. | Candida vaccinii; Starmerella vaccinii |
| E. xylocopis comb. nov. | Starmerella xylocopis |
| Grinbergsozyma gen. nov. | |
| G. bentonensis comb. nov. | Candida bentonensis |
| Limtongella gen. nov. | |
| L. siamensis sp. nov. | Limtongella siamensis |
| L. incommunis comb. nov. | Candida incommunis |
| Starmerella sensu stricto | |
| S. powellii sp. nov. | Candida powellii; Starmerella powellii |
| Tardiomyces gen. nov. | |
| T. blankii comb. nov. | Candida blankie; Tardiomyces blankii |
| T. depauwii sp. nov. | Tardiomyces depauwii |
| T. digboiensis comb. nov. | Candida digboiensis; Tardiomyces digboiensis |
| Westerdijkia gen. nov. | |
| W. sungouii sp. nov. | Candida sungouii |
| Incertae sedis in Dipodascomycetes | |
| Dengshuqunia gen. nov. | |
| D. hispaniensis comb. nov. | Candida hispaniensis |
| Yarrowia | |
| Y. brassicae sp. nov. | Yarrowia brassicae |
| Y. divulgata sp. nov. | Yarrowia divulgata |
| Y. keelungensis sp. nov. | Yarrowia keelungensis |
| Y. phangngaensis sp. nov. | Candida phangngaensis; Yarrowia phangngaensis |
| Pichiomycetes | |
| Pichiales | |
| Pichiaceae | |
| Ogataea | |
| O. arabinofermentans comb. nov. | Candida arabinofermentans |
| O. chonburiensis sp. nov. | Ogataea chonburiensis |
| O. chumphonensis sp. nov. | Candida chumphonensis |
| O. deakii sp. nov. | Ogataea deakii |
| O. ganodermae sp. nov. | Ogataea ganodermae |
| O. histrianica sp. nov. | Ogataea histrianica |
| O. kanchanaburiensis sp. nov. | Ogataea kanchanaburiensis |
| O. kolombanensis sp. nov. | Ogataea kolombanensis |
| O. krabiensis sp. nov. | Candida krabiensis |
| O. maris comb. nov. | Candida maris; Torulopsis maris |
| O. mattraensis sp. nov. | Candida mattraensis |
| O. nakhonphanomensis sp. nov. | Ogataea nakhonphanomensis |
| O. nemodendra comb. nov. | Candida nemodendra; Torulopsis nemodendra |
| O. ortonii sp. nov. | Candida ortonii |
| O. ovalis comb. nov. | Candida ovalis |
| O. piceae comb. nov. | Candida piceae |
| O. pinus comb. nov. | Candida pinus; Torulopsis pinus |
| O. phyllophila sp. nov. | Ogataea phyllophila |
| O. rishirensis comb. nov. | Candida rishirensis |
| O. siamensis sp. nov. | Ogataea siamensis; Pichia siamensis |
| O. sithepensis sp. nov. | Candida sithepensis |
| O. sonorensis comb. nov. | Candida sonorensis; Torulopsis sonorensis |
| O. thermomethanolica sp. nov. | Ogataea thermomethanolica; Pichia thermomethanolica |
| O. thermophila sp. nov. | Candida thermophila |
| O. xyloterini comb. nov. | Candida xyloterini |
| Pichia | |
| P. awuae sp. nov. | Candida awuae |
| P. bruneiensis sp. nov. | Pichia bruneiensis |
| P. chibodasensis sp. nov. | Pichia chibodasensis |
| P. dushanensis sp. nov. | Pichia dushanensis |
| P. insulana sp. nov. | Pichia insulana |
| P. phayaonensis sp. nov. | Candida phayaonensis; Pichia phayaonensis |
| P. sorboxylosa comb. nov. | Candida sorboxylosa |
| P. thaimueangensis sp. nov. | Candida thaimueangensis; Pichia thaimueangensis |
| Ramirezia gen. nov. | |
| R. boidinii comb. nov. | Candida boidinii |
| Wenyingozyma gen. nov. | |
| W. methanosorbosa comb. nov. | Candida methanosorbosa; Torulopsis methanosorbosa |
| W. methylovora comb. nov. | Ogataea methylovora; Pichia methylovora |
| W. naganishii comb. nov. | Ogataea naganishii; Pichia naganishii |
| W. nanaspora comb. nov. | Candida nanaspora |
| W. neixiangensis comb. nov. | Ogataea neixiangensis |
| W. nitratophila comb. nov. | Candida nitratophila; Torulopsis nitratophila |
| W. ramenticola comb. nov. | Ogataea ramenticola; Pichia ramenticola |
| W. succiphila comb. nov. | Candida cellulolytica; Candida succiphila |
| W. suzukii comb. nov. | Candida suzukii |
| W. wangdongensis sp. nov. | Ogataea wangdongensis |
| Xiuguozyma gen. nov. | |
| X. insectalens comb. nov. | Candida insectalens; Torulopsis insectalens |
| X. silvatica comb. nov. | Candida silvatica; Torulopsis silvatica |
| Serinales | |
| Debaryomycetaceae | |
| Candida emend. | |
| C. baotianmanensis sp. nov. | Candida baotianmanensis |
| C. pseudoviswanathii sp. nov. | Candida pseudoviswanathii |
| C. sanyaensis sp. nov. | Candida sanyaensis |
| C. saraburiensis sp. nov. | Candida saraburiensis |
| Chernovozyma gen. nov. | |
| C. aurita comb. nov. | Candida aurita |
| C. palmyrensis comb. nov. | Candida palmyrensis |
| C. sophiae-reginae comb. nov. | Candida sophiae-reginae |
| Debaryomyces | |
| D. psychrophila comb. nov. | Candida psychrophila; Torulopsis psychrophila |
| D. renaianus sp. nov. | Debaryomyces renaii |
| D. vindobonensis sp. nov. | Debaryomyces vindobonensis |
| Dujonia gen. nov. | |
| D. boleticola comb. nov. | Candida boleticola |
| D. oleophila comb. nov. | Candida oleophila |
| D. railenensis comb. nov. | Candida railenensis |
| D. santamariae comb. nov. | Candida santamariae |
| D. schatavii comb. nov. | Candida schatavii; Torulopsis schatavii |
| D. zeylanoides comb. nov. | Candida zeylanoides; Monilia zeylanoides |
| Fermentozyma gen. nov. | |
| F. sake comb. nov. | Candida sake; Eutorulopsis sake |
| F. vespimorsuum comb. nov. | Candida vespimorsuum |
| Glucitozyma gen. nov. | |
| G. multigemmis comb. nov. | Candida multigemmis; Torulopsis multigemmis |
| Hemisphaericaspora gen. nov. | |
| H. elongata comb. nov. | Spathaspora elongata |
| H. gorwiae comb. nov. | Spathaspora gorwiae |
| H. haegerdaliae comb. nov. | Spathaspora haegerdaliae |
| H. insectamans comb. nov. | Candida insectamans; Hemisphaericaspora insectamans |
| H. jiuxiensis comb. nov. | Spathaspora jiuxiensis |
| H. lyxosophila comb. nov. | Candida lyxosophila |
| H. mengyangensis comb. nov. | Spathaspora mengyangensis |
| H. nanyangensis sp. nov. | Hemisphaericaspora nanyangensis |
| H. parajiuxiensis comb. nov. | Spathaspora parajiuxiensis |
| H. roraimensis comb. nov. | Spathaspora roraimensis |
| H. rosae comb. nov. | Spathaspora rosae |
| H. subhashii comb. nov. | Candida subhashii |
| H. xylanilytica sp. nov. | Candida xylanilytica |
| H. xylofermentans comb. nov. | Spathaspora xylofermentans |
| Insectozyma gen. nov. | |
| I. bohioensis comb. nov. | Candida bohioensis |
| I. chauliodis comb. nov. | Candida chauliodis |
| I. coleopterorum comb. nov. | Candida coleopterorum |
| I. corydali comb. nov. | Candida corydali |
| I. morakotiae comb. nov. | Candida morakotiae |
| I. parachauliodis comb. nov. | Candida parachauliodis |
| I. prachuapensis sp. nov. | Candida prachuapensis |
| I. sakaeoensis sp. nov. | Candida sakaeoensis |
| I. verbasci sp. nov. | Candida verbasci |
| I. xiaguanensis comb. nov. | Candida xiaguanensis |
| Intestinozyma gen. nov. | Candida verbasci |
| I. alai comb. nov. | Candida alai |
| Keqinozyma gen. nov. | |
| K. heliconiae comb. nov. | Candida heliconiae |
| K. nonsorbophila comb. nov. | Candida nonsorbophila |
| K. sinolaborantium comb. nov. | Candida sinolaborantium |
| K. temnochilae comb. nov. | Candida temnochilae |
| Lizanozyma gen. nov. | |
| L. gosingicus sp. nov. | Candida gosingica; Scheffersomyces gosingicus |
| L. spartinae comb. nov. | Pichia spartinae; Scheffersomyces spartinae |
| L. thasaensis sp. nov. | Candida thasaensis |
| Lodderomyces | |
| L. cetoniae comb. nov. | Candida cetoniae |
| L. hyderabadensis comb. nov. | Candida hyderabadensis |
| L. jiufengensis comb. nov. | Candida jiufengensis |
| L. margitis comb. nov. | Candida margitis |
| L. metapsilosis comb. nov. | Candida metapsilosis |
| L. orthopsilosis comb. nov. | Candida orthopsilosis |
| L. oxycetoniae comb. nov. | Candida oxycetoniae |
| L. parapsilosis comb. nov. | Candida parapsilosis; Monilia parapsilosis |
| L. pseudojiufengensis comb. nov. | Candida pseudojiufengensis |
| L. theae sp. nov. | Candida theae |
| Millerozyma | |
| M. porticicola comb. nov. | Pichia porticicola |
| M. pseudofarinosa comb. nov. | Candida pseudofarinosa |
| Nothofagozyma gen. nov. | |
| N. chilensis comb. nov. | Candida chilensis |
| Scheffersomyces | |
| S. broadrunensis comb. nov. | Candida broadrunensis |
| S. lignicola sp. nov. | Candida lignicola; Scheffersomyces lignicola |
| Spathaspora sensu stricto | |
| S. brasiliensis sp. nov. | Spathaspora brasiliensis |
| S. suhii sp. nov. | Spathaspora suhii |
| Suhomyces | |
| S. caryicola comb. nov. | Candida caryicola |
| S. rongomai-pounamu comb. nov. | Candida rongomai-pounamu |
| S. tibetensis comb. nov. | Candida tibetensis |
| Suzukiozyma gen. nov. | |
| S. candida comb. nov. | Candida saitoana; Torulopsis candida |
| S. fluviatilis comb. nov. | Candida fluviatilis |
| S. glaebosa comb. nov. | Candida glaebosa |
| S. manassasensis comb. nov. | Candida manassasensis |
| S. palmioleophila comb. nov. | Candida palmioleophila |
| S. pseudoglaebosa comb. nov. | Candida pseudoglaebosa |
| S. sphagnicola comb. nov. | Candida sphagnicola |
| Yamadazyma | |
| Y. andamanensis sp. nov. | Candida andamanensis |
| Y. dushanensis sp. nov. | Yamadazyma dushanensis |
| Y. oceani sp. nov. | Candida oceani; Yamadazyma oceani |
| Y. paraphyllophila sp. nov. | Yamadazyma paraphyllophila |
| Y. phyllophila sp. nov. | Yamadazyma phyllophila |
| Y. jaroonii sp. nov. | Candida jaroonii |
| Y. siamensis sp. nov. | Yamadazyma siamensis |
| Y. songkhlaensis sp. nov. | Candida songkhlaensis |
| Y. ubonensis sp. nov. | Yamadazyma ubonensis |
| Y. vrieseae sp. nov. | Candida vrieseae |
| Zhuliangozyma gen. nov. | |
| Z. blackwelliae comb. nov. | Candida blackwellae |
| Z. boniae comb. nov. | Spathaspora boniae |
| Z. parablackwelliae comb. nov. | Candida parablackwelliae |
| Saccharomycetes | |
| Phaffomycetales | |
| Phaffomycetaceae | |
| Barnettozyma | |
| B. sucrosica sp. nov. | Barnettozyma sucrosica |
| B. xylosica sp. nov. | Barnettozyma xylosica |
| B. xylosiphila sp. nov. | Barnettozyma xylosiphila |
| Gotozyma gen. nov. | |
| G. botsteinii sp. nov. | Barnettozyma botsteinii |
| G. montana comb. nov. | Candida montana |
| G. siamensis comb. nov. | Barnettozyma siamensis |
| Komagataea | |
| K. norvegica comb. nov. | Candida norvegica; Torulopsis norvegica |
| K. qinlingensis comb. nov. | Candida qinlingensis |
| K. salicaria comb. nov. | Barnettozyma salicaria; Pichia salicaria |
| K. wickerhamii comb. nov. | Barnettozyma wickerhamii; Endomycopsis wickerhamii |
| Millerago | |
| M. ficus sp. nov. | Candida ficus |
| Phaffomyces | |
| P. coquimbonensis sp. nov. | Candida coquimbonensis |
| P. orba comb. nov. | Candida orba |
| Wickerhamomycetaceae | |
| Buckleya gen. nov. | |
| B. freyschussii comb. nov. | Candida freyschussii |
| Hansenula | |
| H. queroliae comb. nov. | Wickerhamomyces queroliae |
| H. silvicultrix comb. nov. | Candida silvicultrix |
| H. spegazzinii sp. nov. | Wickerhamomyces spegazzinii |
| H. sylviae sp. nov. | Wickerhamomyces sylviae |
| Liangdongia gen. nov. | |
| L. berthetii comb. nov. | Candida berthetii |
| L. dendrica comb. nov. | Candida dendrica; Torulopsis dendrica |
| L. dryadoides comb. nov. | Hansenula dryadoides; Starmera dryadoides |
| L. ilhagrandensis comb. nov. | Starmera ilhagrandensis |
| L. laemsonensis sp. nov. | Candida laemsonensis |
| L. nongkratonensis comb. nov. | Pichia nongkratonensis |
| L. prunicorticola comb. nov. | Starmera prunicorticola |
| L. quercuum comb. nov. | Pichia quercuum; Starmera quercuum |
| Petasospora | |
| P. adriatica comb. nov. | Candida adriatica |
| P. americana comb. nov. | Cyberlindnera americana; Hansenula bimundalis var. americana |
| P. amylophila comb. nov. | Cyberlindnera amylophila; Pichia amylophila |
| P. bimundalis comb. nov. | Cyberlindnera bimundalis; Hansenula bimundalis |
| P. dasilvae comb. nov. | Cyberlindnera dasilvae |
| P. easanensis sp. nov. | Candida easanensis |
| P. euphorbiae comb. nov. | Cyberlindnera euphorbiae; Pichia euphorbiae |
| P. euphorbiiphila comb. nov. | Cyberlindnera euphorbiiphila; Hansenula euphorbiiphila |
| P. fabianii comb. nov. | Cyberlindnera fabianii; Hansenula fabianii |
| P. hubeiensis comb. nov. | Cyberlindnera hubeiensis |
| P. hungchunana sp. nov. | Candida hungchunana |
| P. japonica comb. nov. | Cyberlindnera japonica; Pichia japonica |
| P. juglandicorticola comb. nov. | Cyberlindnera juglandicorticola |
| P. maesae sp. nov. | Candida maesae |
| P. maritima comb. nov. | Candida maritima; Cyberlindnera maritima; Trichosporon maritimum |
| P. meyerae comb. nov. | Cyberlindnera meyerae; Pichia meyerae |
| P. mississippiensis comb. nov. | Cyberlindnera mississippiensis; Pichia mississippiensis |
| P. mycetangii comb. nov. | Candida mycetangii; Cyberlindnera mycetangii |
| P. nakhonratchasimensis sp. nov. | Candida nakhonratchasimensis; Cyberlindnera nakhonratchasimensis |
| P. pattaniensis sp. nov. | Candida pattaniensis |
| P. petersonii comb. nov. | Cyberlindnera petersonii; Hansenula petersonii |
| P. rhizosphaerae comb. nov. | Cyberlindnera rhizosphaerae; Lindnera rhizosphaerae |
| P. shennongjiaensis | Cyberlindnera shennongjiaensis |
| P. stauntonica sp. nov. | Candida stauntonica |
| P. sylvatica comb. nov. | Cyberlindnera sylvatica |
| P. taoyuanica sp. nov. | Candida taoyuanica |
| P. veronae comb. nov. | Cyberlindnera veronae; Pichia veronae |
| P. wuzhiensis comb. nov. | Cyberlindnera wuzhiensis; Lindnera wuzhiensis |
| P. xishuangbannaensis comb. nov. | Cyberlindnera xishuangbannaensis |
| P. xylebori comb. nov. | Cyberlindnera xylebori |
| P. xylosilytica comb. nov. | Cyberlindnera xylosilytica |
| Ruyongia gen. nov. | |
| R. chambardii comb. nov. | Saccharomyces chambardii; Wickerhamomyces chambardii |
| R. mori comb. nov. | Wickerhamomyces mori |
| R. namnaoensis comb. nov. | Candida namnaoensis |
| R. patagonica comb. nov. | Wickerhamomyces patagonicus |
| R. ponderosae comb. nov. | Candida ponderosae |
| R. rarassimilans comb. nov. | Pichia rarassimilans |
| R. tratensis comb. nov. | Wickerhamomyces tratensis |
| Taiozyma gen. nov. | |
| T. bovis comb. nov. | Pichia bovis; Wickerhamomyces bovis |
| T. dajiaensis sp. nov. | Candida dajiaensis |
| T. odintsovae comb. nov. | Candida odintsovae |
| T. onychis comb. nov. | Pichia onychis; Wickerhamomyces onychis |
| T. peoriensis comb. nov. | Candida peoriensis |
| T. rabaulensis comb. nov. | Pichia rabaulensis; Wickerhamomyces rabaulensis |
| T. yuanshanica sp. nov. | Candida yuanshanica |
| Waltiozyma | |
| W. chaumierensis comb. nov. | Wickerhamomyces chaumierensis |
| W. pijperi comb. nov. | Pichia pijperi; Wickerhamomyces pijperi |
| W. solani comb. nov. | Candida solani |
| W. xylosica sp. nov. | Wickerhamomyces xylosica |
| Wickerhamomyces sensu stricto | |
| W. jianshihensis sp. nov. | Candida jianshihensis |
| W. quercuum comb. nov. | Candida quercuum |
| W. ulmi comb. nov. | Candida ulmi |
| Williopsis | |
| W. culbertsonii sp. nov. | Cyberlindnera culbertsonii |
| W. dauci comb. nov. | Cyberlindnera dauci |
| W. galapagoensis comb. nov. | Cyberlindnera galapagoensis |
| W. jadinii comb. nov. | Candida utilis; Cyberlindnera jadinii; Saccharomyces jadinii |
| W. lachancei comb. nov. | Cyberlindnera lachancei; Pichia lachancei |
| W. maclurae comb. nov. | Cyberlindnera maclurae; Pichia maclurae |
| W. mengyuniae comb. nov. | Candida mengyuniae |
| W. misumaiensis comb. nov. | Cyberlindnera misumaiensis; Pichia misumaiensis |
| W. samutprakarnensis sp. nov. | Cyberlindnera samutprakarnensis |
| W. sargentensis comb. nov. | Cyberlindnera sargentensis; Pichia sargentensis |
| W. takata sp. nov. | Candida takata |
| W. tropicalis comb. nov. | Cyberlindnera tropicalis |
| W. vartiovaarae comb. nov. | Candida vartiovaarae; Cyberlindnera vartiovaarae; Torulopsis vartiovaarae |
| Xingzhongia gen. nov. | |
| X. hampshirensis comb. nov. | Pichia hampshirensis; Wickerhamomyces hampshirensis |
| X. scolytoplatypi comb. nov. | Wickerhamomyces scolytoplatypi |
| X. strasburgensis comb. nov. | Saccharomyces strasburgensis; Wickerhamomyces strasburgensis |
Table 6 List of the genus-specific OGs (unique genes) to use as diagnostic characters for the newly proposed, emended and reinstated genera
| Taxa | OGs to use as diagnostic characters |
| Dipodascomycetes | |
| Casaregolazyma | OG0008493; OG0010589; OG0011717 |
| Daia | OG0017790; OG0024243; OG0024231 |
| Deakozyma | OG0020116; OG0020113; OG0020111 |
| Dengshuqunia | OG0013204; OG0015719; OG0015720 |
| Entelexis | OG0009064; OG0009065 |
| Grinbergsozyma | OG0015427; OG0015488; OG0019665 |
| Tardiomyces | OG0013179; OG0015603; OG0015572 |
| Pichiomycetes | |
| Pichiales | |
| Ramirezia | OG0009746; OG0009748; OG0009751 |
| Wenyingozyma | OG0006088; OG0006166; OG0006167 |
| Xiuguozyma | OG0009801; OG0009803; OG0011903 |
| Serinales | |
| Candida sensu stricto clade | OG0006359; OG0006430 |
| Chernovozyma | OG0009945; OG0009948; OG0010921 |
| Dujonia | OG0009139 |
| Fermentozyma | OG0012707; OG0015120; OG0019183 |
| Glucitozyma | OG0018541; OG0018542; OG0018545 |
| Insectozyma | OG0008208; OG0008209; OG0008210 |
| Intestinozyma | OG0014078; OG0014079; OG0017124 |
| Keqinozyma | OG0018587; OG0018588; OG0018589 |
| Lizanozyma | OG0013736; OG0016664; OG0022956 |
| Nothofagozyma | OG0017645; OG0017647; OG0017651 |
| Suzukiozyma | OG0007615; OG0007616; OG0007620 |
| Zhuliangozyma | OG0009199; OG0009200; OG0012185 |
| Saccharomycetes | |
| Phaffomycetales | |
| Buckleya | OG0010458; OG0010457; OG0008787 |
| Hansenula | OG0006330; OG0006334; OG0006336 |
| Komagataea | OG0007192; OG0007193; OG0007195 |
| Lacusozyma | OG0009727; OG0012681; OG0012683 |
| Liangdongia | OG0007703; OG0007712; OG0007714 |
| Taxa | OGs to use as diagnostic characters |
| Petasospora | OG0005539; OG0005586 |
| Ruyongia | OG0007319; OG0007880; OG0007918 |
| Taiozyma | OG0006108; OG0006424 |
| Waltiozyma | OG0007005; OG0007007; OG0007012 |
| Williopsis | OG0005812; OG0005845; OG0005847 |
| Xingzhongia | OG0012666; OG0012667; OG0012668 |
New taxa in Trichomonascaceae (Dipodascales, Dipodascomycetes)
Casaregolazyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857878
Etymology: the genus is named in honor of S. Casaregola for his contribution to yeast taxonomy.
Type species: Casaregolazyma lundiana (Saks., M. Suzuki, Lumyong, Ohkuma & Chantaw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Candida lundiana clade, which occur in a separate lineage from the Zygoascus sensu stricto clade and the Zygoascus tannicola clade (Fig. 1). Member of the Trichomonascaceae (Dipodascales, Dipodascomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0008493 and OG0010589 (Table 6).
Sexual reproduction not known. Colonies are white, cream, convex, fimbriate. Budding is multilateral. Pseudohyhae are present, hyphae are present or absent. Fermentation is absent.
Note: Casaregolazyma spp. can be distinguished from those of the closely related genera Daia and Zygoascus sensu stricto by lack of fermentation (Table S3).
Species accepted:
Casaregolazyma lundiana (Saks., M. Suzuki, Lumyong, Ohkuma & Chantaw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857879
Holotype: CBS 12271, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. THAILAND, isolated from the raw honey of Homotrigona fimbriata, December 2006.
Basionym: Candida lundiana Saks., M. Suzuki, Lumyong, Ohkuma & Chantaw., Antonie van Leeuwenhoek 101: 636. 2012.
Casaregolazyma patagonica Sangorrín, C.A. Lopes, Belloch, Querol & A.C. Caball. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857880
For a description: see Sangorrínet al., Antonie van Leeuwenhoek 92: 80. 2007.
Holotype: CBS 10443, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ARGENTINA (North Patagonian Region), isolated from walls of fermentation vats and oak barrels in wine cellars, 2005.
Synonym: Candida patagonica Sangorrín, C.A. Lopes, Belloch, Querol & A.C. Caball., Antonie van Leeuwenhoek 92: 80. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Casaregolazyma suthepensis (Saks., M. Suzuki, Lumyong, Ohkuma & Chantaw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857881
Holotype: CBS 12270, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. THAILAND, isolated from raw honey of the Asian cavity-nesting honeybee (Apis cerana), February 2009.
Basionym: Candida suthepensis Saks., M. Suzuki, Lumyong, Ohkuma & Chantaw., Antonie van Leeuwenhoek 101: 637. 2012.
Daia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857911
Etymology: the genus is named in honor of Dr. Yu-Chen Dai for his contributions to the taxonomy of Basidiomycota, especially of macrofungal taxonomy.
Type species: Daia tannicola (F.H. Jacob) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Zygoascus tannicola clade, which occur in a separate lineage from the Zygoascus sensu stricto clade and the Candida lundiana clade (Fig. 1). Member of the Trichomonascaceae (Dipodascales, Dipodascomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0017790 and OG0024231 (Table 6).
Asci are persistent and contain one to four subspherical or hat-shaped ascospores. Colonies are tannish-white, butyrous, glistening. Budding is multilateral. Pseudohyphae are present, but hyphae are observed.
Note: Daia spp. can be distinguished from those of the closely related genera Casaregolazyma and Zygoascus sensu stricto by assimilation of melibiose (Table S3). Unlike Zygoascus sensu stricto, Daia species do not produce septate hyphae.
Species accepted:
Daia ofunaensis (Makig. & Asai) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857912
Holotype: CBS 8129, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. JAPAN, Ofuna, isolated from soil.
Basionym: Hansenula ofunaensis Makig. & Asai, J.Gen. Appl. Microbiol. 22: 200. 1976.
Synonym: Zygoascus ofunaensis (Makig. & Asai) Kurtzman & Robnett, FEMS Yeast Res. 7: 147. 2007.
Daia tannicola (F.H. Jacob) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857914
Holotype: CBS 6065, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. FRANCE, isolated from vegetable tanning fluid.
Basionym: Pichia tannicola F.H. Jacob, Bull. Trimestriel Soc. Mycol. France 85: 111. 1969.
Synonym: Zygoascus tannicola (F.H. Jacob) Kurtzman & Robnett [as 'tannicolus'], FEMS Yeast Res. 7: 147. 2007.
Deakozyma Kurtzman & Robnett ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857882
For a description: see Kurtzman & Robnett, Antonie van Leeuwenhoek 105: 938. 2014.
Synonym: Deakozyma Kurtzman & Robnett, Antonie van Leeuwenhoek 105: 938. 2014. Nom. inval., Art. 40.1 (Melbourne), the type species is invalid as more than one collection in which the type is conserved was specified.
Type species: Deakozyma indianensis Kurtzman & Robnett ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
Species accepted:
Deakozyma indianensis Kurtzman & Robnett ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857883
For a description: see Kurtzman & Robnett, Antonie van Leeuwenhoek 105: 938. 2014.
Holotype: NRRL YB-1937, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. INDIANA, isolated from the eggs of an insect.
Synonym: Deakozyma indianensis Kurtzman & Robnett, Antonie van Leeuwenhoek 105: 938. 2014. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Deakozyma yunnanensis F.L. Hui, K.F. Liu & Xiao J. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857884
For a description: see Zeng et al., Int. J. Syst. Evol. Microbiol. 67: 2438. 2017.
Holotype: CICC 33160, preserved in a metabolically inactive state at the China Centre of Industrial Culture Collection (CICC), Beijing, China. CHINA, Yunnan province, Xishuangbanna tropical rainforest, isolated from rotten wood, July 2016.
Synonym: Deakozyma yunnanensis F.L. Hui, K.F. Liu & Xiao J. Liu, Int. J. Syst. Evol. Microbiol. 67: 2438. 2017. Nom. inval., Art. 35.1 (Shenzhen), the genus name is invalid.
Reinstated genus
Entelexis van der Walt & Johannsen, Antonie van Leeuwenhoek 39: 646. 1973.
MycoBank: MB1824
Type species: Entelexis magnoliae van der Walt & Johannsen, Antonie van Leeuwenhoek 39(4): 646. 1973.
Species accepted:
Entelexis apis (Lavie ex Uden & Vidal-Leir.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857888
Holotype: CBS 2674, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. UNITED KINGDOM, isolated from the trachea of Apis sp. (bee).
Basionym: Torulopsis apis Lavie ex Uden & Vidal-Leir., Yeasts, a taxonomic study, 2nd Edn (Amsterdam): 1245. 1970.
Synonyms: Candida apis (Lavie ex Uden & Vidal-Leir.) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 611. 1978.
Starmerella apis (Lavie ex Uden & Vidal-Leir.) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1340. 2018.
Entelexis geochares (van der Walt, Johannsen & Yarrow) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857889
Holotype: CBS 6870, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from isolated from soil.
Basionym: Torulopsis geochares van der Walt, Johannsen & Yarrow, Antonie van Leeuwenhoek 44: 98. 1978.
Synonyms: Candida geochares (van der Walt, Johannsen & Yarrow) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 612. 1978.
Starmerella geochares (van der Walt, Johannsen & Yarrow) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1341. 2018.
Entelexis gropengiesseri (F.C. Harrison) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857890
Holotype: CBS 156, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. UNKNOWN, isolated from the cocoon of cockroach (Periplaneta orientalis).
Basionym: Torula gropengiesseri F.C. Harrison, Trans. Roy. Soc. Canada, Sect. 5, Biol. Sci. 22: 187. 1928.
Synonyms: Candida gropengiesseri (F.C. Harrison) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 612. 1978.
Starmerella gropengiesseri (F.C. Harrison) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1341. 2018.
Entelexis litoralis (A.R.O. Santos, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857891
Holotype: CBS 14104, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, Bahia, a sand dune ecosystem of the municipality, isolated from the flower of Ipomoea imperati.
Basionym: Starmerella litoralis A.R.O. Santos, Lachance & C.A. Rosa, Int. J. Syst. Evol. Microbiol. 68: 1338. 2018.
Entelexis magnoliae van der Walt & Johannsen, Antonie van Leeuwenhoek 39(4): 646. 1973.
MycoBank: MB313623
Entelexis paramagnoliae (Lodder & Kreger-van Rij) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, nom. nov.
MycoBank: MB857893
Holotype: CBS 166, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. NETHERLANDS, isolated from the flower of Magnolia sp.
Replaced synonym: Torulopsis magnoliae Lodder & Kreger-van Rij, Yeasts, a Taxonomic Study, [Edn 1] (Amsterdam): 671. 1952, non Entelexis magnoliae Van der Walt & Johannsen.
Candida magnoliae (Lodder & Kreger-van Rij) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Starmerella magnoliae (Lodder & Kreger-van Rij) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1341. 2018.
Entelexis potacharoeniae (Nakase, Jindam., Imanishi, Am-In, Ninomiya, H. Kawas. & Limtong) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858031
Holotype: BCC 25963, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Bangkok, Chatuchak, isolated from flower of pink allamanda (Mandevilla sp.), March 2003.
Basionym: Candida potacharoeniae Nakase, Jindam., Imanishi, Am-In, Ninomiya, H. Kawas. & Limtong, J. Gen. Appl. Microbiol. 56: 291. 2010.
Synonym: Starmerella potacharoeniae (Nakase, Jindam., Imanishi, Am-In, Ninomiya, H. Kawas. & Limtong) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1341. 2018.
Entelexis sorbosivorans (S.A. James, C.J. Bond & I.N. Roberts) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857894
Holotype: NCYC 2938, preserved in a metabolically inactive state at the National Collection of Yeast Cultures, Norwich, UK. GREAT BRITAIN, isolated from the sorbitol-to-sorbose continuous fermentation cascade.
Basionym: Candida sorbosivorans S.A. James, C.J. Bond & I.N. Roberts, Int. J. Syst. Evol. Microbiol. 51: 1218. 2001.
Synonym: Starmerella sorbosivorans (S.A. James, C.J. Bond & I.N. Roberts) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1342. 2018.
Entelexis spenceri (Nakase, Jindam., Imanishi, Am-In, Ninomiya, H. Kawas. & Limtong) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857895
Holotype: BCC 15278, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Kanchanaburi, Amphoe Tong Pha Phum, isolated from the flower of the humble-plant (Mimosa pudica), November 2003.
Basionym: Candida spenceri Nakase, Jindam., Imanishi, Am-In, Ninomiya, H. Kawas. & Limtong, J. Gen. Appl. Microbiol. 56: 293. 2010.
Entelexis syriaca Sipiczki ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857897
For a description: see Sipiczki, Antonie van Leeuwenhoek 107: 851. 2014.
Holotype: CBS 13909, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SYRIA, Aleppo, isolated from the flowers of Malva sp.
Synonym: Starmerella syriaca Sipiczki, Antonie van Leeuwenhoek 107: 851. 2014. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Entelexis tilneyi Lachance ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857908
For a description: see Lachance et al., Int. J. Syst. Evol. Microbiol. 51: 1205. 2001.
Holotype: CBS 8794, preserved in a metabolically inactive state at Westerdijk Institute, Utrecht, the Netherlands. Costa Rica, Guanacaste, isolated from a beetle (Conotelus sp.) of a flower of Ipomoea carnea.
Synonyms: Candida tilneyi Lachance, Int. J. Syst. Evol. Microbiol. 51: 1205. 2001. Nom. inval., Art. 40.3 (Melbourne).
Starmerella tilneyi Lachance ex C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1342. 2018. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified; published as a combination, but 'basionym' (Candida tilneyi) is invalid.
Entelexis vaccinii (Tokuoka, Ishit., Goto & Komag.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857909
Holotype: CBS 7318, preserved in a metabolically inactive state at Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from the flower of blueberry (Vaccinium sp.).
Basionym: Candida vaccinii Tokuoka, Ishit., Goto & Komag., J. Gen. Appl. Microbiol. 33: 8. 1987.
Synonym: Starmerella vaccinii (Tokuoka, Ishit., Goto & Komag.) C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1342. 2018.
Entelexis xylocopis (Gouliam., Dimitrov, M. Groenew., M.T. Sm. & Boekhout) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857910
Holotype: CBS 5659, preserved in a metabolically inactive state at Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from the larval feed of an Afrotropical bee Xylocopa caffra.
Basionym: Starmerella xylocopis Gouliam., Dimitrov, M. Groenew., M.T. Sm. & Boekhout, Persoonia 46: 507. 2021.
Grinbergsozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857885
Etymology: the genus is named in honor of J. Grinbergs for his contribution to the study of yeast biodiversity, especially of the Valdivian rain forests in Chile.
Type species: Grinbergsozyma bentonensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida bentonensis, which is closely related to the genera Deakozyma and Limtongella (Figs. 1–2). Member of the Trichomonascaceae (Dipodascales, Dipodascomycetes). The genus is mainly circumscribed by phylogenomic analysis (Fig. 1) and rDNA phylogenetic analysis (Fig. 2), and the presence of genus-specific protein families OG0015427, OG0015488 and OG0019665 (Table 6).
Sexual reproduction not known. Colonies are white, semi-glistening and butyrous. Budding is multilateral. Pseudohyphae are present with elongate blastoconidia, but true hyphae are not observed.
Note: Grinbergsozyma bentonensis differs from species of the closely related genera Deakozyma and Limtongella by growth on 10% NaCl+5% glucose medium (Table S3). Our D1/D2 LSU analysis showed that at least three potential new species belong to Grinbergsozyma (Fig. S5, Table S2).
Species accepted:
Grinbergsozyma bentonensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857886
Holotype: NRRL YB-2364, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Illinois, Benton, isolated from apple cider, October 1950.
Basionym: Candida bentonensis Kurtzman, Antonie van Leeuwenhoek 88: 125. 2005.
Limtongella Sakpunt., Angchuan, Boontham, Khunnamw., Boonmak & Srisuk ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858217
For a description: see Sakpuntoon et al., Microorganisms 8 (27): 11. 2019.
Synonym: Limtongella Sakpunt., Angchuan, Boontham, Khunnamw., Boonmak & Srisuk, Microorganisms 8 (27): 11. 2019. Nom. inval., Art. F.5.1 (Shenzhen), the identifier of a recognized repository was not cited.
Type species: Limtongella siamensis Sakpuntoon, Angchuan, Boontham, Khunnamw., Boonmak & Srisuk ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
Species accepted:
Limtongella incommunis (Y. Ohara, Nonom. & M. Yamaz.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858218
Holotype: CBS 5604, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. JAPAN, isolated from grape must.
Basionym: Candida incommunis Y. Ohara, Nonom. & M. Yamaz., J. Gen. Appl. Microbiol. 11: 274. 1965.
Limtongella siamensis Sakpunt., Angchuan, Boontham, Khunnamw., Boonmak & Srisuk ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858219
For a description: see Sakpuntoon et al., Microorganisms 8 (27): 11. 2019.
Holotype: NBRC 114140, preserved in a metabolically inactive state at NITE Biological Resource Center (NBRC), Tokyo, Japan. THAILAND, Kasetsart University, isolated from the grease trap.
Synonym: Limtongella siamensis Sakpunt., Angchuan, Boontham, Khunnamw., Boonmak & Srisuk, Microorganisms 8 (27): 11. 2019. Nom. inval., Art. 35.1 (Shenzhen), the genus name is invalid as an identifier of a recognized repository was not cited.
Validated taxa of Starmerella
Starmerella powellii Lachance ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857934
For a description: see Lachance et al., Int. J. Syst. Evol. Microbiol. 51: 1206. 2001.
Holotype: CBS 8795, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. COSTA RICA, isolated from the nitidulid beetle (Conotelus sp.).
Synonyms: Candida powellii Lachance, Int. J. Syst. Evol. Microbiol. 51 (3): 1206. 2001. Nom. inval., Art. 40.3 (Melbourne).
Starmerella powellii Lachance ex C.A. Rosa & Lachance, Int. J. Syst. Evol. Microbiol. 68: 1342. 2018. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified; published as a combination, but ‘basionym’ (Candida powellii) is invalid.
Tardiomyces Spruijtenburg, Meis & T. de Groot ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857918
For a description: see Spruijtenburg et al., Infection 52: 1808. 2024.
Synonym: Tardiomyces Spruijtenburg, Meis & T. de Groot, Infection 52: 1808. 2024. Nom. inval., Art. F.5.1 (Shenzhen), the identifier of a recognized repository was not cited.
Type species: Tardiomyces depauwii Spruijtenburg, Meis & T. de Groot ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
Species accepted:
Tardiomyces blankii (H.R. Buckley & Uden) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857915
Holotype: CBS 1898, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. CANADA, isolated from blood of a mink (Putorius vison).
Basionym: Candida blankii H.R. Buckley & Uden, Mycopathol. Mycol. Appl. 36: 259. 1968.
Synonym: Tardiomyces blankii (H.R. Buckley & Uden) Spruijtenburg, Meis & T. de Groot, Infection 52: 1809. 2024. Nom. inval., Arts 35.1, 41.5, F.5.1 (Shenzhen), the identifier of a recognized repository was not cited.
Tardiomyces depauwii Spruijtenburg, Meis & T. de Groot ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857921
For a description: see Spruijtenburg et al., Infection 52: 1808. 2024.
Holotype: CBS 18495, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. UNITED KINGDOM, isolated from human blood.
Synonym: Tardiomyces depauwii Spruijtenburg, Meis & T. de Groot, Infection 52: 1808. 2024. Nom. inval., Art. 35.1, F.5.1 (Shenzhen), the genus name is invalid and the identifier of a recognized repository was not cited.
Tardiomyces digboiensis (G.S. Prasad, Mayilraj, Sood & Ban. Lal) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857923
Holotype: CBS 9800, preserved in a metabolically inactive state in the Westerdijk Fungal Biodiversity Institute (CBS), Utrecht, The Netherlands. INDIA, Assam, Digboi, isolated from acid tar sludge-contaminated soil from the Digboi oil refinery.
Basionym: Candida digboiensis G.S. Prasad, Mayilraj, Sood & Ban. Lal, Int. J. Syst. Evol. Microbiol. 55: 968. 2005.
Synonym: Tardiomyces digboiensis (G.S. Prasad, Mayilraj, Sood & Ban. Lal) Spruijtenburg, Meis & T. de Groot, Infection 52: 1809. 2024. Nom. inval., Arts 35.1, F.5.1 (Shenzhen), the genus name is invalid and the identifier of a recognized repository was not cited.
Westerdijkia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857927
Etymology: the genus is named in honor of Johanna Westerdijk for her contribution to plant pathology, fungal collections, and her role as a leading female scientist and the first female professor in the Netherlands.
Type species: Westerdijkia sungouii Q.M. Wang, B. James, K. Sylvester & Hittinger ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida sungouii, which occurs in a separate branch from the genera Diddensiella and Spencermartinsiella (Fig. 3). Member of the Trichomonascaceae (Dipodascales, Dipodascomycetes). The genus is mainly circumscribed by rDNA phylogenetic analysis.
No sexual stages are observed. Colonies are cream, butyrous, smooth. Budding is multilateral. Hyphae and pseudohyphae are not observed.
Note: Westerdijkia sungouii can be distinguished from species of the closely related genera Diddensiella and Spencermartinsiella by the absence of assimilation of ethanol (Table S3). Our D1/D2 LSU analysis showed Candida sp. BG02-7-18-018A-2-2 (GenBank AY520408) represents a potential new species of Westerdijkia (Fig. S6, Table S2).
Species accepted:
Westerdijkia sungouii Q.M. Wang, B. James, K. Sylvester & Hittinger ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857929
For a description: see Sylvester et al., FEMS Yeast Res. 15: 10. 2015.
Holotype: CBS 13907, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, Wisconsin, isolated from white slime, September 2011.
Synonym: Candida sungouii Q.M. Wang, B. James, K. Sylvester & Hittinger, FEMS Yeast Res. 15: 10. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Incertae sedis in Dipodascales (Dipodascomycetes)
Dengshuqunia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857935
Etymology: the genus is named after Prof. Shu-Qun Deng for his contribution to plant pathogenic fungi.
Type species: Dengshuqunia hispaniensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida hispaniensis, which is phylogenetically positioned in a separate long branch distinct from Yarrowia (Fig. 1). Member of the Dipodascomycetes. The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0013204, OG0015719 and OG0015720 (Table 6).
Sexual reproduction not known. Colonies are white, semi-glistening and butyrous. Pseudohyphae and true hyphae are not present. However, some tapered outgrowths bear blastoconidia. Fermentation is absent.
Note: Dengshuqunia can be distinguished from its closely related genus Yarrowia by its low genome size (about 11 Mb) and GC content (41.69%), whereas all species of Yarrowia have a genome size of about 20 Mb (17% higher than the genome size of Dengshuqunia) and higher GC contents (43.46–50.89%). Our ITS+D1/D2 LSU and D1/D2 LSU analyses showed that four potential new species belong to Dengshuqunia (Fig. 5 and Fig. S8, Table S2).
Species accepted:
Dengshuqunia hispaniensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857936
Holotype: NRRL Y-5580, preserved in a metabolically inactive state at the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. SPAIN, isolated from the larva of longhorn beetle (Spondylis buprestoides) in conifer forest, April 1960.
Basionym: Candida hispaniensis Kurtzman, Antonie van Leeuwenhoek 88: 128. 2005.
Validated taxa in Yarrowia
Yarrowia brassicae F.L. Hui & K.F. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857939
For a description: see Liu et al., Int. J. Syst. Evol. Microbiol. 68: 2026. 2018.
Holotype: CICC 33263, preserved in a metabolically inactive state at the China Centre of Industrial Culture Collection (CICC), Beijing, China. CHINA, Henan province, Nanyang, isolated from traditional Chinese sauerkraut.
Synonym: Yarrowia brassicae F.L. Hui & K.F. Liu, Int. J. Syst. Evol. Microbiol. 68: 2026. 2018. Nom. inval., Art. 40.7 (Shenzhen), more than one collection in which the type is conserved was specified.
Yarrowia divulgata Nagy, Niss, Dlauchy, Arneborg, D.S. Nielsen & G. Péter ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857940
For a description: see Nagy et al., Int. J. Syst. Evol. Microbiol. 63: 4821. 2013.
Holotype: CBS 11013, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. DENMARK, isolated from a bacon processing plant.
Synonym: Yarrowia divulgata Nagy, Niss, Dlauchy, Arneborg, D.S. Nielsen & G. Péter, Int. J. Syst. Evol. Microbiol. 63: 4821. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yarrowia keelungensis Chin F. Chang & S.M. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857941
For a description: see Chang et al., Antonie van Leeuwenhoek 104: 1121. 2013.
Holotype: BCRC 23110, preserved in a metabolically inactive state at the Bioresource Collection and Research Center, Food Industry Research and Development Institute, Hsinchu, Taiwan, China. CHINA, Taiwan province, isolated from the sea-surface microlayer, 2006.
Synonym: Yarrowia keelungensis Chin F. Chang & S.M. Liu, Antonie van Leeuwenhoek 104: 1121. 2013. Nom. inval., Art. 40.1, 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yarrowia phangngaensis Limtong, Yongman., H. Kawas. & T. Seki ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB857942
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 58: 517. 2008.
Holotype: BCC 21231, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Phang-Nga Province, Mu Ko Ra-Ko Prathong National Park, isolated from water in a mangrove forest.
Synonyms: Candida phangngaensis Limtong, Yongman., H. Kawas. & T. Seki [as ‘phangngensis’], Int. J. Syst. Evol. Microbiol. 58: 517. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yarrowia phangngaensis (Limtong, Yongman., H. Kawas. & T. Seki) Gouliam., R.A. Dimitrov, M.T. Sm. & M. Groenew., Persoonia 39: 289. 2017. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified; published as a combination but 'basionym' (Candida phangngaensis) invalid.
New taxa in Pichiaceae (Pichiales, Pichiomycetes)
New combinations and validated taxa in Ogataea
Ogataea arabinofermentans (Kurtzman & Dien) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858229
Holotype: NRRL YB-2248, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Alaska, isolated from insect frass from dead Larix.
Basionym: Candida arabinofermentans Kurtzman & Dien, Antonie van Leeuwenhoek 74: 241. 1998.
Ogataea chonburiensis Limtong, Srisuk, Yongman., Yurim. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858010
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 58: 304. 2008.
Holotype: BCC 21227, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Chonburi province, isolated from the soil.
Synonym: Ogataea chonburiensis Limtong, Srisuk, Yongman., Yurim. & Nakase, Int. J. Syst. Evol. Microbiol. 58: 304. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea chumphonensis Limtong, Koowadj., Jindam. & Yongman. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858011
For a description: see Koowadjanakul et al., Antonie van Leeuwenhoek 100: 213. 2011.
Holotype: BCC 42667, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, isolated from the leaves of the tree.
Synonym: Candida chumphonensis Limtong, Koowadj., Jindam. & Yongman., Antonie van Leeuwenhoek 100: 213. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea deakii G. Péter, Dlauchy & Čadež ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858012
For a description: see Čadež et al., Int. J. Syst. Evol. Microbiol. 63: 3121. 2013.
Holotype: CBS 12735, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. HUNGARY, Pilis mountains, isolated brown rotten wood of beech (Fagus sylvatica), 2003.
Synonym: Ogataea deakii G. Péter, Dlauchy & Čadež, Int. J. Syst. Evol. Microbiol. 63: 3121. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea ganodermae F.Y. Bai & Z.H. Ji ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862204
For a description: see Ji & Bai, Int. J. Syst. Evol. Microbiol. 58: 1504. 2008.
Holotype: CGMCC 2.3435, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hunan province, Mangshan mountain, isolated from the a basidiocarp of Ganoderma sp., October 2006.
Synonym: Ogataea ganodermae F.Y. Bai & Z.H. Ji, Int. J. Syst. Evol. Microbiol. 58: 1504. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea histrianica Čadež & G. Péter ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858013
For a description: see Čadež et al., Int. J. Syst. Evol. Microbiol. 63: 3120. 2013.
Holotype: CBS 12779, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SLOVENI, Ankaran, isolated from olive oil sediment of mixed varieties, 2011.
Synonym: Ogataea histrianica Čadež & G. Péter, Int. J. Syst. Evol. Microbiol. 63: 3120. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea kanchanaburiensis Limtong, Kaewwich. & M. Groenew. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862206
For a description: see Limtong et al., Antonie van Leeuwenhoek 103(3): 554. 2012.
Holotype: BCC 47626, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Kanchanaburi province, Mueang District, Wang Dong sub district, isolated from the leaves of the mango tree (Mangifera indica).
Synonym: Ogataea kanchanaburiensis Limtong, Kaewwich. & M. Groenew., Antonie van Leeuwenhoek 103(3): 554. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea kolombanensis Čadež & G. Péter ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858014.
For a description: see Čadež et al., Int. J. Syst. Evol. Microbiol. 63: 3119. 2013.
Holotype: CBS 12778, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SLOVENI, Koper, isolated from olive oil sediment of the ‘Istrska Belica’ variety, 2009.
Synonym: Ogataea kolombanensis Čadež & G. Péter, Int. J. Syst. Evol. Microbiol. 63: 3119. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea krabiensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858230
For a description: see Limtong et al., J. Gen. Appl. Microbiol. 50: 123. 2004.
Holotype: TISTR 5820, preserved in a metabolically inactive state at TISTR Culture Collection Bangkok MIRCEN, Thailand Institute of Science and Technological Research, Bangkok 10900, Thailand. THAILAND, Krabi province, isolated from soil, 2000.
Synonym: Candida krabiensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato, J. Gen. Appl. Microbiol. 50: 123. 2004. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea maris (Uden & Zobell) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858231
Holotype: CBS 5151, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. AUSTRALIA, Torres Strait, isolated from seawater.
Basionym: Torulopsis maris Uden & Zobell, Antonie van Leeuwenhoek 28: 281. 1962.
Synonym: Candida maris (Uden & Zobell) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Ogataea mattraensis Limtong, Koowadj., Jindam. & Yongman. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858015
For a description: see Koowadjanakul et al., Antonie van Leeuwenhoek 100: 214. 2011.
Holotype: BCC 42668, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, isolated from leaves of tree.
Synonym: Candida mattraensis Limtong, Koowadj., Jindam. & Yongman. [as ‘mattranensis’], Antonie van Leeuwenhoek 100: 214. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea nakhonphanomensis Limtong, Srisuk, Yongman., Yurim. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858016
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 58: 305. 2008.
Holotype: BCC 21228, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Nakhon-Phanom province, isolated from a tree exudate.
Synonym: Ogataea nakhonphanomensis Limtong, Srisuk, Yongman., Yurim. & Nakase, Int. J. Syst. Evol. Microbiol. 58: 305. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea nemodendra (van der Walt, Klift & D.B. Scott) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858232
Holotype: CBS 6280, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from tunnels of Xyleborus aemulus in Rapanea melanophloeos.
Basionym: Torulopsis nemodendra van der Walt, Klift & D.B. Scott, Antonie van Leeuwenhoek 37: 468. 1971.
Synonym: Candida nemodendra (van der Walt, Klift & D.B. Scott) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Ogataea ortonii Lachance ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858233
For a description: see Lachance et al., FEMS Yeast Res. 1: 90. 2001.
Holotype: CBS 8843, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. COSTA RICA, isolated from the flux of old fustic tree (Maclura tinctoria).
Synonym: Candida ortonii Lachance, FEMS Yeast Res. 1: 90. 2001. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea ovalis (Kumam. & M. Yamam.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858234
Holotype: CBS 7298, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from soil.
Basionym: Candida ovalis Kumam. & M. Yamam., Trans. Mycol. Soc. Japan 27: 391. 1987.
Ogataea phyllophila Koowadj., Jindam., Yongman. & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858017
For a description: see Koowadjanakul et al., Antonie van Leeuwenhoek 100: 211. 2011.
Holotype: BCC 42666, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Chumphon province, Chumphon Islands National Park, isolated from the leaves of a tree, May 2009.
Synonym: Ogataea phyllophila Koowadj., Jindam., Yongman. & Limtong, Antonie van Leeuwenhoek 100: 211. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea piceae (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858235
Holotype: CBS 8701, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, Alaska, isolated from insect frass in Sitka spruce (Picea sitchensis).
Basionym: Candida piceae Kurtzman, Can. J. Microbiol. 46: 54. 2000.
Ogataea pinus (Lodder & Kreger-van Rij) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858237
Holotype: CBS 970, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SWEDEN, isolated from the water-logged heart of a pine tree.
Basionym: Torulopsis pinus Lodder & Kreger-van Rij, Yeasts, a taxonomic study, [Edn 1] (Amsterdam): 671. 1952.
Synonyms: Candida pinus (Lodder & Kreger-van Rij) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Paratorulopsis pinus (Lodder & Kreger-van Rij) E.K. Novák & Zsolt, Acta Bot. Acad. Sci. Hung. 7: 141. 1961.
Ogataea rishiriensis (Nakase, Imanishi, Ninomiya & M. Takash.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858238
Holotype: NBRC 105027, preserved in a metabolically inactive state at NITE Biological Resource Center (NBRC), Tokyo, Japan. JAPAN, Hokkaido, isolated from the soil in a pine forest.
Basionym: Candida rishiriensis Nakase, Imanishi, Ninomiya & M. Takash. [as 'rishirensis'], J. Gen. Appl. Microbiol. 56: 171. 2010.
Ogataea siamensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858018
For a description: see Limtong et al., J. Gen. Appl. Microbiol. 50: 121. 2004.
Holotype: JCM 12264, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. THAILAND, Kanchanaburi province, isolated from the flower of Ervatamia coronaria, 2001.
Synonyms: Pichia siamensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato, J. Gen. Appl. Microbiol. 50: 121. 2004. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea siamensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato ex Limtong, Srisuk, Yongman., Yurim. & Nakase, Int. J. Syst. Evol. Microbiol. 58: 306. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea sithepensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858239
For a description: see Limtong et al., J. Gen. Appl. Microbiol. 50: 125. 2004.
Holotype: JCM 12265, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. THAILAND, Petchabun province, Sithep Historical Park, isolated from soil, 2000.
Synonym: Candida sithepensis Limtong, Srisuk, Yongman., H. Kawas., Yurim., Nakase & N. Kato, J. Gen. Appl. Microbiol. 50: 125. 2004. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea sonorensis (M.W. Mill., Phaff, M. Miranda, Heed & Starmer) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858240
Holotype: CBS 6792, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. MEXICO, isolated from the tissue of the organ-pipe cactus (Lemaireocereus thurberi).
Basionym: Torulopsis sonorensis M.W. Mill., Phaff, M. Miranda, Heed & Starmer, Int. J. Syst. Bacteriol. 26: 88. 1976.
Synonym: Candida sonorensis (M.W. Mill., Phaff, M. Miranda, Heed & Starmer) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Ogataea thermomethanolica Limtong, Srisuk, Yongman., Yurim., Nakase & N. Kato ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858019
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 55: 2226. 2005.
Holotype: BCC 16875, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Pathalung province, isolated from soil.
Synonyms: Pichia thermomethanolica Limtong, Srisuk, Yongman., Yurim., Nakase & N. Kato, Int. J. Syst. Evol. Microbiol. 55: 2226. 2005. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea thermomethanolica Limtong, Srisuk, Yongman., Yurim., Nakase & N. Kato ex Limtong, Srisuk, Yongman., Yurim. & Nakase, Int. J. Syst. Evol. Microbiol. 58: 306. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea thermophila K.S. Shin, Y.K. Shin, J.H. Yoon & Y.H. Park ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858020
For a description: see Shin et al., Int. J. Syst. Evol. Microbiol. 51: 2168. 2001.
Holotype: JCM 10994, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. SOUTH KOREA, isolated from china clay.
Synonyms: Candida thermophila K.S. Shin, Y.K. Shin, J.H. Yoon & Y.H. Park, Int. J. Syst. Evol. Microbiol. 51: 2168. 2001. Nom. inval., Art. 40.7(Melbourne), more than one collection in which the type is conserved was specified.
Ogataea thermophila G. Péter, Tornai-Leh., K.S. Shin & Dlauchy, FEMS Yeast Res. 7: 495. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Ogataea xyloterini (S.O. Suh & J.J. Zhou) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858021
Holotype: ATCC 62898, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, Wisconsin, Palmyra, isolated from Xyloterinus politus.
Basionym: Candida xyloterini S.O. Suh & J.J. Zhou, Int. J. Syst. Evol. Microbiol. 60: 1704. 2010.
New combination and validated taxa in Pichia
Pichia awuae D.S. Nielsen, M. Jakobsen & Jespersen ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858022
For a description: see Nielsen et al., Int. J. Syst. Evol. Microbiol. 60: 1464. 2010.
Holotype: CBS 11011, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. GHANA, isolated from fermenting cocoa beans.
Synonym: Candida awuae D.S. Nielsen, M. Jakobsen & Jespersen [as ‘awuaii’], Int. J. Syst. Evol. Microbiol. 60: 1464. 2010. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia bruneiensis Sipiczki ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858023
For a description: see Sipiczki, Int. J. Syst. Evol. Microbiol. 62: 3103. 2012.
Holotype: CBS 12611, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRUNEI, Borneo, isolated from the flowers of Hibiscus.
Synonym: Pichia bruneiensis Sipiczki, Int. J. Syst. Evol. Microbiol. 62: 3103. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia chibodasensis R. Kobay., A. Kanti & H. Kawas. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858024
For a description: see Kobayashi et al., Int. J. Syst. Evol. Microbiol. 67: 1025. 2017.
Holotype: NBRC 111569, preserved in a metabolically inactive state at the Biological Resource Center, NITE (NBRC), Tokyo, Japan. INDONESIA, West Java, Chibodas Botanic Garden, isolated from soil and decayed wood, December 2014.
Synonym: Pichia chibodasensis R. Kobay., A. Kanti & H. Kawas., Int. J. Syst. Evol. Microbiol. 67: 1025. 2017. Nom. inval., Art. 40.7 (Shenzhen), more than one collection in which the type is conserved was specified.
Pichia dushanensis F.L. Hui, Y.C. Ren, S.T. Liu & Ying Li ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858025
For a description: see Ren et al., Int. J. Syst. Evol. Microbiol. 65: 2877. 2015.
Holotype: CBS 13912, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Dushan Forest Park in Nanyang, isolated from the gut of insect larvae.
Synonym: Pichia dushanensis F.L. Hui, Y.C. Ren, S.T. Liu & Ying Li, Int. J. Syst. Evol. Microbiol. 65: 2877. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia insulana Ganter, Cardinali & Boundy-Mills ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858028
For a description: see Ganter et al., Int. J. Syst. Evol. Microbiol. 60: 1003. 2010.
Holotype: CBS 11169, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CURAÇAO, isolated from a rot pocket in a Cereus repandus.
Synonym: Pichia insulana Ganter, Cardinali & Boundy-Mills, Int. J. Syst. Evol. Microbiol. 60: 1003. 2010. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia phayaonensis Limtong, Nitiyon, Kaewwich., Jindam., Am-In & Yongman. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858029
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 62: 2789. 2012.
Holotype: BCC 47634, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Phayao province, isolated from soil, December 2008.
Synonyms: Candida phayaonensis Limtong, Nitiyon, Kaewwich., Jindam., Am-In & Yongman., Int. J. Syst. Evol. Microbiol. 62: 2789. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia phayaonensis Limtong, Nitiyon, Kaewwich., Jindam., Am-In & Yongman. ex H.Y. Zhu, L.C. Guo & F.Y. Bai, Int. J. Syst. Evol. Microbiol. 74: 006306, 7. 2024. Nom. inval., Art. 40.1 (Shenzhen), published as a combination but ‘basionym’ (Candida phayaonensis) invalid.
Pichia sorboxylosa (Nakase) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858241
Holotype: CBS 6378, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from pineapple (Ananas comosus).
Basionym: Candida sorboxylosa Nakase, J. Gen. Appl. Microbiol. 17: 392. 1971.
Pichia thaimueangensis Limtong, Yongman., H. Kawas. & T. Seki ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858030
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 57: 651. 2007.
Holotype: BCC 21229, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Phang-Nga province, Kho Lumpee-Haad Thaimueang National Park, isolated from water in a mangrove forest.
Synonyms: Candida thaimueangensis Limtong, Yongman., H. Kawas. & T. Seki, Int. J. Syst. Evol. Microbiol. 57: 651. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Pichia thaimueangensis Limtong, Yongman., H. Kawas. & T. Seki ex H.Y. Zhu, L.C. Guo & F.Y. Bai, Int. J. Syst. Evol. Microbiol. 74: 006306, 7. 2024. Nom. inval., Art. 40.1 (Shenzhen), published as a combination, but ‘basionym’ (Candida thaimueangensis) is invalid.
Ramirezia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858227
Etymology: the genus is named in honor of C. Ramírez, who described the type species of this genus.
Type species: Ramirezia boidinii (C. Ramírez) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida boidinii, which is in a separate long branch from the other genera in Pichiales (Fig. 6). Member of the Pichiaceae (Pichiales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0009746, OG0009748 and OG0009751 (Table 6).
Sexual reproduction not known. Colonies are white to cream, butyrous. Budding is multilateral. Pseudohyphae with blastoconidia are present. Septate hyphae may be present. Coenzyme Q-7 is formed.
Note: Our ITS analysis showed that two potential new species belong to Ramirezia (Fig. S10, Table S2).
Species accepted:
Ramirezia boidinii (C. Ramírez) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858228
Holotype: CBS 2428, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SPAIN, isolated from tanning fluid.
Basionym: Candida boidinii C. Ramírez [as 'boidini'], Microbiol. Esp. 6: 251. 1953.
Wenyingozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB857943
Etymology: the genus is named in honor of Wen-Ying Zhuang for her contributions to the taxonomy of Ascomycota, especially of Pezizomycetes.
Type species: Wenyingozyma ramenticola (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for species in the Ogataea clade 2 (i.e., the Ogataea naganishii subclade, the Candida methanosorbosa subclade, the Ogataea ramenticola subclade and the single-species Ogataea methylovora) which is in a separate clade from the Ogataea clade 1 (Fig. 6). Member of the Pichiaceae (Pichiales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0006088, OG0006166 and OG0006167 (Table 6).
Both teleomorphic and anamorphic species occur. Asci originate from conjugation between a cell and its bud or between independent cells, and produce one to four hat-shaped ascospores. Colonies are cream to white, butyrous. Budding is multilateral. Neither pseudohyphae nor true hyphae are formed. Where known, coenzyme Q-7 is formed.
Species accepted:
Wenyingozyma methanosorbosa (S. Abe & Yokote) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858220
Holotype: CBS 7029, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from soil.
Basionym: Torulopsis methanosorbosa S. Abe & Yokote, J. Ferment. Technol. 52: 203. 1974.
Synonym: Candida methanosorbosa (S. Abe & Yokote) J.A. Barnett, R.W. Payne & Yarrow, Yeasts: Characteristics and Identification (Cambridge): 732. 1983.
Wenyingozyma methylovora (Kumam. & Seriu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858221
Holotype: CBS 7300, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from a rotted tree.
Basionym: Pichia methylovora Kumam. & Seriu, Trans. Mycol. Soc. Japan 27: 394. 1987.
Synonym: Ogataea methylovora (Kumam. & Seriu) Kurtzman & Robnett [as 'methylivora'], FEMS Yeast Res. 10: 357. 2010.
Wenyingozyma naganishii (K. Kodama) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858222
Holotype: CBS 6429, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from exudate from a camellia (Camellia japonica).
Basionym: Pichia naganishii K. Kodama, J. Ferment. Technol. 52: 9. 1974.
Synonym: Ogataea naganishii (K. Kodama) Kurtzman & Robnett, FEMS Yeast Res. 10: 357. 2010.
Wenyingozyma nanaspora (Saëz & Rodr. Mir.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857944
Holotype: CBS 7200, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. FRANCE, isolated from hair of capuchin monkey (Cebus apella).
Basionym: Candida nanaspora Saëz & Rodr. Mir., Bull. Trimestriel Soc. Mycol. France 104: 214. 1988.
Wenyingozyma neixiangensis (F.L. Hui & Jun Zheng) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857945
Holotype: CICC 33166, preserved in a metabolically inactive state at the China Centre of Industrial Culture Collection (CICC), Beijing, China. CHINA, Henan province, Baotianman Nature Reserve, isolated from rotting wood, September 2016.
Basionym: Ogataea neixiangensis F.L. Hui & Jun Zheng, Int. J. Syst. Evol. Microbiol. 67: 3041. 2017.
Wenyingozyma nitratophila (Shifrine & Phaff) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857946
Holotype: CBS 2027, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, California, isolated from bark beetle (Dendroctonus monticolae).
Basionym: Torulopsis nitratophila Shifrine & Phaff, Mycologia 48: 48. 1956.
Synonym: Candida nitratophila (Shifrine & Phaff) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Wenyingozyma ramenticola (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB857947
Holotype: NRRL YB-1985, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Florida, Wilma, isolated from frass in a tunnel of a beetle from Pinus palustris.
Basionym: Pichia ramenticola Kurtzman, Canad. J. Microbiol. 46: 51. 2000.
Synonym: Ogataea ramenticola (Kurtzman) Kurtzman & Robnett, FEMS Yeast Res. 10: 357. 2010.
Wenyingozyma succiphila (J.D. Lee & Komag.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858223
Holotype: CBS 8003, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from sap of a peach tree (Amygdalus persica).
Basionym: Candida succiphila J.D. Lee & Komag., Int. J. Syst. Bacteriol. 30: 518. 1980.
Synonym: Candida cellulolytica Nakase, M. Suzuki, M. Takash. & Hamam., J. Gen. Appl. Microbiol. 40: 521. 1994.
Note: C. succiphila and C. cellulolytica have the same LSU D1/D2 sequences and near 100% genome sequence similarity; therefore, C. cellulolytica is assigned as a synonym of C. succiphila.
Wenyingozyma suzukii (G. Péter, Tornai-Leh., Fülöp & Dlauchy) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858008
Holotype: CBS 9253, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, isolated from the bark of an unidentified tree.
Basionym: Candida suzukii G. Péter, Tornai-Leh., Fülöp & Dlauchy, Antonie van Leeuwenhoek 84: 149. 2003.
Wenyingozyma wangdongensis Limtong, Kaewwich. & M. Groenew. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858009
For a description: see Limtong et al., Antonie van Leeuwenhoek 103: 555. 2012.
Holotype: BCC 42667, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Kanchanaburi province, isolated from leaves of wine grape (Vitis vinifera), July 2009.
Synonym: Ogataea wangdongensis Limtong, Kaewwich. & M. Groenew., Antonie van Leeuwenhoek 103: 555. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Xiuguozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858224
Etymology: the genus is named in honor of Xiu-Guo Zhang for his contribution to fungal taxonomy, especially of Hyphomycetes.
Type species: Xiuguozyma insectalens (D.B. Scott, van der Walt & Klift) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species of the Candida insectalens clade, which is in a separate clade from Allodekkera and Brettanomyces in the phylogenomic tree (Fig. 6). Member of Pichiaceae (Pichiales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0009801, OG0009803 and OG001190 (Table 6).
Sexual reproduction not known. Colonies are white to cream, butyrous, smooth, convex. Budding is multilateral. Fermentation is absent. Coenzyme Q-9 is formed.
Note: Xiuguozyma species can be distinguished from species of the closely related genus Brettanomyces by the lack of growth with 0.01% cycloheximide (Table S3). Ecologically, the new genus Xiuguozyma appears to be related to insects.
Species accepted:
Xiuguozyma insectalens (D.B. Scott, van der Walt & Klift) Q.M. Wang, Boekhout, A.M. Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858225
Holotype: CBS 6036, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from tunnels of Crossotarsus extrernedentatus in Ficus sycomorus.
Basionym: Torulopsis insectalens D.B. Scott, van der Walt & Klift, Antonie van Leeuwenhoek 37: 467. 1971.
Synonym: Candida insectalens (D.B. Scott, van der Walt & Klift) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Xiuguozyma silvatica (van der Walt, Klift & D.B. Scott) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858226
Holotype: CBS 6277, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from tunnels of Crossotarsus extrernedentatus in Ficus sycomorus.
Basionym: Torulopsis silvatica van der Walt, Klift & D.B. Scott, Antonie van Leeuwenhoek 37: 465. 1971.
Synonym: Candida silvatica (van der Walt, Klift & D.B. Scott) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
New taxa, reinstated and emended genera in Debaryomycetaceae (Serinales, Pichiomycetes)
Candida Berkhout, nom. cons., emend. Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai
Type species: Candida vulgaris Berkhout.
This genus is emended to accommodate the species in the Candida sensu stricto clade including Candida albicans, Candida dubliniensis and Candida tropicalis (Fig. 9). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The emended genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0006430 and OG0006359 (Table 6).
Both teleomorphic and anamorphic species occur in this genus. Asci are unconjugated asci with one or two ellipsoidal to elongate ascospores. Colonies are white to cream, smooth and butyrous. Budding is multilateral. Pseudohyphae and septate hyphae are present or not. Where known, coenzyme Q-9 is formed.
Species accepted:
Candida albicans (C.P. Robin) Berkhout, Schimmelgesl. Monilia: 44. 1923.
MycoBank: MB256187
Synonyms: Oidium albicans C.P. Robin, Hist. Nat. Veg. Paras.: 488. 1853.
Dematium albicans (C.P. Robin) E. Laurent, Bull. Soc. Belge Microscop. 16: 14. 1889.
Monilia albicans (C.P. Robin) Zopf, Pilze: 478. 1890. Nom. illeg., Art. 53.1, non Monilia albicans Pers. 1822.
Myceloblastanon albicans (C.P. Robin) M. Ota, Jap. J. Derm. Urol. 27: 170. 1927.
Mycotorula albicans (C.P. Robin) Langeron & Talice, Ann. Parasitol. Humaine Comp. 10: 44. 1932.
Parasaccharomyces albicans (C.P. Robin) Mello & L.G. Fern., Arq. Hig. Patol. Exot. 6: 207-316. 1918.
Procandida albicans (C.P. Robin) E.K. Novák & Zsolt, Acta Bot. Acad. Sci. Hung. 7: 133. 1961. Nom. inval., Art. 41.5 (Melbourne).
Saccharomyces albicans (C.P. Robin) Reess, Sitzungsber. Phys.-Med. Soc. Erlangen 9: 195. 1877.
Syringospora albicans (C.P. Robin) C.W. Dodge, Medic. Mycol.: 274. 1935.
Candida aquae-textoris Vallini, Frassinetti & Scorzetti [as 'aquaetextoris'], Int. J. Syst. Bacteriol. 47: 336. 1997.
MycoBank: MB436997
Candida baotianmanensis Y.C. Ren, L.L. Xu, Lin Zhang & F.L. Hui ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858036
For a description: see Ren et al., Int. J. Syst. Evol. Microbiol. 65: 3583. 2015.
Holotype: CBS 13915, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Baotianman Mountain in Nanyang, isolated from the gut of Nitidula carnaria.
Synonym: Candida baotianmanensis Y.C. Ren, L.L. Xu, Lin Zhang & F.L. Hui, Int. J. Syst. Evol. Microbiol. 65: 3583. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Candida buenavistaensis S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 96. 2008.
MycoBank: MB505577
Candida dubliniensis D.J. Sullivan, Western., K.A. Haynes, Dés.E. Benn. & D.C. Coleman [as 'dublinionensis'], Microbiology (Reading) 141: 1519. 1995.
MycoBank: MB254786
Candida frijolesensis S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 96. 2008.
MycoBank: MB505950
Candida gigantensis S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 92. 2008.
MycoBank: MB505574
Candida kantuleensis Nitiyon, Khunnamw., Lertwatt. & Limtong, Int. J. Syst. Evol. Microbiol. 68: 2316. 2018.
MycoBank: MB824179
Candida labiduridarum S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 97. 2008.
MycoBank: MB505578
Candida maltosa Komag., Nakase & Katsuya, J. Gen. Appl. Microbiol., 10: 327. 1964.
MycoBank: MB327445
Candida neerlandica Kurtzman, Robnett & Yarrow, Antonie van Leeuwenhoek 80: 81. 2001.
MycoBank: MB456708
Candida pellucidi Glushakova, Tomashevskaya & Kachalkin [as 'pellucida'], Persoonia 44: 369. 2020.
MycoBank: MB836057
Candida pseudoviswanathii Y.C. Ren, L.L. Xu, Lin Zhang & F.L. Hui ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858037
For a description: see Ren et al., Int. J. Syst. Evol. Microbiol. 65: 3584. 2015.
Holotype: CBS 13916, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Nanyang, isolated from the gut of a beetle larva (Dorcus curvidens).
Synonym: Candida pseudoviswanathii Y.C. Ren, L.L. Xu, Lin Zhang & F.L. Hui, Int. J. Syst. Evol. Microbiol. 65: 3584. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Candida sanyaensis F.L. Hui, Q.H. Niu, T. Ke & Ying X. Li ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858038
For a description: see Hui et al., Antonie van Leeuwenhoek 103: 50. 2013.
Holotype: CBS 12637, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Hainan province, Sanya, isolated from soil.
Synonym: Candida sanyaensis F.L. Hui, Q.H. Niu, T. Ke & Ying X. Li, Antonie van Leeuwenhoek 103: 50. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Candida saraburiensis Nitiyon, Boonmak, Am-In, Jindam., H. Kawas., Yongman. & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858039
For a description: see Nitiyon et al., Int. J. Syst. Evol. Microbiol. 61: 465. 2011.
Holotype: CBS 11696, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Saraburi province, isolated from decaying corncobs.
Synonym: Candida saraburiensis Nitiyon, Boonmak, Am-In, Jindam., H. Kawas., Yongman. & Limtong, Int. J. Syst. Evol. Microbiol. 61: 465. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Candida sojae Nakase, M. Suzuki, M. Takash., Yozo Miyak., Kagaya, Fukaz. & Komag., J. Gen. Appl. Microbiol. 40: 163. 1994.
MycoBank: MB363475
Candida tetrigidarum S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 97. 2008.
MycoBank: MB505573
Candida tropicalis (Castell.) Berkhout, Schimmelgesl. Monilia: 44. 1923.
MycoBank: MB280770
Synonyms: Oidium tropicale Castell., Philipp. J. Sci., B, Med. Sci. 5: 202. 1910.
Candida vulgaris Berkhout, Schimmelgesl. Monilia: 42. 1923 (TYPE of the genus Candida).
Candida viswanathii R.S. Sandhu & H.S. Randhawa, Mycopath. Mycol. Appl. 18: 179. 1962.
MycoBank: MB294056
Synonym: Candida viswanathii T.S. Viswan. & H.S. Randhawa, Sci. Cult. 25: 86. 1959. Nom. inval., Art. 39.1 (Melbourne).
Candida yunnanensis F.L. Hui & L.N. Huang, Int. J. Syst. Evol. Microbiol. 69: 2778. 2019.
MycoBank: MB830819
Chernovozyma Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, gen. nov.
MycoBank: MB858242
Etymology: the genus is named in honor of I. Yu. Chernov for his contribution to yeast taxonomy and ecology.
Type species: Chernovozyma aurita (A.V. Polyak. & Chernov) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai.
This genus is proposed for the Candida aurita clade, which occurs as a separate lineage closely related to Kurtzmaniella and the Candida railenensis clade in Debaryomycetaceae (Serinales, Pichiomycetes) (Fig. 12). The genus is mainly circumscribed by the phylogenomic and rDNA phylogenetic analyses, and the presence of genus-specific protein families OG0009945, OG0009948 and OG0010921 (Table 6).
Sexual reproduction not known. Colonies are white, cream, butyrous. Budding is multilateral. Pseudohyphae is produced, but hyphae are not present.
Note: Chernovozyma spp. are different from those of the closely related genus Dujonia by the lack of growth on 0.1% cycloheximide (Table S3).
Species accepted:
Chernovozyma aurita (A.V. Polyak. & Chernov) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858243
Holotype: CBS 9724, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SIBERIA, isolated from the soil of a peat bog.
Basionym: Candida aurita A.V. Polyak. & Chernov, Microbiology, Moscow 71: 332. 2002.
Chernovozyma palmyrensis (S.O. Suh & J.J. Zhou) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858244
Holotype: ATCC 62899, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, Wisconsin, Palmyra, isolated from Xyloterinus politus.
Basionym: Candida palmyrensis S.O. Suh & J.J. Zhou, Int. J. Syst. Evol. Microbiol. 60: 1706. 2010.
Chernovozyma sophiae-reginae (C. Ramírez & A.E. González) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858245
Holotype: CBS 8175, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHILE, isolated from rotten wood of Chilean laurel (Laurelia sempervirens).
Basionym: Candida sophiae-reginae C. Ramírez & A.E. González, Mycopathologia 88: 93. 1984.
New combinations and validated taxa in Debaryomyces
Debaryomyces psychrophilus (Goto, Sugiy. & Iizuka) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858088
Holotype: CBS 5956, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ANTARCTICA, isolated from the dung of Spheniscidae.
Basionym: Torulopsis psychrophila Goto, Sugiy. & Iizuka, Mycologia 61: 761. 1969.
Synonym: Candida psychrophila (Goto, Sugiy. & Iizuka) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Debaryomyces renaianus C.F. Lee & Y.R. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858089
For a description: see Lee et al., Bot. Studies (Taipei) 50: 326. 2009.
Holotype: CBS 10891, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Nantou, Renai, isolated from forest soil.
Synonym: Debaryomyces renaii C.F. Lee & Y.R. Liu, Bot. Studies (Taipei) 50: 326. 2009. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Debaryomyces vindobonensis Lopandić, Rents. & Prillinger ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858090
For a description: see Lopandić et al., J. Gen. Appl. Microbiol. 59: 54. 2013.
Holotype: CBS 11666, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. AUSTRIA, Vienna, isolated from the wastewater treatment plant.
Synonym: Debaryomyces vindobonensis Lopandić, Rents. & Prillinger, J. Gen. Appl. Microbiol. 59: 54. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Dujonia Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, gen. nov.
MycoBank: MB848190
Etymology: the genus is named in honor of Dr. Bernard Dujon for his contribution to comparative genomics and genetics of ascomycetous yeasts, and his leading role in the French Genolevures consortium.
Type species: Dujonia schatavii (Kock.-Krat. & Ondrush.) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai.
This genus is proposed for the Candida railenensis clade, which occurs as a separate lineage closely related to the genus Kurtzmaniella and the Candida aurita clade in Debaryomycetaceae (Figs. 12–13). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by the phylogenomics analysis, and the presence of genus-specific protein family OG0009139 (Table 6).
Sexual reproduction not known. Colonies are white, creamy, butyrous. Budding is multilateral. Hyphae are present or not. Pseudohyphae are produced.
Species accepted:
Dujonia boleticola (Nakase) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848192
Holotype: CBS 6420, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from the fruiting body of an earthstar mushroom (Astraeus hygrometricus).
Basionym: Candida boleticola Nakase, J. Gen. Appl. Microbiol. 17: 473. 1971.
Dujonia oleophila (Montrocher) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848193
Holotype: CBS 2219, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ITALY, isolated from olives (Olea europea).
Basionym: Candida oleophila Montrocher, Rev. Mycol. (Paris) 32: 73. 1967.
Dujonia railenensis (C. Ramírez & A.E. González) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848194
Holotype: CBS 8164, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHILE, isolated from the rotten trunk of southern beech (Nothofagus dombeyii).
Basionym: Candida railenensis C. Ramírez & A.E. González, Mycopathologia 88: 55. 1984.
Dujonia santamariae (Montrocher) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848195
Holotype: CBS 4515, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SPAIN, isolated from the sugar solution.
Basionym: Candida santamariae Montrocher, Rev. Mycol. (Paris) 32: 77. 1967.
Dujonia schatavii (Kock.-Krat. & Ondrush.) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848196
Holotype: CBS 6452, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CZECHOSLOVAKIA, isolated from the basidiocarp of a polypore (Fomitopsis pinicola).
Basionym: Torulopsis schatavii Kock.-Krat. & Ondrush., Biológia (Bratislava) 26: 483. 1971.
Synonym: Candida schatavii (Kock.-Krat. & Ondrush.) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Dujonia zeylanoides (Castell.) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848197
Holotype: CBS 619, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SRI LANKA, isolated from blastomycotic macroglossia.
Basionym: Monilia zeylanoides Castell., J. Trop. Med. Hyg. 23: 17. 1920.
Synonym: Candida zeylanoides (Castell.) Langeron & Guerra, Ann. Parasitol. Humaine Comp. 16: 501. 1938.
Fermentozyma Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, gen. nov.
MycoBank: MB848183
Etymology: the genus is named after the fermentation ability of the type species.
Type species: Fermentozyma sake (Saito & M. Ota) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai.
This genus is proposed for the lineage with Candida sake, which occurs in a separate branch from Teunomyces (Fig. 9). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis (Fig. 9) and rDNA-based phylogenetic analysis (Fig. 10), and the presence of genus-specific protein families OG0012707, OG0015120 and OG0019183 (Table 6).
Sexual reproduction not known. Colonies white or cream, soft. Budding is multilateral. Pseudohyphae present or not.
Note: Fermentozyma can be distinguished from its closely related genus Teunomyces by genome size and GC content (Table S3). Fermentozyma has a 14 Mb genome size with GC 38.76%, whereas Teunomyces has a low genome size (10–13Mb) and higher GC content (41.03–46.16%). Our ITS+D1/D2 LSU analysis showed that Candida sp. KBP Y-6292 (GenBank OP941477) may represent the third species of Fermentozyma (Fig. 10, Table S2).
Species accepted:
Fermentozyma sake (Saito & M. Ota) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848184
Holotype: CBS 159, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from sake-moto.
Basionym: Eutorulopsis sake Saito & M. Ota, J. Brew. 12: 166. 1934.
Synonym: Candida sake (Saito & M. Ota) Uden & H.R. Buckley ex S.A. Mey. & Ahearn, Mycotaxon 17: 298. 1983.
Fermentozyma vespimorsuum (Padamsee, B.S. Weir, M.E. Petterson & P.K. Buchanan) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848185
Holotype: PDD 105304. NEW ZEALAND, isolated from the surface of the cup fungus.
Basionym: Candida vespimorsuum Padamsee, B.S. Weir, M.E. Petterson & P.K. Buchanan, Persoonia 38: 349. 2017.
Glucitozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858042
Etymology: the genus is named based on the utilization of D-glucitol of species in this lineage.
Type species: Glucitozyma multigemmis (Buhagiar) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida multigemmis, which occurs in a separate long branch from the other taxa in the Serinales (e.g., Metschwnikowiaceae and Debaryomycetaceae) (Fig. 12). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0018541, OG0018542 and OG0018545 (Table 6).
Sexual reproduction not known. Colonies are white, smooth, creamy, semi-glossy. Pseudohyphae and hyphae are absent. Coenzyme Q-9 is formed.
Note: Glucitozyma differs from its closely related genus Meyerozyma by lower 32.6% GC content, whereas the genus Meyerozyma has a higher 42.5–50% GC content (Table S3). The ITS blast analysis demonstrated that Candida sp. CPD-35-1 (GenBank MZ701688) differs from C. multigemmis by 14 nt (2%) in the ITS region, which indicates that Candida sp. CPD-35-1 represents a potential new member of Glucitozyma (Fig. S12).
Species accepted:
Glucitozyma multigemmis (Buhagiar) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858043
Holotype: CBS 6524, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. UK, isolated from raspberry (Rubus idaeus).
Basionym: Torulopsis multigemmis Buhagiar, J. Gen. Microbiol. 86: 7. 1975.
Synonym: Candida multigemmis (Buhagiar) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Hemisphaericaspora F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858044
For a description: see Hui et al., PLoS ONE 9: e103737, 7. 2014.
Synonym: Hemisphaericaspora F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu, PLoS ONE 9: e103737, 7. 2014. Nom. inval., Art. 40.1 (Melbourne). The name of the type species of Hemisphaericaspora is invalid, therefore, the genus name Hemisphaericaspora is also invalid without a type.
Type species: Hemisphaericaspora nanyangensis F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
Species accepted:
Hemisphaericaspora elongata (C.Y. Chai & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858046
Holotype: NYNU 18115, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Jinghong, Mengyang town, isolated from rotting wood from a tropical rainforest, August 2018.
Basionym: Spathaspora elongata C.Y. Chai & F.L. Hui, MycoKeys 75: 36. 2020.
Hemisphaericaspora gorwiae (M.R. Lopes, C.G. Morais, R.M. Cadete, C. Fonseca, Kominek, Hittinger, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858047
Holotype: UFMG-CM-Y312, preserved in a metabolically inactive state at the Collection of Microorganisms and Cells of Federal University of Minas Gerais (UFMG), Belo Horizonte, Minas Gerais, Brazil. BRAZIL, isolated from rotting wood.
Basionym: Spathaspora gorwae M.R. Lopes, C.G. Morais, R.M. Cadete, C. Fonseca, Kominek, Hittinger, Lachance & C.A. Rosa [as ‘gorwiae’], FEMS Yeast Res. 16: fow044, 10. 2016.
Hemisphaericaspora haegerdaliae (M.R. Lopes, C.G. Morais, R.M. Cadete, C. Fonseca, Kominek, Hittinger, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858048
Holotype: UFMG-CM-Y303, preserved in a metabolically inactive state at the the Collection of Microorganisms and Cells of Federal University of Minas Gerais (UFMG), Belo Horizonte, Minas Gerais, Brazil. BRAZIL, isolated from rotting wood.
Basionym: Spathaspora haegerdaliae M.R. Lopes, C.G. Morais, R.M. Cadete, C. Fonseca, Kominek, Hittinger, Lachance & C.A. Rosa [as ‘hagerdaliae’], FEMS Yeast Res. 16: fow044, 10. 2016.
Hemisphaericaspora insectamans (D.B. Scott, van der Walt & Klift) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858049
Holotype: CBS 6033, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from frass of beetle larvae in gum Arabic tree (Acacia nilotica).
Basionym: Candida insectamans D.B. Scott, van der Walt & Klift, Mycopathol. Mycol. Appl. 47: 226. 1972.
Synonym: Hemisphaericaspora insectamans (D.B. Scott, van der Walt & Klift) F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu, PLoS ONE 9: e103737, 8. 2014. Nom. inval., Art. 35.1, the genus name is invalid (Melbourne).
Hemisphaericaspora jiuxiensis (C.Y. Chai & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858050
Holotype: NYNU 17416, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Honghe prefecture, Luxi county, Jiuxi Mountain Forest Park, isolated from rotting wood, July 2017.
Basionym: Spathaspora jiuxiensis C.Y. Chai & F.L. Hui, MycoKeys 75: 40. 2020.
Hemisphaericaspora lyxosophila (van der Walt, N.P. Ferreira & Steyn) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858051
Holotype: CBS 8194, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from the surface of the woodland soil.
Basionym: Candida lyxosophila van der Walt, N.P. Ferreira & Steyn, Antonie van Leeuwenhoek 53: 93. 1987.
Hemisphaericaspora mengyangensis (C.Y. Chai & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858052
Holotype: NYNU 17741, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Jinghong, Mengyang town, isolated from rotting wood from a tropical rainforest, July 2017.
Basionym: Spathaspora mengyangensis C.Y. Chai & F.L. Hui, MycoKeys 75: 38. 2020.
Hemisphaericaspora nanyangensis F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858053
For a description: see Hui et al., PLoS ONE 9: e103737, 7. 2014.
Holotype: CBS 13020, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Nanyang, Baotianman Nature Reserve, isolated from the frass of beetle larvae, July 2013.
Synonym: Hemisphaericaspora nanyangensis F.L. Hui, Y.C. Ren, Liang Chen, Ying Li, Lin Zhang & Q.H. Niu, PLoS ONE 9: e103737, 7. 2014. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Hemisphaericaspora parajiuxiensis (C.Y. Chai & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858054
Holotype: NYNU 16747, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Honghe prefecture, Luxi county, Jiuxi Mountain Forest Park, isolated from rotting wood, July 2016.
Basionym: Spathaspora parajiuxiensis C.Y. Chai & F.L. Hui, MycoKeys 75: 41. 2020.
Hemisphaericaspora roraimensis (R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858216
Holotype: UFMG-XMD 23.2. BRAZIL, Roraima, isolated from rotting wood in the Amazonian forest ecosystem.
Basionym: Spathaspora roraimensis R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa [as ‘roraimanensis’], Antonie van Leeuwenhoek 103: 428. 2013.
Hemisphaericaspora rosae (C.Y. Chai & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858055
Holotype: NYNU 17934, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Jinghong, Mengyang town, isolated from rotting wood in a tropical rainforest, July 2017.
Basionym: Spathaspora rosae C.Y. Chai & F.L. Hui, MycoKeys 75: 43. 2020.
Hemisphaericaspora subhashii (M. Groenew., Sigler & S.E. Richardson) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858056
Holotype: CBS 10753, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CANADA, Ontario, isolated from the peritoneal dialysis fluid of man.
Basionym: Candida subhashii M. Groenew., Sigler & S.E. Richardson, Med. Mycol. 47: 308. 2009.
Hemisphaericaspora xylanilytica Boonmak, Limtong, Jindam., Am-in, Yongman., Nakase & H. Kawas. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858057
For a description: see Boonmak et al., Int. J. Syst. Evol. Microbiol. 61: 1231. 2011.
Holotype: CBS 11761, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Nakhon Ratchasima, National Corn and Sorghum Research Center, isolated from decayed corn cobs.
Synonym: Candida xylanilytica Boonmak, Limtong, Jindam., Am-in, Yongman., Nakase & H. Kawas., Int. J. Syst. Evol. Microbiol. 61: 1231. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Hemisphaericaspora xylofermentans (R.M. Cadete, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858058
Holotype: UFMG-HMD23.3. BRAZIL, Roraima, isolated from rotting wood in the Amazonian forest ecosystem.
Basionym: Spathaspora xylofermentans R.M. Cadete, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa, Antonie van Leeuwenhoek 103: 430. 2013.
Insectozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858064
Etymology: the genus name is based on the ecological origin of most species that were isolated from insects.
Type species: Insectozyma corydali (N.H. Nguyen, S.O. Suh & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species of the Candida corydali clade, which occur in a separate clade from the Candida sensu stricto clade, the Lodderomyces clade, Nematodospora, Scheffersomyces, the Candida blackwelliae clade, the Hemisphaericaspora clade, and the single-species lineage Scheffersomyces stambukii (Fig. 9). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0008208, OG0008209 and OG0008210 (Table 6).
Sexual reproduction not known. Colonies are white to cream, pinkish, smooth and butyrous. Budding is multilateral. Pseudohyphae and septate hyphae are present or not. Where known, coenzyme Q-9 is formed.
Species accepted:
Insectozyma bohioensis (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858065
Holotype: CBS 9897, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. PANAMA, Barro Colorado Island, isolated from the gut of an unidentified click beetle.
Basionym: Candida bohioensis S.O. Suh, N.H. Nguyen & M. Blackw. [as 'bohiensis'], FEMS Yeast Res. 8: 94. 2008.
Insectozyma chauliodis (N.H. Nguyen, S.O. Suh & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858066
Holotype: NRRL Y-27909, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Louisiana, Livingston Parish, isolated from the surface of a male fishfly (Chauliodes pectinicornis).
Basionym: Candida chauliodis N.H. Nguyen, S.O. Suh & M. Blackw. [as 'chauliodes'], Mycologia 99: 847. 2008 [2007].
Insectozyma coleopterorum (F.L. Hui & X.J. Liu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858067
Holotype: CBS 14180, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Baotianman mountain, isolated from Anomala heydeni.
Basionym: Candida coleopterorum F.L. Hui & X.J. Liu, Int. J. Syst. Evol. Microbiol. 66: 4888. 2016.
Insectozyma corydali (N.H. Nguyen, S.O. Suh & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858068
Holotype: NRRL Y-27910, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Louisiana, Livingston Parish, isolated from the surface of a female Corydalus cornutus.
Basionym: Candida corydali N.H. Nguyen, S.O. Suh & M. Blackw., Mycologia 99: 849. 2008.
Insectozyma morakotiae (Nakase, Jindam., Ninomiya, Imanishi & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858069
Holotype: BCC 7718, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Nakhonratchasima province, Khao-Yai National Park, isolated fruit body of an unidentified mushroom, November 2000.
Basionym: Candida morakotiae Nakase, Jindam., Ninomiya, Imanishi & H. Kawas., J. Gen. Appl. Microbiol. 55: 98. 2009.
Insectozyma parachauliodis (F.L. Hui & X.J. Liu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858070
Holotype: CBS 13928, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Baotianman mountain, isolated from beetle larva, September 2014.
Basionym: Candida parachauliodis F.L. Hui & X.J. Liu [as ‘parachauliodes’], Int. J. Syst. Evol. Microbiol. 66: 4888. 2016.
Insectozyma prachuapensis Boonmak, Nitiyon, Am-in, Jindam., H. Kawas., Yongman. & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858071
For a description: see Nitiyon et al., Int. J. Syst. Evol. Microbiol. 61: 466. 2011.
Holotype: CBS 11024, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Prachuap Khiri Khan province, isolated from water collected from a mangrove forest.
Synonym: Candida prachuapensis Boonmak, Nitiyon, Am-in, Jindam., H. Kawas., Yongman. & Limtong, Int. J. Syst. Evol. Microbiol. 61: 466. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Insectozyma sakaeoensis Limtong, Koowadj., Jindam., Yongman. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858072
For a description: see Limtong et al., Antonie van Leeuwenhoek 102: 227. 2012.
Holotype: CBS 12318, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Sa Kaeo province, isolated from the phylloplane of Burma pad tree (Pterocarpus indicus).
Synonym: Candida sakaeoensis Limtong, Koowadj., Jindam., Yongman. & Nakase, Antonie van Leeuwenhoek 102: 227. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Insectozyma verbasci Sipiczki ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858073
For a description: see Sipiczki, Antonie van Leeuwenhoek 103: 573. 2012 [2013].
Holotype: CBS 12699, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. GEORGIA, Tbilisi, isolated from the flowers of Verbascum.
Synonym: Candida verbasci Sipiczki, Antonie van Leeuwenhoek 103: 573. 2012 [2013]. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Insectozyma xiaguanensis (F.L. Hui & X.J. Liu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858074
Holotype: CBS 13923, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Baotianman mountain, isolated from Anomala corpulenta, August 2014.
Basionym: Candida xiaguanensis F.L. Hui & X.J. Liu, Int. J. Syst. Evol. Microbiol. 66: 4887. 2016.
Intestinozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858059
Etymology: the genus is named based on the ecological origin of the species that was isolated from the gut of a beetle.
Type species: Intestinozyma alai (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida alai, which occurs in a separate long branch from the Spathaspora sensu stricto clade and the Hemisphaericaspora clade (Fig. 9). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0014078, OG0014079 and OG0017124 (Table 6).
Sexual reproduction not known. Colonies are cream, smooth, butyrous, shiny. Budding is multilateral. Pseudohyphae and true hyphae are not present.
Note: Intestinozyma alai differs from species of the closely related genus Spathaspora by lack of assimilation of erythritol and N-acetyl-D-glucosamine (Table S3). Ecologically, this new genus appears to be associated with the gut of insects. Our ITS analysis showed that Candida sp. B53C (GenBank MW165503) may represent another new member of the genus Intestinozyma (Fig. S11, Table S2).
Species accepted:
Intestinozyma alai (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858062
Holotype: NRRL Y-27739, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Louisiana, Livingston Parish, isolated from the gut of the blind click beetle (Alaus myops).
Basionym: Candida alai S.O. Suh, N.H. Nguyen & M. Blackw., FEMS Yeast Res. 8: 95. 2008.
Keqinozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858075
Etymology: the genus is named in honor of Dr. Ke-Qin Zhang for his contribution to the studies of biocontrol fungi against root-knot nematode.
Type species: Keqinozyma sinolaborantium (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Candida nonsorbophila clade, which occur in a separate clade from the Yamadazyma sensu stricto clade, the Yamadazyma epiphylla clade, the Yamadazyma olivae clade, and the Yamadazyma triangularis clade (Figs. 12–13). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis (Fig. 12) and rDNA phylogenetic analysis (Fig. 13), and the presence of genus-specific protein families OG0018587, OG0018588 and OG0018589 (Table 6).
Sexual reproduction not known. Colonies are white to cream, smooth and butyrous. Budding is multilateral. Pseudohyphae are present, but hyphae are not observed. Where known, coenzyme Q-9 is formed.
Note: Ecologically, this new genus seems to be associated with the insects or the plant-water interface.
Species accepted:
Keqinozyma heliconiae (Ruivo, Pagnocca, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858076
Holotype: CBS 10000, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, São Paulo, isolated from water accumulated in the flower bracts of Heliconia velloziana.
Basionym: Candida heliconiae Ruivo, Pagnocca, Lachance & C.A. Rosa, Int. J. Syst. Evol. Microbiol. 56: 1148. 2006.
Keqinozyma nonsorbophila (Nakase, Jindam., Am-in, Ninomiya, H. Kawas. & Limtong) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858077
Holotype: BCC 25963, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Center for Genetic Engineering and Biotechnology (BIOTEC), Pathumthani, Thailand. THAILAND, Ranong Province, Laem Son National Park, isolated from water in mangrove forest, May 2006.
Basionym: Candida nonsorbophila Nakase, Jindam., Am-in, Ninomiya, H. Kawas. & Limtong, FEMS Yeast Res. 9: 665. 2009.
Keqinozyma sinolaborantium (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858078
Holotype: NRRL Y-27765, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. PANAMA, Barro Colorado Island, isolated from the gut of Amphix.
Basionym: Candida sinolaborantium S.O. Suh, N.H. Nguyen & M. Blackw., Mycol. Res. 109: 1053. 2005.
Keqinozyma temnochilae (S.O. Suh, N.H. Nguyen & M. Blackw.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858079
Holotype: NRRL Y-27763, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. PANAMA, Barro Colorado Island, isolated from the gut of Temnochila.
Basionym: Candida temnochilae S.O. Suh, N.H. Nguyen & M. Blackw., Mycol. Res. 109: 1052. 2005.
Lizanozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858032
Etymology: the genus is named in honor of the late artist Lizan Freijsen for her contribution to fungi-inspired textile art. She contributed to bridge the gap between science and art. Her master piece ‘The Fungal Wall’ is on display in the museum Micropia in Amsterdam and a new work is expected for ‘Het Groote Museum’, also in Amsterdam. In addition, she has been a great inspirator for young artists.
Type species: Lizanozyma spartinae (Ahearn, Yarrow & Meyers) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Scheffersomyces gosingicus clade, which occur in a separate clade from Priceomyces, Schwanniomyces and other genera in Serinales (Figs. 9–10). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0013736 and OG0022956 (Table 6).
Both teleomorphic and anamorphic species occur in this genus. Asci are unconjugated or conjugated from the pairing of complementary mating types. Asci produce one, but usually two, hat-shaped ascospores. Colonies are white to tannish-white, smooth, butyrous, glistening. Budding is multilateral. Pseudohyphae are formed, but true hyphae do not develop. D-xylose is not assimilated. Where known, coenzyme Q-9 is formed.
Note: Lizanozyma spp. are distinct from those of the closely related genus Priceomyces by the lack of assimilation of galactose and the ability to ferment glucose. Lizanozyma spp. differs from Schwanniomyces spp. by the lack of xylose assimilation (Table S3).
Species accepted:
Lizanozyma gosingica C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858033
For a description: see Chang et al., Int. J. Syst. Evol. Microbiol. 61: 692. 2011.
Holotype: CBS 11433, preserved in a metabolically inactive state at Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Nanto, isolated from soil.
Synonyms: Candida gosingica C.F. Lee, Int. J. Syst. Evol. Microbiol. 61: 692. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Scheffersomyces gosingicus H. Urbina & M. Blackw., PLoS ONE 7: e39128, 7. 2012. Nom. inval., Art. 40.1 (Melbourne), published as a combination, but ‘basionym’ (Candida gosingica) is invalid and bibliographic error in the basionym reference.
Lizanozyma spartinae (Ahearn, Yarrow & Meyers) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858034
Holotype: NRRLY-7322, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Louisiana, Barataria Bay, isolated from oyster grass (Spartina alterniflora) marshes.
Basionym: Pichia spartinae Ahearn, Yarrow & Meyers, Antonie van Leeuwenhoek 36: 503. 1970.
Synonym: Scheffersomyces spartinae (Ahearn, Yarrow & Meyers) Kurtzman & M. Suzuki [as ‘spartiniae’], Mycoscience 51: 9. 2010.
Lizanozyma thasaensis Poomtien, Jindam., Limtong, Pinphan. & Thaniy. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858035
For a description: see Poomtien et al., Antonie van Leeuwenhoek 103: 236. 2012 [2013].
Holotype: CBS 12529, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Chumporn province, Thasae, a soil sediment pond of a palm oil biodiesel production plant, May 2009.
Synonym: Candida thasaensis Poomtien, Jindam., Limtong, Pinphan. & Thaniy. [as ‘thasaenensis’], Antonie van Leeuwenhoek 103: 236. 2012 [2013]. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
New combinations and validated taxa of Lodderomyces
Lodderomyces cetoniae (Gouliam., R.A. Dimitrov, M.T. Sm., M. Groenew. & Boekhout) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858091
Holotype: IMB1R2, preserved in a metabolically inactive state at the yeast collection of the Institute of Microbiology, Sofia, Bulgaria. BULGARIA, East Rhodopes, isolated from the beetles Cetonia aurata.
Synonym: Candida cetoniae Gouliam., R.A. Dimitrov, M.T. Sm., M. Groenew. & Boekhout, Fungal Biol. 120: 188. 2016.
Lodderomyces hyderabadensis (R.Sreen. Rao, Bhadra, N.N. Kumar & Shivaji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858092
Holotype: CBS 10444, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. INDIA, AP, Hyderabad, isolated from wine grapes.
Basionym: Candida hyderabadensis R.Sreen. Rao, Bhadra, N.N. Kumar & Shivaji, FEMS Yeast Res. 7: 491. 2007.
Lodderomyces jiufengensis (F.Y. Bai & Z.H. Ji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858093
Holotype: CGMCC 2.3688, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Beijing, Jiufeng mountain, isolated from the gut of Oxycetonia jucunda, July 2007.
Basionym: Candida jiufengensis F.Y. Bai & Z.H. Ji, Antonie van Leeuwenhoek 95: 30. 2009.
Lodderomyces margitis (F.L. Hui & X.J. Liu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858097
Holotype: CBS 14175, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Baotianman mountain, isolated from Margites fulvidus, August 2015.
Basionym: Candida margitis F.L. Hui & X.J. Liu, Int. J. Syst. Evol. Microbiol. 66: 4887. 2016.
Lodderomyces metapsilosis (Tavanti, A. Davidson, Gow, M. Maiden & Odds) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858098
Holotype: ATCC 96144, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, Washington, isolated from the hand of Homo sapiens.
Basionym: Candida metapsilosis Tavanti, A. Davidson, Gow, M. Maiden & Odds, J. Clin. Microbiol. 43: 290. 2005.
Lodderomyces orthopsilosis (Tavanti, A. Davidson, Gow, M. Maiden & Odds) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858099
Holotype: ATCC 96139, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, Texas, isolated from the tip of the catheter.
Basionym: Candida orthopsilosis Tavanti, A. Davidson, Gow, M. Maiden & Odds, J. Clin. Microbiol. 43: 290. 2005.
Lodderomyces oxycetoniae (F.Y. Bai & Z.H. Ji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858100
Holotype: CGMCC 2.3656, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Beijing, Baiwangshan mountain, isolated from the gut of Oxycetonia jucunda, July 2007
Basionym: Candida oxycetoniae F.Y. Bai & Z.H. Ji, Antonie van Leeuwenhoek 95: 29. 2009.
Lodderomyces parapsilosis (Ashford) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858101
Holotype: CBS 604, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. PUERTO RICO, isolated from sprue.
Basionym: Monilia parapsilosis Ashford, Amer. J. Trop. Med. 8: 518. 1928.
Synonyms: Candida parapsilosis (Ashford) Langeron & Talice, Ann. Parasitol. Humaine Comp. 10: 54. 1932.
Mycocandida parapsilosis (Ashford) C.W. Dodge, Medic. Mycol.: 294. 1935.
Mycotorula parapsilosis (Ashford) Cif. & Redaelli, Atti Ist. Bot. Lab. Crittog. Univ. Pavia, sér. 5 3: 47. 1943.
Lodderomyces pseudojiufengensis (F.Y. Bai & Z.H. Ji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858102
Holotype: CGMCC 2.3693, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Beijing, Jiufeng mountain, isolated from the gut of Oxycetonia jucunda, July 2007.
Basionym: Candida pseudojiufengensis F.Y. Bai & Z.H. Ji, Antonie van Leeuwenhoek 95: 30. 2009.
Lodderomyces theae C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858103
For a description: see Chang et al., Int. J. Food Microbiol. 153: 13. 2012.
Holotype: CBS 12239, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, isolated from tea drink.
Synonym: Candida theae C.F. Lee, Int. J. Food Microbiol. 153: 13. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
New combinations of Millerozyma
Millerozyma porticicola (Ninomiya, Mikata, Nakagiri, Nakase & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858104
Holotype: NBRC 100302, preserved in a metabolically inactive state at Biological Resource Center, NITE (NBRC), Tokyo, Japan. JAPAN, Wakavama Pref., isolated from the gallery of Indocryphalus pubipennis infesting Quercus acutissima.
Basionym: Pichia porticicola Ninomiya, Mikata, Nakagiri, Nakase & H. Kawas., J. Gen. Appl. Microbiol. 56: 284. 2010.
Millerozyma pseudofarinosa (S. Mallet, S. Weiss, N. Jacques & Casarég.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858105
Holotype: NCYC 386, preserved in a metabolically inactive state at Biological Resource Center, NITE (NBRC), Tokyo, Japan. JAPAN, isolated from an unknown substrate.
Basionym: Candida pseudofarinosa S. Mallet, S. Weiss, N. Jacques & Casarég., PLoS ONE 7: e35842, 9. 2012.
Nothofagozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858080
Etymology: the genus is named based on the ecological origin of the species that were isolated from decomposed wood of Nothofagus.
Type species: Nothofagozyma chilensis (Grinb. & Yarrow) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida chilensis, which is located in a separate branch closely related to Cephaloascus (Fig. 12). Member of the Cephaloascaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0017645, OG0017647 and OG0017651 (Table 6).
Sexual reproduction is not known. Colonies are cream, shiny, smooth and butyrous. Budding is multilateral. Hyphae may be present. Coenzyme Q-9 is formed.
Note: Nothofagozyma chilensis assimilates sucrose, maltose, melezitose, methyl-α-D-glucoside, D-ribose, erythritol, hexadecane, nitrate and nitrite, whereas species of its closely related genus Cephaloascus do not (Table S3). Our D1/D2 LSU analysis showed that Candida cf. chilensis CBS 11766 (GenBank FN824503) represents a potential new species of Nothofagozyma (Fig. S9, Table S2).
Species accepted:
Nothofagozyma chilensis (Grinb. & Yarrow) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858081
Holotype: CBS 5719, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. Chile, isolated from decomposed wood of the southern beech tree (Nothofagus sp.).
Basionym: Candida chilensis Grinb. & Yarrow, Antonie van Leeuwenhoek 36: 144. 1970.
New combinations and validated taxa of Scheffersomyces
Scheffersomyces broadrunensis (S.O. Suh & J.J. Zhou) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858106
Holotype: ATCC MYA-4650, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, VA, Broad Run, Bull Run Mountain, isolated from the gut of an unidentified tenebrionid beetle on a rotten log.
Basionym: Candida broadrunensis S.O. Suh & J.J. Zhou, Int. J. Syst. Evol. Microbiol. 63: 4334. 2013.
Scheffersomyces lignicola Jindam., Limtong, Yongman., Tuntir., Potach., H. Kawas. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858107
For a description: see Jindam. et al., FEMS Yeast Res. 7: 1412. 2007.
Holotype: NBRC 102564, preserved in a metabolically inactive state at Biological Resource Center, NITE (NBRC), Tokyo, Japan. THAILAND, isolated from insect frass.
Synonyms: Candida lignicola Jindam., Limtong, Yongman., Tuntir., Potach., H. Kawas. & Nakase, FEMS Yeast Res. 7: 1412. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Scheffersomyces lignicola Jindam., Limtong, Yongman., Tuntir., Potach., H. Kawas. & Nakase ex H. Urbina & M. Blackw., PLoS ONE 7: e39128, 9. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified; published as a combination, but 'basionym' (Candida lignicola) is invalid.
Validated taxa in Spathaspora
Spathaspora brasiliensis R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858108
For a description: see Cadete et al., Antonie van Leeuwenhoek 103: 426. 2013.
Holotype: CBS 12679, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, isolated from rotting wood.
Synonym: Spathaspora brasiliensis R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa, Antonie van Leeuwenhoek 103: 426. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Spathaspora suhii R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858109
For a description: see Cadete et al., Antonie van Leeuwenhoek 103: 428. 2013.
Holotype: CBS 12680, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, isolated from rotting wood.
Synonym: Spathaspora suhii R.M. Cadete, Zilli, M.J.S. Vital, F.C.O. Gomes, Stambuk, Lachance & C.A. Rosa, Antonie van Leeuwenhoek 103: 428. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
New combinations in Suhomyces
Suhomyces caryicola (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858110
Holotype: NRRL YB-1499, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, IL, Peoria, isolated from a nut pod of a pignut hickory tree (Carya glabra).
Basionym: Candida caryicola Kurtzman, FEMS Yeast Res. 1: 182. 2001.
Suhomyces rongomai-pounamu (Padamsee, B.S. Weir, M.E. Petterson & P.K. Buchanan) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858111
Holotype: PDD 105303. NEW ZEALAND, Auckland, The Gardens, Totara Park, isolated from agaric mushroom surface, March 2016.
Basionym: Candida rongomai-pounamu Padamsee, B.S. Weir, M.E. Petterson & P.K. Buchanan, Persoonia 38: 347. 2017.
Suhomyces tibetensis (F.Y. Bai & Z.W. Wu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858112
Holotype: CGMCC 2.3072, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Tibet, Linzhi, isolated from the flower, July 2004.
Basionym: Candida tibetensis F.Y. Bai & Z.W. Wu, Int. J. Syst. Evol. Microbiol. 56: 1154. 2006.
Suzukiozyma Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, gen. nov.
MycoBank: MB848205
Etymology: the genus is named in honor of M. Suzuki for his contribution to yeast taxonomy.
Type species: Suzukiozyma glaebosa (Komag. & Nakase) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai.
This genus is proposed for the species in the Candida glaebosa clade, which occur in a separate clade from the genus Diutina (Fig. 9, Daniel et al. 2014). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0007615 and OG0007620 (Table 6).
Sexual reproduction not known. Colonies are white to cream, butyrous. Budding is multilateral. Hyphae not produced, pseudohyphae are present or not.
Note: Suzukiozyma differs from its closely related genus Diutina by a lower GC content (31.16–41.24%), whereas the genus Diutina has a higher GC content (41.23–53.05%) (Table S3).
Species accepted:
Suzukiozyma candida (Lodder) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848211
Holotype: CBS 940, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from the atmosphere.
Basionym: Torulopsis candida Lodder, Verh. Kon. Ned. Akad. Wetensch., Afd. Natuurk. 32: 163. 1934.
Synonyms: Torula candida Saito, J. Jap. Bot. 1: 41 1922. Nom. illeg., Art. 53.1, homonym, non Torula candida Opiz, 1855.
Cryptococcus candidus (Lodder) C.E. Skinner, Amer. Midl. Naturalist 43: 249. 1950.
Candida saitoana Nakase & M. Suzuki, J. Gen. Appl. Microbiol. 31: 85. 1985.
Suzukiozyma fluviatilis (L.R. Hedrick) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848206
Holotype: CBS 6776, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, Indiana, isolated from polluted river water.
Basionym: Candida fluviatilis L.R. Hedrick, Antonie van Leeuwenhoek 42: 329. 1976.
Suzukiozyma glaebosa (Komag. & Nakase) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848207
Holotype: CBS 5691, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from frozen cuttlefish.
Basionym: Candida glaebosa Komag. & Nakase, J. Gen. Appl. Microbiol. 11: 262. 1965.
Suzukiozyma manassasensis (S.O. Suh & J.J. Zhou) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848208
Holotype: ATCC MYA-4652, preserved in a metabolically inactive state at the American Type Culture Collection (ATCC), Manassas, USA. USA, VA, Broad Run, Bull Run Mountain, isolated from the gut of Xylopinus saperdioides.
Basionym: Candida manassasensis S.O. Suh & J.J. Zhou, Int. J. Syst. Evol. Microbiol. 63: 4336. 2013.
Suzukiozyma palmioleophila (Nakase & Itoh) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848209
Holotype: JCM 5218, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. JAPAN, isolated from soil.
Basionym: Candida palmioleophila Nakase & Itoh, J. Gen. Appl. Microbiol. 34: 496. 1988.
Suzukiozyma pseudoglaebosa (M. Suzuki & Nakase) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848210
Holotype: JCM 2168, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. JAPAN, isolated from soil.
Basionym: Candida pseudoglaebosa M. Suzuki & Nakase, Bull. Jap. Fed. Cult. Coll. 9: 130. 1993.
Suzukiozyma sphagnicola (Kachalkin & Yurkov) Q.M. Wang, Yurkov, Boekhout & F.Y. Bai, comb. nov.
MycoBank: MB848212
Holotype: CBS 11774, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. RUSSIA, Tver region, Central Forest State Biosphere Reserve, isolated from Sphagnum girgensohnii.
Basionym: Candida sphagnicola Kachalkin & Yurkov, Antonie van Leeuwenhoek 102: 40. 2011.
New combinations and validated taxa in Yamadazyma
Yamadazyma andamanensis Am-In, Limtong, Yongman. & Jindam. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858113
For a description: see Am-In et al., Int. J. Syst. Evol. Microbiol. 61: 459. 2011.
Holotype: CBS 10859, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Ranong province, Laem Son National Park, isolated from the estuarine water of the mangrove forest.
Synonym: Candida andamanensis Am-In, Limtong, Yongman. & Jindam., Int. J. Syst. Evol. Microbiol. 61: 459. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma dushanensis F.L. Hui, Yun Wang, Y.C. Ren & Ying Li ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858116
For a description: see Wang et al., Curr. Microbiol. 71: 272. 2015.
Holotype: CBS 13914, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Nanyang, Dushan Forest Park, isolated from rotten wood, June 2014.
Synonym: Yamadazyma dushanensis F.L. Hui, Yun Wang, Y.C. Ren & Ying Li, Curr. Microbiol. 71: 272. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma jaroonii Imanishi, Jindam., Mikata, Nagak., Potach., Tantich. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862196
For a description: see Imanishi et al., Antonie van Leeuwenhoek 94(2): 273. 2007.
Holotype: CBS 10790, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Kao-Yaow, isolated from insect frass, March 2001.
Synonym: Candida jaroonii Imanishi, Jindam., Mikata, Nagak., Potach., Tantich. & Nakase, Antonie van Leeuwenhoek 94(2): 273. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma oceani Burgaud & G. Barbier ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858117
For a description: see Burgaud et al., Antonie van Leeuwenhoek 100: 79. 2011.
Holotype: CBS 11857, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ATLANTIC OCEAN, Mid-Atlantic Ridg, isolated from an unidentified deep-sea coral.
Synonyms: Candida oceani Burgaud & G. Barbier, Antonie van Leeuwenhoek 100: 79. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma oceani Burgaud & G. Barbier ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4776. 2025. Nom. inval., Art. 40.1 (Shenzhen); published as a combination, but 'basionym' (Candida oceani) is invalid.
Yamadazyma paraphyllophila Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858118
For a description: see Kaewwichian et al., Antonie van Leeuwenhoek 103: 786. 2012.
Holotype: CBS 9928, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Pindung, NanJen-Shan Natural Reserve, isolated from the phylloplane of hengchun pencilwood (Dysoxylum hongkongense).
Synonym: Yamadazyma paraphyllophila Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong, Antonie van Leeuwenhoek 103: 786. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma phyllophila Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858119
For a description: see Kaewwichian et al., Antonie van Leeuwenhoek 103: 786. 2012.
Holotype: CBS 12572, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Nakhon Ratchasima province, isolated from the phylloplane of corn (Zea mays).
Synonym: Yamadazyma phyllophila Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong, Antonie van Leeuwenhoek 103: 786. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma siamensis Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858120
For a description: see Kaewwichian et al., Antonie van Leeuwenhoek 103: 784. 2012.
Holotype: CBS 12573, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Phetchabun province, isolated from the phylloplane sugarcane (Saccharum officinarum).
Synonym: Yamadazyma siamensis Kaewwich., Yongman., H. Kawas., P.H. Wang & Limtong, Antonie van Leeuwenhoek 103: 784. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma songkhlaensis Imanishi, Jindam., Mikata, Nagak., Potach., Tantich. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862197
For a description: see Imanishi et al., Antonie van Leeuwenhoek 94(2): 274. 2007.
Holotype: CBS 10791, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Nam Tok Tone Nga-Chang Waterfall, isolated from insect frass, March 2001.
Synonym: Candida songkhlaensis Imanishi, Jindam., Mikata, Nagak., Potach., Tantich. & Nakase, Antonie van Leeuwenhoek 94(2): 274. 2007. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma ubonensis Junyapate, Jindam. & Limtong ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858125
For a description: see Junyapate et al., Antonie van Leeuwenhoek 105: 477. 2014.
Holotype: CBS 12859, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Ubon Ratchathani province, isolated from the tree bark.
Synonym: Yamadazyma ubonensis Junyapate, Jindam. & Limtong, Antonie van Leeuwenhoek 105: 477. 2014. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Yamadazyma vrieseae Landell & P. Valente ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862198
For a description: see Landell et al., Int. J. Syst. Evol. Microbiol. 60 (1): 247. 2010.
Holotype: CBS 10829, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, Rio Grande do Sul, Itapuã Park, isolated from the tank water of bromeliads.
Synonym: Candida vrieseae Landell & P. Valente, Int. J. Syst. Evol. Microbiol. 60(1): 247. 2010. Nom. inval., Art. 40.7 (Shenzhen), more than one collection in which the type is conserved was specified.
Zhuliangozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858084
Etymology: the genus is named in honor of Dr. Zhu-Liang Yang for his contribution to fungal taxonomy, especially macrofungal taxonomic system and economic mushrooms.
Type species: Zhuliangozyma blackwelliae (F.Y. Bai & Z.H. Ji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Candida blackwelliae clade, which occur in a separate clade from the Candida sensu stricto, Lodderomyces clade, Nematodospora, the Candida corydali clade, the Hemisphaericaspora clade, and the single-species lineage Scheffersomyces stambukii (Fig. 9). Member of the Debaryomycetaceae (Serinales, Pichiomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0009199 and OG0009200 (Table 6).
Both teleomorphic and anamorphic species occur in this genus. Unconjugated asci are formed from single cells with a single greatly elongated ascospore with tapered and curved ends. Colonies are white to cream, smooth, butyrous. Budding is multilateral. Pseudohyphae are present or not.
Note: Zhuliangozym differs from the closely related Candida/Lodderomyces lineage by a higher GC content (50.46–53.88%), whereas all genera in the Candida/Lodderomyces lineage have a GC content lower than 45% (Table S3).
Species accepted:
Zhuliangozyma blackwelliae (F.Y. Bai & Z.H. Ji) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858085
Holotype: CGMCC 2.3639, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Beijing, Baiwangshan mountain, isolated from the gut of Trichius succinctus, July 2007.
Basionym: Candida blackwelliae F.Y. Bai & Z.H. Ji [as 'blackwellae'], Antonie van Leeuwenhoek 95: 28. 2009.
Zhuliangozyma boniae (C.G. Morais, T.M. Batista, J. Kominek, G.R. Franco, C. Fonseca, C.T. Hittinger, M.A. Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858086
Holotype: UFMG-CM-Y306, preserved in a metabolically inactive state at the Collection of Microorganisms and Cells of Federal University of Minas Gerais (UFMG), Belo Horizonte, Minas Gerais, Brazil. BRAZIL, Rio Doce Ecological State Park, isolated from rotting wood.
Basionym: Spathaspora boniae C.G. Morais, T.M. Batista, J. Kominek, G.R. Franco, C. Fonseca, C.T. Hittinger, M.A. Lachance & C.A. Rosa, Int. J. Syst. Evol. Microbiol. 67: 3804. 2017.
Zhuliangozyma parablackwelliae (F.L. Hui & L.N. Huang) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858087
Holotype: NYNU 17763, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Xishuangbanna Tropical Rainforest, isolated from rotting wood.
Basionym: Candida parablackwelliae F.L. Hui & L.N. Huang, Int. J. Syst. Evol. Microbiol. 69: 2779. 2019.
New taxa and reinstated genus in Phaffomycetaceae (Phaffomycetales, Saccharomycetes)
Validated taxa in Barnettozyma
Barnettozyma sucrosica Imanishi, A. Yamaz. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858131
For a description: see Imanishi et al., J. Gen. Appl. Microbiol. 56: 449. 2010.
Holotype: CBS 11512, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, Rishiri Island, isolated from soil, July 2007.
Synonym: Barnettozyma sucrosica Imanishi, A. Yamaz. & Nakase, J. Gen. Appl. Microbiol. 56: 449. 2010. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Barnettozyma xylosica R. Kobay., A. Kanti & H. Kawas. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858132
For a description: see Kobayashi et al., Int. J. Syst. Evol. Microbiol. 67: 3974. 2017.
Holotype: NBRC 111558, preserved in a metabolically inactive state at Biological Resource Center, NITE (NBRC), Tokyo, Japan. INDONESIA, West Java, Chibodas Botanical Garden, isolated from litter, December 2014.
Synonym: Barnettozyma xylosica R. Kobay., A. Kanti & H. Kawas., Int. J. Syst. Evol. Microbiol. 67: 3974. 2017. Nom. inval., Art. 40.7 (Shenzhen), more than one collection in which the type is conserved was specified.
Barnettozyma xylosiphila R. Kobay., A. Kanti & H. Kawas. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858133
For a description: see Kobayashi et al., Int. J. Syst. Evol. Microbiol. 67: 3974. 2017.
Holotype: NBRC 110202, preserved in a metabolically inactive state at Biological Resource Center, NITE (NBRC), Tokyo, Japan. INDONESIA, West Sumatra, Bung Hatta Botanical Garden, isolated from decayed wood, May 2013.
Synonym: Barnettozyma xylosiphila R. Kobay., A. Kanti & H. Kawas., Int. J. Syst. Evol. Microbiol. 67: 3974. 2017. Nom. inval., Art. 40.7 (Shenzhen), more than one collection in which the type is conserved was specified.
Gotozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858126
Etymology: the genus is named in honor of S. Goto for his contribution to yeast taxonomy.
Type species: Gotozyma montana (Goto & Oguri) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Barnettozyma siamensis clade, which occur in a separate clade from the genera Barnettozyma sensu stricto, Millerago, the Barnettozyma wickerhamii clade and the Phaffomyces clade (Figs. 15–16). Member of the Phaffomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis and rDNA phylogenetic analysis.
Sexual reproduction not known. Colonies are white to cream, butyrous. Budding is multilateral. Pseudohyphae are present or not. True hyphae are absent. Where known, coenzyme Q-7 is formed.
Note: Gotozyma spp. have higher GC% (37.35–37.83%) compared with its closely related genus Phaffomyces (31.11–32.3%).
Species accepted:
Gotozyma botsteinii G. Arrey, G.S. Li, R. Murphy, L. Guimaraes, S. Alizadeh, M. Poulsen & B. Regenberg ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858129
For a description: see Arrey et al., G3, Genes, Genomes, Genetics 11 (12, jkab342): 7. 2021.
Holotype: CBS 16679, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. IVORY COAST, isolated from hindgut of Macrotermes bellicosus.
Synonym: Barnettozyma botsteinii G. Arrey, G.S. Li, R. Murphy, L. Guimaraes, S. Alizadeh, M. Poulsen & B. Regenberg, G3, Genes, Genomes, Genetics: 11 (12, jkab342): 7. 2021. Nom. inval., Art. 40.8 (Shenzhen), a statement that the culture is preserved in a metabolically inactive state was not included in the protologue.
Gotozyma montana (Goto & Oguri) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858127
Holotype: CBS 8057, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from the wild grape (Vitis coignetiae).
Basionym: Candida montana Goto & Oguri, J. Gen. Appl. Microbiol. 29: 88. 1983.
Gotozyma siamensis (Polburee & Limtong) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858128
Holotype: DMKU-UbN24(1). THAILAND, Ubon Ratchathani province, Na Chaluai district, isolated from soil.
Basionym: Barnettozyma siamensis Polburee & Limtong, Int. J. Syst. Evol. Microbiol. 64: 3055. 2014.
Reinstated genus
Komagataea Y. Yamada, M. Matsuda, K. Maeda, Sakak. & Mikata, Biosc. Biotechn. Biochem. 58: 1243. 1994.
MycoBank: MB27298
Type species: Komagataea pratensis (Babeva & Reshetova) Y. Yamada, M. Matsuda, K. Maeda, Sakak. & Mikata.
Species accepted:
Komagataea norvegica (Reiersöl) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858249
Holotype: CBS 4239, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. Norway, isolated from sputum.
Basionym: Torulopsis norvegica Reiersöl [as 'norwegica'], Antonie van Leeuwenhoek 24: 111. 1958.
Synonym: Candida norvegica (Reiersöl) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 613. 1978.
Komagataea pratensis (Babeva & Reshetova) Y. Yamada, M. Matsuda, K. Maeda, Sakak. & Mikata, Biosc. Biotechn. Biochem. 58: 1243. 1994.
MycoBank: MB362653
Basionym: Williopsis pratensis Babeva & Reshetova, Mikrobiologiya 48: 1041. 1979.
Synonym: Barnettozyma pratensis (Babeva & Reshetova) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 948. 2008.
Komagataea qinlingensis (F.Y. Bai & H.Z. Lu) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858136
Holotype: CGMCC 2.2524, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Shanxi province, Qinling, isolated from soil, October 2002.
Basionym: Candida qinlingensis F.Y. Bai & H.Z. Lu, Int. J. Syst. Evol. Microbiol. 54: 1411. 2004.
Komagataea salicaria (Phaff, M.W. Mill. & J.F.T. Spencer) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858250
Holotype: CBS 5456, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, isolated from slime fluxes of willows (Salix sp.).
Basionym: Pichia salicaria Phaff, M.W. Mill. & J.F.T. Spencer, Antonie van Leeuwenhoek 30: 139. 1964.
Synonym: Barnettozyma salicaria (Phaff, M.W. Mill. & J.F.T. Spencer) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 948. 2008.
Komagataea wickerhamii (van der Walt) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858137
Holotype: NRRL Y-2435, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. SOUTH AFRICA, isolated from the larval gut of Cossidae.
Basionym: Endomycopsis wickerhamii van der Walt, Antonie van Leeuwenhoek 25: 347. 1959.
Synonym: Barnettozyma wickerhamii (Van der Walt) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8(6): 948. 2008.
New taxon in Millerago
Millerago ficus F.L. Hui, Q.H. Niu, T. Ke & Zheng Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858248
For a description: see Hui et al., Int. J. Syst. Evol. Microbiol. 62(11): 2807. 2012.
Holotype: CBS 12638, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Henan province, Nanyang, isolated from the gut of larvae of Apriona germari in the trunk of Ficus carica.
Synonym: Candida ficus F.L. Hui, Q.H. Niu, T. Ke & Zheng Liu, Int. J. Syst. Evol. Microbiol. 62(11): 2807. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
New combination and validated taxa in Phaffomyces
Phaffomyces coquimbonensis Cardinali, Antonielli, L. Corte, Roscini & Ganter ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858134
For a description: see Cardinali et al., Int. J. Syst. Evol. Microbiol. 62: 3068. 2012.
Holotype: CBS 12348, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. PICHIDANGUI, isolated from a necrotic Echinopsis chiloensis.
Synonym: Candida coquimbonensis Cardinali, Antonielli, L. Corte, Roscini & Ganter, Int. J. Syst. Evol. Microbiol. 62: 3068. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Phaffomyces orbus (Starmer, Phaff, Ganter & Lachance) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858135
Holotype: UCD-FST 84-833.1. AUSTRALIA, Queensland, isolated from necrotic cladodes of Opuntia stricta.
Basionym: Candida orba Starmer, Phaff, Ganter & Lachance, Int. J. Syst. Evol. Microbiol. 51: 701. 2001.
New taxa and reinstated genus in Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes)
Buckleya Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858138
Etymology: the genus is named in honor of H.R. Buckley for her contribution to yeast taxonomy and ecology.
Type species: Buckleya freyschussii (H.R. Buckley & Uden) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the single-species lineage Candida freyschussii, which occurs in a separate long branch from the genus Cyberlindnera sensu stricto clade, the Williopsis clade, and the Wickerhamomyces hampshirensis clade (Figs. 15–16). Member of the Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis, and the presence of genus-specific protein families OG0010458, OG0010457 and OG0008787 (Table 6).
Sexual reproduction not known. Colonies are white to cream, smooth and glistening. Budding is multilateral. Pseudohyphae occur, but true hyphae not observed. Coenzyme Q-7 is formed.
Note: Our D1/D2 LSU analysis showed that Candida sp. NIAH-01 (GenBank AB703242) represents a potential new species of Buckleya (Fig. S13, Table S2).
Species accepted:
Buckleya freyschussii (H.R. Buckley & Uden) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858139
Holotype: CBS 2162, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SWEDEN, isolated from wet wood pulp.
Basionym: Candida freyschussii H.R. Buckley & Uden, Mycopath. Mycol. Appl. 36: 263. 1968.
New combinations and validated taxa in Hansenula
Hansenula queroliae (C.A. Rosa, P.B. Morais, Lachance & Pimenta) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858175
Holotype: CBS 10936, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, Tocantins, isolated from larva of Anastrepha mucronata from fruit of Peritassa campestris.
Basionym: Wickerhamomyces queroliae C.A. Rosa, P.B. Morais, Lachance & Pimenta, Int. J. Syst. Evol. Microbiol. 59: 1234. 2009.
Hansenula silvicultrix (van der Walt, D.B. Scott & Klift) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858176
Holotype: CBS 6269, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from frass of bostrichid beetle (Sinoxylon ruficorne) infesting sweet thorn (Acacia karroo).
Basionym: Candida silvicultrix van der Walt, D.B. Scott & Klift, Mycopathol. Mycol. Appl. 47: 234. 1972.
Hansenula spegazzinii Masiulionis & Pagnocca ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858177
For a description: see Masiulionis & Pagnocca, Int. J. Syst. Evol. Microbiol. 66: 2144. 2016.
Holotype: CBS 12756, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ARGENTINA, Santa Fe province, Santurce, isolated from the fungus garden of an ant nest of Acromyrmex lundii, September 2009.
Synonym: Wickerhamomyces spegazzinii Masiulionis & Pagnocca, Int. J. Syst. Evol. Microbiol. 66: 2144. 2016. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Hansenula sylviae Moschetti & J.P. Samp. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858178
For a description: see Francesca et al., Int. J. Syst. Evol. Microbiol. 63: 4828. 2013.
Holotype: CBS 12888, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ITALY, Sicily, island of Ustica, isolated from a trans-Saharan migratory bird (Sylvia communis), 2012.
Synonym: Wickerhamomyces sylviae Moschetti & J.P. Samp., Int. J. Syst. Evol. Microbiol. 63: 4828. 2013. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Liangdongia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858142
Etymology: the genus is named in honor of Liang-Dong Guo for his contribution of fungal taxonomy and ecology, especially of endophytic and mycorrhizal fungi.
Type species: Liangdongia dryadoides (D.B. Scott & van der Walt) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Starmera dryadoides clade, which occur in a separate clade distinct from the Starmera sensu stricto clade (Fig. 15). Member of the Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0007703 and OG0007714 (Table 6).
Both teleomorphic and anamorphic species occur. Asci are unconjugated, or originate from conjugation between independent cells or between a cell and its bud, with one to four hat-shaped or hemispheroidal ascospores. Colonies are white to cream, tannish-yellow, smooth, glistening and butyrous. Budding is multilateral. Pseudohyphae are present, but true hyphae are not observed. Where known, coenzyme Q-7 is formed.
Note: Liangdongia spp. are different from those of the closely related genus Starmera sensu stricto by their ability to grow on 0.1% cycloheximide (Table S3).
Species accepted:
Liangdongia berthetii (Boidin, Pignal, Mermiér & Arpin) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858143
Holotype: CBS 5452, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CAMEROON, isolated from gum arabic on a tree.
Basionym: Candida berthetii Boidin, Pignal, Mermiér & Arpin, Cah. Maboké 1: 100. 1963.
Synonym: Starmera berthetii (Boidin, Pignal, Mermiér & Arpin) H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4761. 2025.
Liangdongia dendrica (van der Walt, Klift & D.B. Scott) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858144
Holotype: CBS 6151, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. South Africa, isolated from frass of Cerambycidae larvae infesting Cryptocarya latifolia.
Basionym: Torulopsis dendrica van der Walt, Klift & D.B. Scott, Antonie van Leeuwenhoek 37: 461. 1971.
Synonyms: Candida dendrica (van der Walt, Klift & D.B. Scott) S.A. Mey. & Yarrow, Int. J. Syst. Bacteriol. 28: 612. 1978.
Starmera dendrica (Van der Walt, Klift & D.B. Scott) H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4761. 2025.
Liangdongia dryadoides (D.B. Scott & van der Walt) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858145
Holotype: CBS 6154, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from tunnels of Platypus externedentatus in Ficus sycomorus and Ficus hippopotami.
Basionym: Hansenula dryadoides D.B. Scott & van der Walt, Antonie van Leeuwenhoek 37: 171. 1971.
Synonym: Starmera dryadoides (D.B. Scott & van der Walt) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 951. 2008.
Liangdongia ilhagrandensis (C.G. Morais, A.R.O. Santos, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858146
Holotype: CBS 16316, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, Atlantic Rain Forest, isolated from rotting wood.
Basionym: Starmera ilhagrandensis C.G. Morais, A.R.O. Santos, Lachance & C.A. Rosa, Int. J. Syst. Evol. Microbiol. 70: 4382. 2020.
Liangdongia laemsonensis Am-In, Limtong, Yongman. & Jindam. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858147
For a description: see Am-In et al., Int. J. Syst. Evol. Microbiol. 61: 458. 2011.
Holotype: BCC 35154, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Ranong province, Laem Son National Park, isolated from estuarine water collected from a mangrove forest.
Synonyms: Candida laemsonensis Am-In, Limtong, Yongman. & Jindam., Int. J. Syst. Evol. Microbiol. 61: 458. 2011. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Starmera laemsonensis Am-In, Limtong, Yongman. & Jindam. ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4762. 2025.
Liangdongia nongkratonensis (Nakase & Jindam.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858148
Holotype: BCC 11772, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Nakhonratchasima province, Nong Kratone, isolated from insect frass in a tropical rain forest, February 2001.
Basionym: Pichia nongkratonensis Nakase & Jindam., Mycoscience 46: 193. 2005.
Synonym: Starmera nongkratonensis (Nakase & Jindam.) H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4762. 2025.
Liangdongia prunicorticola (Y.J. Qiu, H.Y. Zhu & F.Y. Bai) Q.M. Wang, comb. nov.
MycoBank: MB862244
Holotype: CGMCC 2.8558, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hubei province, Shennongjia Forestry District, Tangchaogumu, isolated from bark of a Prunus brachypoda tree, April, 2024.
Basionym: Starmera prunicorticola Y.J. Qiu, H.Y. Zhu & F.Y. Bai, Mycosphere 16(1): 4759. 2025.
Note: The ITS+LSU D1/D2 sequence analysis showed that S. prunicorticola was closely related to Starmera quercuum (Qiu et al. 2025), indicating that S. prunicorticola belongs to the Starmera dryadoides clade; therefore, this species was transferred to the genus Liangdongia.
Liangdongia quercuum (Phaff & E.P. Knapp) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858149
Holotype: CBS 2283, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, California, Mather, isolated from slime flux of a black oak (Quercus kelloggii).
Basionym: Pichia quercuum Phaff & E.P. Knapp [as ‘quercibus’], Antonie van Leeuwenhoek 22: 126. 1956.
Synonym: Starmera quercuum (Phaff & E.P. Knapp) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 951. 2008.
Reinstated genus
Petasospora Boidin & Abadie, Bull. Trimestriel Soc. Mycol. France 70: 364. 1955. [1954]
MycoBank: MB3845
Type species: Petasospora rhodanensis (C. Ramírez & Boidin) Boidin & Abadie, Bull. Trimestriel Soc. Mycol. France 70: 365. 1955. [1954]
Species accepted:
Petasospora adriatica (Čadež, Cardinali, Ciafardini & G. Péter) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858182
Holotype: CBS 12504, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SLOVENIA, Koper, isolated from the sediment of extra virgin olive oil.
Basionym: Candida adriatica Čadež, Cardinali, Ciafardini & G. Péter, Int. J. Syst. Evol. Microbiol. 62: 2299. 2012.
Synonym: Cyberlindnera adriatica (Čadež, Cardinali, Ciafardini & G. Péter) H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Petasospora americana (Wick.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858183
Holotype: NRRL Y-2156, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, California, isolated from frass of Insecta, under bark of Pinus jeffreyi.
Basionym: Hansenula bimundalis var. americana Wick., Mycopathol. Mycol. Appl. 26(1): 97. 1965.
Synonym: Cyberlindnera americana (Wick.) Minter, Mycotaxon 110: 473. 2009.
Petasospora amylophila (Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858184
Holotype: NRRL YB-1287, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Michigan, isolated from frass in insect tunnels in loblolly pine (Pinus taeda).
Basionym: Pichia amylophila Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson, Int. J. Syst. Bacteriol. 30(1): 209. 1980.
Synonym: Cyberlindnera amylophila (Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson) Minter, Mycotaxon 110: 473. 2009.
Petasospora bimundalis (Wick. & Santa María) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858185
Holotype: NRRL Y-5343, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. SPAIN, isolated from larva of the beetle Ergates faber in Scotch pine (Pinus sylvestris).
Basionym: Hansenula bimundalis Wick. & Santa María, Mycopathol. Mycol. Appl. 26(1): 96. 1965.
Synonym: Cyberlindnera bimundalis (Wick. & Santa María) Minter, Mycotaxon 110: 474. 2009.
Petasospora dasilvae (K.O. Barros, R.M. Souza, Palladino, R.M. Cadete, A.R.O. Santos, Góes-Neto, Berkov, Zilli, M.J.S. Vital, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858186
Holotype: CBS 16129, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. BRAZIL, Roraima, São João da Baliza, isolated from rotting wood.
Basionym: Cyberlindnera dasilvae K.O. Barros, R.M. Souza, Palladino, R.M. Cadete, A.R.O. Santos, Góes-Neto, Berkov, Zilli, M.J.S. Vital, Lachance & C.A. Rosa, Int. J. Syst. Evolut. Biol. 71(9, no. 4986): 7. 2021.
Petasospora easanensis Jindam., Thuy & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858187
For a description: see Jindamorakot et al., J. Gen. Appl. Microbiol. 50: 263. 2004.
Holotype: JCM 12476, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. THAILAND, Amnat Charoen, Nong Laung, isolated from insect frass, February 1997.
Synonym: Candida easanensis Jindam., Thuy & Nakase, J. Gen. Appl. Microbiol. 50: 263. 2004. Nom. inval., Art. 40.6 (Melbourne).
Cyberlindnera easanensis Jindam., Thuy & Nakase ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Petasospora euphorbiae (van der Walt & A. Opperman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858188
Holotype: CBS 8033, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, Groblersdal, isolated from the insect-infested spurge (Euphorbia ingens).
Basionym: Pichia euphorbiae Van der Walt & A. Opperman, Antonie van Leeuwenhoek 49: 55. 1983.
Synonym: Cyberlindnera euphorbiae (van der Walt & A. Opperman) Minter, Mycotaxon 110: 474. 2009.
Petasospora euphorbiiphila (van der Walt) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858189
Holotype: CBS 8083, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, Transvaal, isolated from the insect-infested decaying tissue of spurge (Euphorbia ingens).
Basionym: Hansenula euphorbiiphila Van der Walt [as 'euphorbiaphila'], Antonie van Leeuwenhoek 48: 467. 1982.
Synonym: Cyberlindnera euphorbiiphila (Van der Walt) Minter, Mycotaxon 110: 474. 2009.
Petasospora fabianii (Wick.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858197
Holotype: NRRL Y-1871, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Illinois, isolated from a fermentor inoculated with Aerobacter aerogenes for the production of butylene glycol.
Basionym: Hansenula fabianii Wick., Mycopathol. Mycol. Appl. 26(1): 84. 1965.
Synonym: Cyberlindnera fabianii (Wick.) Minter, Mycotaxon 110: 474. 2009.
Petasospora hubeiensis (Y.J. Qiu, H.Y. Zhu & F.Y. Bai) Q.M. Wang, comb. nov.
MycoBank: MB862241
Holotype: CGMCC 2.8539, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hubei province, Shennongjia Forestry District, Tanjiawan, isolated from bark of a tree, July, 2023.
Basionym: Cyberlindnera hubeiensis Y.J. Qiu, H.Y. Zhu & F.Y. Bai, Mycosphere 16(1): 4732. 2025.
Note: The ITS+LSU D1/D2 sequence analysis showed C. hubeiensis was closely related to Cyberlindnera mycetangii, Cyberlindnera rhodanensis and Cyberlindnera rhizosphaerae (Qiu et al. 2025), indicating that C. hubeiensis belongs to the Petasospora clade; therefore, this species was transferred into the genus Petasospora.
Petasospora hungchunana C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858198
For a description: see Chang et al., Antonie van Leeuwenhoek 102: 17. 2012.
Holotype: CBS 12243, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Pingtung, Hengchun, isolated from decayed wood, October 2009.
Synonym: Candida hungchunana C.F. Lee, Antonie van Leeuwenhoek 102: 17. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Petasospora juglandicorticola (Y.J. Qiu, H.Y. Zhu & F.Y. Bai) Q.M. Wang, comb. nov.
MycoBank: MB862242
Holotype: CGMCC 2.8538, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hubei province, Shennongjia Forestry District, Duidongping, isolated from the bark of a Juglans tree, April, 2024.
Basionym: Cyberlindnera juglandicorticola Y.J. Qiu, H.Y. Zhu & F.Y. Bai, Mycosphere 16(1): 4734. 2025.
Note: The ITS+LSU D1/D2 sequence analysis showed that C. juglandicorticola was closely related to Cyberlindnera americana, Cyberlindnera bimundalis and Cyberlindnera nakhonratchasimensis (Qiu et al. 2025), indicating that C. juglandicorticola belongs to the Petasospora clade; therefore, this species was transferred into the genus Petasospora.
Petasospora japonica (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858199
Holotype: NRRL YB-2750, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. JAPAN, isolated from the frass of Abies firma.
Basionym: Pichia japonica Kurtzman, Mycologia 79(3): 413. 1987.
Synonym: Cyberlindnera japonica (Kurtzman) Minter, Mycotaxon 110: 474. 2009.
Petasospora maesae C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858200
For a description: see Chang et al., Antonie van Leeuwenhoek 102: 12. 2012.
Holotype: CBS 12240, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Kaohsiung, Maolin, isolated from the leaf of Maesa japonica, January 2008.
Synonym: Candida maesae C.F. Lee [as ‘maesa’], Antonie van Leeuwenhoek 102: 12. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Petasospora maritima (Siepmann) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858201
Holotype: CBS 5107, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. Atlantic Ocean, isolated from the eggs of shrimp.
Basionym: Trichosporon maritimum Siepmann, Veröff. Inst. Meeresf. Bremerhaven 8: 89. 1962.
Synonyms: Cyberlindnera maritima (Siepmann) Brysch-Herzb., Dlauchy, M. Seidel & G. Péter, Int. J. Syst. Evol. Microbiol. 71(2, no. 4477): 6. 2021.
Candida maritima (Siepmann) Uden & H.R. Buckley, Mycotaxon 17: 298. 1983.
Petasospora meyerae (van der Walt) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858202
Holotype: CBS 7076, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, Groblersdal, isolated from rotting insect-infested spurge (Euphorbia ingens).
Basionym: Pichia meyerae van der Walt, Antonie van Leeuwenhoek 48(4): 385. 1982.
Synonym: Cyberlindnera meyerae (van der Walt) Minter, Mycotaxon 110: 475. 2009.
Petasospora mississippiensis (Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858203
Holotype: NRRL YB-1294, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Mississippi, isolated from the insect frass, loblolly pine (Pinus taeda).
Basionym: Pichia mississippiensis Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson, Int. J. Syst. Bacteriol. 30: 212. 1980.
Synonym: Cyberlindnera mississippiensis (Kurtzman, M.J. Smiley, C.J. Johnson, Wick. & Fuson) Minter, Mycotaxon 110: 475. 2009.
Petasospora mycetangii (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858204
Holotype: NRRL Y-6843, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Kansas, isolated from the mycetangia of ambrosia beetles.
Basionym: Candida mycetangii Kurtzman, Int. J. Syst. Evol. Microbiol. 50: 400. 2000.
Synonym: Cyberlindnera mycetangii (Kurtzman) Brysch-Herzb., Dlauchy, M. Seidel & G. Péter, Int. J. Syst. Evol. Microbiol. 71(2, no. 4477): 6. 2021.
Petasospora nakhonratchasimensis Jindam. & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858205
For a description: see Jindamorakot et al., J. Gen. Appl. Microbiol. 50: 266. 2004.
Holotype: JCM 12474, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. THAILAND, isolated from the insect frass.
Synonyms: Candida nakhonratchasimensis Jindam. & Nakase, J. Gen. Appl. Microbiol. 50: 266. 2004. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Cyberlindnera nakhonratchasimensis Jindam. & Nakase ex Brysch-Herzb., Dlauchy, M. Seidel & G. Péter, Int. J. Syst. Evol. Microbiol. 71(2, no. 4477): 6. 2021. Nom. inval., Art. 40.6 (Melbourne), published as a combination, but 'basionym' (Candida nakhonratchasimensis) is invalid.
Petasospora pattaniensis Jindam., Duy & Nakase ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858206
For a description: see Jindamorakot et al., J. Gen. Appl. Microbiol. 50: 265. 2004.
Holotype: JCM 12475, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. THAILAND, isolated from the insect frass.
Synonym: Candida pattaniensis Jindam., Duy & Nakase, J. Gen. Appl. Microbiol. 50: 265. 2004. Nom. inval., Art. 40.6 (Melbourne).
Cyberlindnera pattaniensis Jindam., Duy & Nakase ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Petasospora petersonii (Wick.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858207
Holotype: NRRL YB-3808, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Illinois, isolated from the lung tissue of a human cadaver preserved with embalming fluid.
Basionym: Hansenula petersonii Wick., Mycologia 56(3): 404. 1964.
Synonym: Cyberlindnera petersonii (Wick.) Minter, Mycotaxon 110: 475. 2009.
Petasospora rhizosphaerae (Mestre, C.A. Rosa & S.B. Fontenla) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858208
Holotype: CRUB 1796, preserved as a lyophilized preparation in the Centro Regional Universitario Bariloche (CRUB), Bariloche, Argentina. ARGENTINA, Rio Negro, isolated from a Nothofagus pumilio forest.
Basionym: Lindnera rhizosphaerae Mestre, C.A. Rosa & S.B. Fontenla, Int. J. Syst. Evol. Microbiol. 61(4): 988. 2011.
Synonym: Cyberlindnera rhizosphaerae (Mestre, C.A. Rosa & S.B. Fontenla) P.M. Kirk & Offord, Index Fungorum 9: 1. 2012.
Petasospora rhodanensis (C. Ramírez & Boidin) Boidin & Abadie, Bull. Trimestriel. Soc. Mycol. France 70: 365. 1955. [1954]
MycoBank: MB302613
Basionym: Saccharomyces rhodanensis C. Ramírez & Boidin, Rev. Mycol. (Paris) 18(2): 152. 1953.
Synonym: Cyberlindnera rhodanensis (C. Ramírez & Boidin) Minter, Mycotaxon 110: 475. 2009.
Petasospora shennongjiaensis (Y.J. Qiu, H.Y. Zhu & F.Y. Bai) Q.M. Wang, comb. nov.
MycoBank: MB862243
Holotype: CGMCC 2.8543, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hubei province, Shennongjia Forestry District, Shennong Tianchi, isolated from pieces of wood/bark from a tree, April, 2024.
Basionym: Cyberlindnera shennongjiaensis Y.J. Qiu, H.Y. Zhu & F.Y. Bai, Mycosphere 16(1): 4735. 2025.
Note: The ITS+LSU D1/D2 sequence analysis showed that C. shennongjiaensis was closely related to Cyberlindnera mycetangii, Cyberlindnera maritima and Cyberlindnera sylvatica (Qiu et al. 2025), indicating that C. shennongjiaensis belongs to the Petasospora clade; therefore, this species was transferred into the genus Petasospora.
Petasospora stauntonica C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858209
For a description: see Chang et al., Antonie van Leeuwenhoek 102: 19. 2012.
Holotype: CBS 12241, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Kaohsiung, Taoyuan, isolated from the leaf of Stauntonia purpurea, May 2008.
Synonym: Candida stauntonica C.F. Lee, Antonie van Leeuwenhoek 102: 19. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Cyberlindnera stauntonica C.F. Lee ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Petasospora sylvatica (Brysch-Herzb., Dlauchy, M. Seidel & G. Péter) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858210
Holotype: CBS 16335, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. GERMANY, Württemberg, isolated from a slime flux of Fagus sylvatica, 2019.
Basionym: Cyberlindnera sylvatica Brysch-Herzb., Dlauchy, M. Seidel & G. Péter, Int. J. Syst. Evol. Microbiol. 71(2, no. 4477): 5. 2021.
Petasospora taoyuanica C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB862180
For a description: see Chang et al., Antonie van Leeuwenhoek 102(1): 16. 2012.
Holotype: CBS 12242, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Kaohsiung, Taoyuan, isolated from soil, May 2008.
Synonym: Candida taoyuanica C.F. Lee, Antonie van Leeuwenhoek 102(1): 16. 2012. Nom. inval., Art. 40.7 (Melbourne).
Cyberlindnera taoyuanica C.F. Lee ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Petasospora veronae (K. Kodama) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858211
Holotype: NRRL Y-7818, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA.
Basionym: Pichia veronae K. Kodama, J. Ferment. Technol. 52(9): 612. 1974.
Synonym: Cyberlindnera veronae (K. Kodama) Minter, Mycotaxon 110: 476. 2009.
Petasospora wuzhiensis (S.A. Wang & F.Y. Bai) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858212
Holotype: CGMCC 2.3480, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Hainan province, Wuzhi mountain, isolated from a leaf of herbaceous plant, November 2006.
Basionym: Lindnera wuzhiensis S.A. Wang & F.Y. Bai, J. Gen. Appl. Microbiol. 56: 411. 2011.
Synonym: Cyberlindnera wuzhiensis (S.A. Wang & F.Y. Bai) P.M. Kirk & Offord, Index Fungorum 9: 1. 2012.
Petasospora xishuangbannaensis (Jun Zheng, Y.F. Lu, X.J. Liu & F.L. Hui) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858213
Holotype: NYNU 16752, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Yunnan province, Mengla, Menglun, isolated from rotting wood.
Basionym: Cyberlindnera xishuangbannaensis Jun Zheng, Y.F. Lu, X.J. Liu & F.L. Hui, Int. J. Syst. Evol. Microbiol. 67(12): 5052. 2017.
Petasospora xylebori (Ninomiya, Mikata, H. Kajim. & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858214
Holotype: NBRC 11048, preserved in a metabolically inactive state at NITE Biological Resource Center (NBRC), Tokyo, Japan. JAPAN, Miyagi prefecture, isolated from a gallery in Fagus crenata infested by an ambrosia beetle (Xyleborus sp.).
Basionym: Cyberlindnera xylebori Ninomiya, Mikata, H. Kajim. & H. Kawas., Int. J. Syst. Evol. Microbiol. 63(7): 2708. 2013.
Petasospora xylosilytica (R.M. Cadete, C.F. Lee, Kurtzman, Zilli, M.J.S. Vital, Lachance & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858215
Holotype: NRRL YB-2097, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, South Carolina, isolated from wood frass.
Basionym: Cyberlindnera xylosilytica R.M. Cadete, C.F. Lee, Kurtzman, Zilli, M.J.S. Vital, Lachance & C.A. Rosa, Int. J. Syst. Evol. Microbiol. 65: 2970. 2015.
Ruyongia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858150
Etymology: the genus is named after Dr. Ru-Yong Zheng for her contribution to fungal taxonomy, especially of Mucoromycetes.
Type species: Ruyongia chambardii (C. Ramírez & Boidin) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Wickerhamomyces chambardii clade, which occurs in a separate clade from other lineages in the Phaffomycetales (Fig. 15). Member of the Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0007319 and OG0007880 (Table 6).
Both teleomorphic and anamorphic species occur. Asci are unconjugated and form one to four hat-shaped ascospores. Colonies are white to tannish-white, butyrous, glistening. Budding is multilateral. Pseudohyphae are present or not, but true hyphae are not observed. Where known, coenzyme Q-7 is formed.
Species accepted:
Ruyongia chambardii (C. Ramírez & Boidin) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858152
Holotype: CBS 1900, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ITALY, isolated from tanning liquor.
Basionym: Saccharomyces chambardii C. Ramírez & Boidin [as ‘chambardi’], Bull. Mens. Soc. Linn. Lyon 23: 152. 1954.
Synonym: Wickerhamomyces chambardii (C. Ramírez & Boidin) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Ruyongia mori (F.L. Hui, Liang Chen, X.Y. Chu, Niu & T. Ke) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858154
Holotype: NYNU 1216, preserved in a metabolically inactive state at Nanyang University, Henan province, China. CHINA, Henan province, Nanyan, isolated from the trunk of a white mulberry tree (Morus alba).
Basionym: Wickerhamomyces mori F.L. Hui, Liang Chen, X.Y. Chu, Niu & T. Ke, Int. J. Syst. Evol. Microbiol. 63: 1177. 2013.
Ruyongia namnaoensis (Nakase, Jindam., Am-In, Ninomiya & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858155
Holotype: BCC 15093, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Petchabun province, Nam Nao National Park, isolated from insect frass of an unidentified tree, November 2001.
Basionym: Candida namnaoensis Nakase, Jindam., Am-In, Ninomiya & H. Kawas., J. Gen. Appl. Microbiol. 58: 150. 2012.
Ruyongia patagonica (V. de García, Brizzio, C.A. Rosa, Libkind & van Broock) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858156
Holotype: CBS 11398, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. ARGENTINA, Patagonia, Nahuel Huapi National Park, isolated from sap exudate on cut branches of Nothofagus dombeyi and glacier meltwater river.
Basionym: Wickerhamomyces patagonicus V. de García, Brizzio, C.A. Rosa, Libkind & van Broock, Int. J. Syst. Evol. Microbiol. 60: 1695. 2010.
Ruyongia ponderosae (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858158
Holotype: NRRL YB-2307, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Washington, Twisp, Chelan National Forest, isolated from the frass of an unidentified beetle that had tunneled into a Ponderosa pine (Pinus ponderosa).
Basionym: Candida ponderosae Kurtzman, Antonie van Leeuwenhoek 79: 360. 2001.
Ruyongia rarassimilans (Endoh, M. Suzuki, Omoto & Benno) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858159
Holotype: JCM 14993, preserved in a metabolically inactive state at the Japan Collection of Microorganisms (JCM), Ibaraki, Japan. JAPAN, Kyoto Pref., isolated from the body surface of Platypus quercivorus, 2004.
Basionym: Pichia rarassimilans Endoh, M. Suzuki, Omoto & Benno, J. Gen. Appl. Microbiol. 54: 183. 2008.
Ruyongia tratensis (Nakase, Jindam., Am-In, Ninomiya & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858160
Holotype: BCC 15102, preserved in a metabolically inactive state at the BIOTEC Culture Collection (BCC), National Science and Technology Development Agency (NSTDA), Pathumthani, Thailand. THAILAND, Trat province, Waeru Mangrove Forest, isolated from a flower of Sonneratia caseolaris, January 2002.
Basionym: Wickerhamomyces tratensis Nakase, Jindam., Am-In, Ninomiya & H. Kawas., J. Gen. Appl. Microbiol. 58: 148. 2012.
Taiozyma Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858163
Etymology: the genus is named in honor of Dr. Fang-Lan Tai for his contribution to fungal taxonomy.
Type species: Taiozyma bovis (Uden & Carmo Souza) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Wickerhamomyces bovis clade, which occurs in a separate clade from other lineages in the Phaffomycetales and closely related to Phaffomycetaceae (Fig. 15). Member of the Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0006108 and OG0006424 (Table 6).
Both teleomorphic and anamorphic species occur. Asci unconjugated and generally with one to four hat-shaped ascospores. Colonies are tannish-white, creamy to brownish, butyrous, smooth. True hyphae are not observed. Pseudohyphae are present or not. Where known, coenzyme Q-7 is formed.
Species accepted:
Taiozyma bovis (Uden & Carmo Souza) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858164
Holotype: CBS 2616, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. PORTUGAL, isolated from the cecum of a cow.
Basionym: Pichia bovis Uden & Carmo Souza, J. Gen. Microbiol. 16: 385. 1957.
Synonym: Wickerhamomyces bovis (Uden & Carmo Souza) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Taiozyma dajiaensis C.F. Lee & C.H. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858165
For a description: see Liu et al., FEMS Yeast Res. 8: 818. 2008.
Holotype: CBS 10590, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Taichung, Dajiae, isolated from forest soil, 2006.
Synonym: Candida dajiaensis C.F. Lee & C.H. Liu, FEMS Yeast Res. 8: 818. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Taiozyma odintsovae (Babeva, Reshetova, Blagod. & Galimova) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858166
Holotype: CBS 6026, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. RUSSIA, isolated from sap of birch (Betula verrucosa).
Basionym: Candida odintsovae Babeva, Reshetova, Blagod. & Galimova, Mikrobiologiya 58: 632. 1989.
Taiozyma onychis (Yarrow) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858167
Holotype: CBS 5587, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. NETHERLANDS, isolated from nail infection of Homo sapiens.
Basionym: Pichia onychis Yarrow, Antonie van Leeuwenhoek 31: 465. 1965.
Synonym: Wickerhamomyces onychis (Yarrow) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Taiozyma peoriensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858168
Holotype: NRRL YB-1497, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Illinois, isolated from the stump of Ulmus.
Basionym: Candida peoriensis Kurtzman [as ‘peoriaensis’], Antonie van Leeuwenhoek 79: 359. 2001.
Taiozyma rabaulensis (Soneda & S. Uchida) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858169
Holotype: CBS 6797, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. PAPUA NEW GUINEA, New Britain, Rabaul, isolated from feces of an African snail.
Basionym: Pichia rabaulensis Soneda & S. Uchida, Bull. Natl. Sci. Mus. Tokyo, 14: 451. 1971.
Synonym: Wickerhamomyces rabaulensis (Soneda & S. Uchida) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Taiozyma yuanshanica C.F. Lee & Chun H. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858170
For a description: see Liu et al., FEMS Yeast Res. 8: 818. 2008.
Holotype: CBS 10589, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Ilan, Yuanshan, isolated from forest soil, 2006.
Synonym: Candida yuanshanica C.F. Lee & Chun H. Liu [as ‘yuanshanicus’], FEMS Yeast Res. 8: 818. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Reinstated genus
Waltiozyma H.B. Mull. & Kock, S. African J. Sci. 82: 491. 1986.
MycoBank: MB22415
Type species: Waltiozyma mucosa (Wick. & Kurtzman) H.B. Mull. & Kock
Species accepted:
Waltiozyma chaumierensis (M. Groenew., V. Robert & M.T. Sm.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858251
Holotype: CBS 8565, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. GUYANA, La Chaumiere, isolated from the surface of a flower.
Basionym: Wickerhamomyces chaumierensis M. Groenew., V. Robert & M.T. Sm., Int. J. Syst. Evol. Microbiol. 61: 2018. 2011.
Waltiozyma mucosa (Wick. & Kurtzman) H.B. Mull. & Kock, S. African J. Sci. 82: 491. 1986.
MycoBank: MB432175
Basionym: Pichia mucosa Wick. & Kurtzman, Mycologia 63: 1014. 1971.
Synonym: Wickerhamomyces mucosus (Wick. & Kurtzman) Kurtzman, Robnett & Basehoar-Powers [as ‘mucosa’], FEMS Yeast Res. 8: 952. 2008.
Waltiozyma pijperi (van der Walt & Tscheuschner) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858252
Holotype: CBS 2887, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. SOUTH AFRICA, isolated from buttermilk.
Basionym: Pichia pijperi van der Walt & Tscheuschner, Antonie van Leeuwenhoek 23: 189. 1957.
Synonym: Wickerhamomyces pijperi (van der Walt & Tscheuschner) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Waltiozyma solani (Lodder & Kreger-van Rij) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858253
Holotype: CBS 1908, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THE NETHERLANDS, isolated from the potato-starch mill.
Basionym: Candida solani Lodder & Kreger-van Rij, Yeasts, a taxonomic study, [Edn 1] (Amsterdam): 672. 1952.
Waltiozyma xylosica Limtong, Nitiyon, Kaewwich., Jindam., Am-In & Yongman ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858254
For a description: see Limtong et al., Int. J. Syst. Evol. Microbiol. 62: 2790. 2012.
Holotype: CBS 12320, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Phayao province, Mueang Phayao district, isolated from soil, December 2008.
Synonym: Wickerhamomyces xylosicus Limtong, Nitiyon, Kaewwich., Jindam., Am-In & Yongman. [as ‘xylosica’], Int. J. Syst. Evol. Microbiol. 62: 2790. 2012. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
New combinations and validated taxa in Wickerhamomyces
Wickerhamomyces jianshihensis C.F. Lee & C.H. Liu ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858179
For a description: see Liu et al., FEMS Yeast Res. 8: 820. 2008.
Holotype: CBS 10588, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Hsinchu, Jianshih, isolated from forest soil, 2006.
Synonym: Candida jianshihensis C.F. Lee & C.H. Liu, FEMS Yeast Res. 8: 820. 2008. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Wickerhamomyces quercuum (Nakase) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858180
Holotype: CBS 6422, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. JAPAN, isolated from the exudate of konara oak (Quercus serrata).
Basionym: Candida quercuum Nakase, J. Gen. Appl. Microbiol. 17(6): 476. 1971.
Wickerhamomyces ulmi (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858181
Holotype: CBS 8670, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, Illinois, isolated from insect frass on elm (Ulmus sp.).
Basionym: Candida ulmi Kurtzman, Int. J. Syst. Evol. Microbiol. 50: 402. 2000.
Reinstated genus
Williopsis Zender, Bull. Soc. Bot. Genève 17: 298. 1925.
MycoBank: MB5781
Type species: Williopsis saturnus (Klöcker) Zender.
Species accepted:
Williopsis culbertsonii Q.M. Wang, Hulfachor, K. Sylvester & Hittinger ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858256
For a description: see Sylvester et al., FEMS Yeast Res. 15: 10. 2015.
Holotype: CBS 13898, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, Michigan, isolated from soil, associated with Fagus grandifolia.
Synonym: Cyberlindnera culbertsonii Q.M. Wang, Hulfachor, K. Sylvester & Hittinger, FEMS Yeast Res. 15: 10. 2015. Nom. inval., Art. 40.7 (Melbourne), more than one collection in which the type is conserved was specified.
Williopsis dauci (A.M. Glushakova, M.A. Tomashevskaya & Kachalkin) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858283
Holotype: KBP Y-6686, preserved in a metabolically inactive state. RUSSIA, Moscow region, from a carrot sample bought on the local market, February 2020.
Basionym: Cyberlindnera dauci A.M. Glushakova, M.A. Tomashevskaya & Kachalkin, Persoonia 45: 343. 2020.
Williopsis galapagoensis (Guamán-Burneo, R.M. Cadete, P. Portero, E.J. Carvajal & C.A. Rosa) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858284
Holotype: CLQCA-24SC-025, preserved in a metabolically inactive state at the Catholic University Yeasts Collection (CLQCA), Galápagos. GALÁPAGOS, Santa Cruz Island, isolated from rotting wood.
Basionym: Cyberlindnera galapagoensis Guamán-Burneo, R.M. Cadete, P. Portero, E.J. Carvajal & C.A. Rosa, Antonie van Leeuwenhoek 108: 927. 2015.
Williopsis jadinii (Sartory, R. Sartory, J. Weill & J. Mey.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858257
Holotype: NRRL Y-1542, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. FRANCE, isolated from pus from a human abscess.
Basionym: Saccharomyces jadinii Sartory, R. Sartory, J. Weill & J. Mey., Compt. Rend. Hebd. Séances Acad. Sci. 194: 1688. 1932.
Synonyms: Cyberlindnera jadinii (Sartory, R. Sartory, J. Weill & J. Mey.) Minter, Mycotaxon 110: 474. 2009.
Candida utilis (Henneberg) Lodder & Kreger-van Rij, Yeasts, a taxonomic study, [Edn 1] (Amsterdam): 546. 1952.
Williopsis lachancei (Phaff, Starmer & Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858258
Holotype: NRRL Y-27008, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Hawaii, isolated from the rotting bark of Tetraplasandra hawaiiensis.
Basionym: Pichia lachancei Phaff, Starmer & Kurtzman, Int. J. Syst. Bacteriol. 49: 1296. 1999.
Synonym: Cyberlindnera lachancei (Phaff, Starmer & Kurtzman) Minter, Mycotaxon 110: 474. 2009.
Williopsis maclurae (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858259
Holotype: NRRL Y-5377, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, Illinois, Peoria, isolated from decaying fruits of the osage orange tree (Maclura pomifera).
Basionym: Pichia maclurae Kurtzman, Int. J. Syst. Evol. Microbiol. 50: 398. 2000.
Synonym: Cyberlindnera maclurae (Kurtzman) Minter, Mycotaxon 110: 475. 2009.
Williopsis mengyuniae (Jian He & Bo Chen) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858261
Holotype: CGMCC 2.3681, preserved in a metabolically inactive state at the China General Microbiological Culture Collection Center (CGMCC), Beijing, China. CHINA, Jiangsu province, isolated from sulfonylurea-contaminated soil, September 2006.
Basionym: Candida mengyuniae Jian He & Bo Chen, Int. J. Syst. Evol. Microbiol. 59: 1240. 2009.
Williopsis misumaiensis (Y. Sasaki & Tak. Yoshida ex Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858260
Holotype: NRRL Y-17389, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. JAPAN, isolated from orchard soil.
Basionym: Pichia misumaiensis Y. Sasaki & Tak. Yoshida ex Kurtzman, Int. J. Syst. Evol. Microbiol. 50: 399. 2000.
Synonym: Cyberlindnera misumaiensis (Y. Sasaki & Tak. Yoshida ex Kurtzman) Minter, Mycotaxon 110: 475. 2009.
Williopsis mrakii (Wick.) G.I. Naumov & Vustin, Dokl. Akad. Nauk S.S.S.R. 259: 721. 1981.
MycoBank: MB108625
Basionym: Hansenula mrakii Wick., Tech. Bull. U.S. Dep. Agric. 1029: 40. 1951.
Synonym: Cyberlindnera mrakii (Wick.) Minter, Mycotaxon 110: 475. 2009.
Williopsis samutprakarnensis Poomtien, Jindam., Limtong, Pinphan. & Thaniy. ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858263
For a description: see Poomtien et al., Antonie van Leeuwenhoek 103: 235. 2013.
Holotype: CBS 12528, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Samutprakarn province, Bangplee, isolated from the wastewater of a cosmetic factory, February 2009.
Synonym: Cyberlindnera samutprakarnensis Poomtien, Jindam., Limtong, Pinphan. & Thaniy., Antonie van Leeuwenhoek 103: 235. 2013. Nom. inval., Art. 40.7 (Melbourne).
Williopsis sargentensis (Wick. & Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858264
Holotype: NRRL YB-4139, preserved as a lyophilized preparation in the ARS Culture Collection, National Center for Agricultural Utilization Research, Peoria, IL, USA. USA, New Hampshire, Peterborough, isolated from the water of a small lake.
Basionym: Pichia sargentensis Wick. & Kurtzman, Mycologia 63: 1016. 1971.
Synonym: Cyberlindnera sargentensis (Wick. & Kurtzman) Minter, Mycotaxon 110: 476. 2009.
Williopsis saturnus (Klöcker) Zender, Bulletin de la Société Botanique de Genéve 17: 298. 1925.
MycoBank: MB277517
Basionym: Saccharomyces saturnus Klöcker, Meddel. Carlsberg Lab. 6: 77. 1903.
Synonyms: Cyberlindnera saturnus (Klöcker) Minter, Mycotaxon 110: 476. 2009.
Lindnera saturnus (Klöcker) Kurtzman, Robnett & Bas.-Powers, FEMS Yeast Res. 8: 951. 2008.
Williopsis suaveolens (Klöcker) G.I. Naumov, Vustin & Babeva, Mikrobiologiya 54: 242. 1985.
MycoBank: MB103900
Basionym: Pichia suaveolens Klöcker, Zentralblatt für Bakteriologie und Parasitenkunde, Abteilung 2 35: 371. 1912.
Synonyms: Cyberlindnera suaveolens (Klöcker) Minter, Mycotaxon 110: 476. 2009.
Lindnera suaveolens (Klöcker) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 951. 2008.
Williopsis saturnus var. suaveolens (Klöcker) Kurtzman, Antonie van Leeuwenhoek 60: 18. 1991.
Williopsis subsufficiens (Wick.) Vustin, G.I. Naumov, Babeva & T.I. Naumova, Dokl. Akad. Nauk SSSR 267: 1481-1484. 1982.
MycoBank: MB456651
Basionym: Hansenula saturnus var. subsufficiens Wick., Mycopath. Mycol. Appl. 37: 30. 1969.
Synonyms: Cyberlindnera subsufficiens (Wick.) Minter, Mycotaxon 110: 476. 2009.
Lindnera subsufficiens (Wick.) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 951. 2008.
Williopsis saturnus var. subsufficiens (Wick.) Kurtzman, Antonie van Leeuwenhoek 60: 18. 1991.
Williopsis takata C.F. Lee ex Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, sp. nov.
MycoBank: MB858267
For a description: see Chang et al., Antonie van Leeuwenhoek 102: 14. 2012.
Holotype: CBS 12244, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. CHINA, Taiwan province, Nantou, Shueili, isolated from soil, November 2007.
Synonyms: Candida takata C.F. Lee, Antonie van Leeuwenhoek 102: 14. 2012. Nom. inval., Art. 40.7 (Melbourne).
Cyberlindnera takata C.F. Lee ex H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4737. 2025.
Williopsis tropicalis (Boontham, Limtong, C.A. Rosa, M.R. Lopes, M.J.S. Vital & Srisuk) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858268
Holotype: CBS 14558, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. THAILAND, Samutprakarn province, isolated from the soil.
Basionym: Cyberlindnera tropicalis Boontham, Limtong, C.A. Rosa, M.R. Lopes, M.J.S. Vital & Srisuk, Int. J. Syst. Evol. Microbiol. 67: 2572. 2017.
Williopsis vartiovaarae (Capr.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858269
Holotype: CBS 4289, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. FINLAND, isolated from the soil.
Basionym: Torulopsis vartiovaarae Capr., Can. J. Microbiol. 7: 683. 1961.
Synonyms: Candida vartiovaarae (Capr.) Uden & H.R. Buckley [as 'vartriovaarai'], Mycotaxon 17: 298. 1983.
Cyberlindnera vartiovaarae (Capr.) H.Y. Zhu, Y.J. Qiu, P.J. Han & F.Y. Bai, Mycosphere 16(1): 4738. 2025.
Xingzhongia Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, gen. nov.
MycoBank: MB858171
Etymology: the genus is named in honor of Dr. Xing-Zhong Liu for his contribution to fungal taxonomy.
Type species: Xingzhongia hampshirensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai.
This genus is proposed for the species in the Wickerhamomyces hampshirensis clade, which occur as a separate clade from the genus Cyberlindnera, the Williopsis clade, and the single-species lineages Candida freyschussii (Figs. 15–16). Member of the Wickerhamomycetaceae (Phaffomycetales, Saccharomycetes). The genus is mainly circumscribed by phylogenomic analysis, the genome-based metrics AAI, POCP and PAPO (Table 1), and the presence of genus-specific protein families OG0012666 and OG0012668 (Table 6).
Asci are unconjugated, or result from conjugation between a cell and its bud, and they have one to four hat-shaped ascospores. Colonies are white to cream, slightly glistening and butyrous. Budding is multilateral. Pseudohyphae occur, but true hyphae do not. Where known, coenzyme Q-7 is formed.
Note: Xingzhongia spp. have lower GC% (36.81–39.18%) compared to its closely related genera, i.e., Cyberlindnera (39.64–51.79%) and Buckleya (42.53%).
Species accepted:
Xingzhongia hampshirensis (Kurtzman) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858172
Holotype: CBS 7208, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. USA, New Hampshire, Camp Sargent, isolated from frass in a dead, cut oak (Quercus sp.).
Basionym: Pichia hampshirensis Kurtzman, Mycologia 79: 412. 1987.
Synonym: Wickerhamomyces hampshirensis (Kurtzman) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Xingzhongia scolytoplatypi (Ninomiya, Mikata, H. Kajim. & H. Kawas.) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858173
Holotype: NBRC 11029, preserved in a metabolically inactive state at NITE Biological Resource Center (NBRC), Tokyo, Japan. JAPAN, Iwate Prefecture, isolated from the gallery of Scolytoplatypus shogun in Fagus crenata.
Basionym: Wickerhamomyces scolytoplatypi Ninomiya, Mikata, H. Kajim. & H. Kawas., Int. J. Syst. Evol. Microbiol. 63: 2708. 2013.
Xingzhongia strasburgensis (C. Ramírez & Boidin) Q.M. Wang, Boekhout, Yurkov, G. Péter & F.Y. Bai, comb. nov.
MycoBank: MB858174
Holotype: CBS 2939, preserved in a metabolically inactive state at the Westerdijk Institute, Utrecht, the Netherlands. FRANCE, isolated from tanned leather.
Basionym: Saccharomyces strasburgensis C. Ramírez & Boidin, Rev. Mycol. (Paris) 18: 154. 1953.
Synonym: Wickerhamomyces strasburgensis (C. Ramírez & Boidin) Kurtzman, Robnett & Basehoar-Powers, FEMS Yeast Res. 8: 952. 2008.
Acknowledgments
This study was supported by grants No. 31961133020 and No. 32370015 from the National Natural Science Foundation of China (NSFC), No. 2021FY100900 and 2023FY101300 from the Ministry of Science and Technology of China, No. 521000981388 from the Advanced Talents Incubation Program of Hebei University.
Author contributions
Wang QM and Boekhout T conceived and designed the project. Zhao XM and Liu F performed the phylogenomic and genomic metrics analyses. Liu MM, Bai J and Zhao YJ worked on the taxonomy and compared the phenotypic data. Liu JH, Zhang YX, Wang JC, Zhang XH, Cui TX, Liu ZQ and Li HZ edited the phylogenetic trees. Bensch K worked on nomenclatural matters. Blackwell M supplied the phenotypic data of Candida sp. BG02-7-18-018A-2-2. Wang QM, Boekhout T and Liu MM wrote the paper. Wang QM, Boekhout T, Yurkov A, Liu MM, Péter G and Bai FY revised and edited the paper. The authors are solely responsible for the content of this work.
ORCID
Miao-Miao Liu: https://orcid.org/0009-0001-1001-4612
Xue-Mei Zhao: https://orcid.org/0009-0004-0682-7436
Qi-Ming Wang: https://orcid.org/0000-0002-2180-8135
Conflict of interest statement
The authors declare no other competing interests.
Data availability
The tree and sequence alignment files, and the genome (or subgenome) datasets for calculating the impact of reductive evolution, HTGs, hybridization (or alloaneuploidy) and introgressions on the values of genomic metrics are deposited in the Figshare repository: https://doi.org/10.6084/m9.figshare.28558733.
Supplementary Information
The online version contains supplemental information available at https://doi.org/10.65390/fdiv.2026.136006.
Rights and permissions
The Author(s) 2026. Published by BioAcademic Press on behalf of Kunming Institute of Botany,Chinese Academy of Sciences (CAS) and Mushroom Research Foundation. This is an open accessarticle under the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0), which permits use, distribution and reproduction in any medium, provided the originalwork is properly cited.
References
- Aphalo PJ (2025) ggpmisc: miscellaneous extensions to ‘ggplot2’.
https://doi.org/10.32614/CRAN.package.ggpmisc - Avesani M, Zapparoli G, Jindamorakot S, Limtong S (2024) Yamadazyma oleae f.a. sp. nov. and Yamadazyma molendinolei f.a. sp. nov., two novel ascomycetous yeast species isolated from olive oil mills in Italy, and reassignment of 11 Candida species to the genus Yamadazyma. International Journal of Systematic and Evolutionary Microbiology 74.
https://doi.org/10.1099/ijsem.0.006592 - Bab'eva IP, Lisichkina GA, Maksimova IA, Reshetova IS, Terenina EE, Iu Chernov I (2000) A new yeast species Candida anutae sp. nov. from the fruiting bodies of agaricus. Microbiology 69: 225–228.
https://doi.org/10.1007/BF02756204 - Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA et al. (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology 19: 455–477.
https://doi.org/10.1089/cmb.2012.0021 - Berkhout CM (1923) De schimmelgeslachten Monilia, Oidium, Oospora en Torula. Dissertation, University of Utrecht.
- Bhunjun CS, Chen YJ, Phukhamsakda C, Boekhout T, Groenewald JZ, McKenzie EHC, Francisco EC, Frisvad JC, Groenewald M, Hurdeal VG, Luangsa-Ard J, Perrone G, Visagie CM, Bai FY, Błaszkowski J et al. (2024) What are the 100 most cited fungal genera? Studies in Mycology 108: 1–411.
https://doi.org/10.3114/sim.2024.108.01 - Bhunjun CS, Niskanen T, Suwannarach N, Wannathes N, Chen YJ, McKenzie EHC, Maharachchikumbura SSN, Buyck B, Zhao CL, Fan YG, Zhang JY, Dissanayake AJ, Marasinghe DS, Jayawardena RS, Kumla J et al. (2022) The numbers of fungi: are the most speciose genera truly diverse? Fungal Diversity 114: 387–462.
https://doi.org/10.1007/s13225-022-00501-4 - Billon-Grand G (1989) A new ascosporogenous yeast genus: Yamadazyma gen. nov. Mycotaxon 35: 201–204.
- Boekhout T, Aime MC, Begerow D, Gabaldón T, Heitman J, Kemler M, Khayhan K, Lachance MA, Louis EJ, Sun S, Vu D, Yurkov A (2021) The evolving species concepts used for yeasts: from phenotypes and genomes to speciation networks. Fungal Diversity 109: 27–55.
https://doi.org/10.1007/s13225-021-00475-9 - Boidin J, Abadie F (1954) Les levures des liqueurs tannantes végétales; leur action sur les tanins pyrogalliques. Bulletin de la Société mycologique de France 70: 353–383.
- Buchfink B, Reuter K, Drost HG (2021) Sensitive protein alignments at tree-of-life scale using DIAMOND. Nature Methods 18: 366–368.
https://doi.org/10.1038/s41592-021-01101-x - Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) BLAST+: architecture and applications. BMC Bioinformatics 10: 421.
https://doi.org/10.1186/1471-2105-10-421 - Chai CY, Gao WL, Yan ZL, Hui FL (2022) Four new species of Trichomonascaceae (Saccharomycetales, Saccharomycetes) from Central China. MycoKeys 90: 1–18.
https://doi.org/10.3897/mycokeys.90.83829 - Chen S, Zhou Y, Chen Y, Gu J (2018) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34: i884–i890.
https://doi.org/10.1093/bioinformatics/bty560 - Crous PW, Wingfield MJ, Burgess TI, Hardy GESJ, Barber PA, Alvarado P, Barnes CW, Buchanan PK, Heykoop M, Moreno G, Thangavel R, van der Spuy S, Barili A, Barrett S, Cacciola SO et al. (2017) Fungal Planet description sheets: 558–624. Persoonia 38: 240–384.
https://doi.org/10.3767/003158517X698941 - D'Angiolo M, De Chiara M, Yue JX, Irizar A, Stenberg S, Persson K, Llored A, Barré B, Schacherer J, Marangoni R, Gilson E, Warringer J, Liti G (2020) A yeast living ancestor reveals the origin of genomic introgressions. Nature 587: 420–425.
https://doi.org/10.1038/s41586-020-2889-1 - Daniel HM, Lachance MA, Kurtzman CP (2014) On the reclassification of species assigned to Candida and other anamorphic ascomycetous yeast genera based on phylogenetic circumscription. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 106: 67–84.
https://doi.org/10.1007/s10482-014-0170-z - Daniel HM, Redhead SA, Schnürer J, Naumov GI, Kurtzman CP (2012) (2049-2050) Proposals to conserve the name Wickerhamomyces against Hansenula and to reject the name Saccharomyces sphaericus (Ascomycota: Saccharomycotina). TAXON 61: 459–461.
https://doi.org/10.1002/tax.612015 - de Vega C, Albaladejo RG, Guzmán B, Steenhuisen SL, Johnson SD, Herrera CM, Lachance MA (2017) Flowers as a reservoir of yeast diversity: description of Wickerhamiella nectarea f.a. sp. nov., and Wickerhamiella natalensis f.a. sp. nov. from South African flowers and pollinators, and transfer of related Candida species to the genus Wickerhamiella as new combinations. FEMS Yeast Research 17.
https://doi.org/10.1093/femsyr/fox054 - Draper NR, Smith H (1998) Applied regression analysis. Wiley Series in Probability and Statistics.
- Emms DM, Kelly S (2019) OrthoFinder: phylogenetic orthology inference for comparative genomics. Genome Biology 20: 238.
https://doi.org/10.1186/s13059-019-1832-y - Felsenstein J (1985) Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39: 783–791.
https://doi.org/10.1111/j.1558-5646.1985.tb00420.x - García-Acero AM, Batista TM, Souza GFL, Santos ARO, Souza DL, Franco GR, Velásquez-Lozano ME, Yamamoto D, Toki W, Lachance MA, Rosa CA (2024) Description of Millerago gen. nov. based on taxonomic analysis, with two new species, Millerago phaffii f.a., sp. nov. and Millerago galiae f.a., sp. nov. International Journal of Systematic and Evolutionary Microbiology 74.
https://doi.org/10.1099/ijsem.0.006565 - Glushakova AM, Maximova IA, Kachalkin AV, Yurkov AM (2010) Ogataea cecidiorum sp. nov., a methanol-assimilating yeast isolated from galls on willow leaves. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 98: 93–101.
https://doi.org/10.1007/s10482-010-9433-5 - Gonçalves C, Marques M, Gonçalves P (2022) Contrasting strategies for sucrose utilization in a floral yeast clade. mSphere 7: e0003522.
https://doi.org/10.1128/msphere.00035-22 - Gonçalves P, Gonçalves C, Brito PH, Sampaio JP (2020) The Wickerhamiella/Starmerella clade-A treasure trove for the study of the evolution of yeast metabolism. Yeast 37: 313–320.
https://doi.org/10.1002/yea.3463 - Gouliamova DE, Dimitrov RA, Smith MT (2016) DNA barcoding revealed Nematodospora valgi gen. nov., sp. nov. and Candida cetoniae sp. nov. in the Lodderomyces clade. Fungal Biology 20: 179–190.
https://doi.org/10.1016/j.funbio.2015.05.008 - Groenewald M, Hittinger CT, Bensch K, Opulente DA, Shen XX, Li Y, Liu C, LaBella AL, Zhou X, Limtong S, Jindamorakot S, Gonçalves P, Robert V, Wolfe KH, Rosa CA et al. (2023) A genome-informed higher rank classification of the biotechnologically important fungal subphylum Saccharomycotina. Studies in Mycology 105: 1–22.
https://doi.org/10.3114/sim.2023.105.01 - Hawksworth DL (2011) A new dawn for the naming of fungi: impacts of decisions made in Melbourne in July 2011 on the future publication and regulation of fungal names. International Mycological Association Fungus 2: 155–162.
https://doi.org/10.5598/imafungus.2011.02.02.06 - Holland SL, Dyer PS, Bond CJ, James SA, Roberts IN, Avery SV (2011) Candida argentea sp. nov., a copper and silver resistant yeast species. Fungal Biology 115: 909–918.
https://doi.org/10.1016/j.funbio.2011.07.004 - Hyde KD, Noorabadi MT, Thiyagaraja V, He MQ, Johnston PR, Wijesinghe SN, Armand A, Biketova AY, Chethana KWT, Erdoğdu M, Ge ZW, Groenewald JZ, Hongsanan S, Kušan I, Leontyev DV et al. (2024) The 2024 Outline of Fungi and fungus-like taxa. Mycosphere 15: 5146–6239.
https://doi.org/10.5943/mycosphere/15/1/25 - Jensen RH, Arendrup MC (2011) Candida palmioleophila: characterization of a previously overlooked pathogen and its unique susceptibility profile in comparison with five related species. Journal of Clinical Microbiology 49: 549–556.
https://doi.org/10.1128/JCM.02071-10 - Jensen RH, Astvad KM, Silva LV, Sanglard D, Jørgensen R, Nielsen KF, Mathiasen EG, Doroudian G, Perlin DS, Arendrup MC (2015) Stepwise emergence of azole, echinocandin and amphotericin B multidrug resistance in vivo in Candida albicans orchestrated by multiple genetic alterations. Journal of Antimicrobial Chemotherapy 70: 2551–2555.
https://doi.org/10.1093/jac/dkv140 - Katiyar S, Pfaller M, Edlind T (2006) Candida albicans and Candida glabrata clinical isolates exhibiting reduced echinocandin susceptibility. Antimicrobial Agents and Chemotherapy 50: 2892–2894.
https://doi.org/10.1128/AAC.00349-06 - Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30: 772–780.
https://doi.org/10.1093/molbev/mst010 - Kucukates E, Gultekin NN, Alisan Z, Hondur N, Ozturk R (2016) Identification of Candida species and susceptibility testing with Sensititre YeastOne microdilution panel to 9 antifungal agents. Saudi Medical Journal 37: 750–757.
https://doi.org/10.15537/smj.2016.7.13412 - Kudryavtsev VI (1960) Die Systematik der Hefen. Akademie Verlag, Berlin.
- Kurtzman CP (1984a) The yeasts, a taxonomic study. In: Kreger-van Rij NJW (ed) Hansenula H. et P. Sydow, 3rd edn. Elsevier, Amsterdam, pp 165–213.
https://doi.org/10.1016/B978-0-444-80421-1.50010-3 - Kurtzman CP (1984b) The yeasts, a taxonomic study. In: Kreger-van Rij NJW (ed) Pichia Hansen, 3rd edn. Elsevier, Amsterdam, pp 295–378.
https://doi.org/10.1016/B978-0-444-80421-1.50010-3 - Kurtzman CP (1984c) Synonymy of the yeast genera Hansenula and Pichia demonstrated through comparisons of deoxyribonucleic acid relatedness. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 50: 209–217.
https://doi.org/10.1007/BF02342132 - Kurtzman CP (1998) The yeasts, a taxonomic study. In: Kurtzman CP and Fell JW (ed) Pichia E.G. Hansen emend. Kurtzman, 4th edn. Elsevier, Amsterdam, pp 273–352.
https://doi.org/10.1016/B978-044481312-1/50046-0 - Kurtzman CP (2003) Phylogenetic circumscription of Saccharomyces, Kluyveromyces and other members of the Saccharomycetaceae, and the proposal of the new genera Lachancea, Nakaseomyces, Naumovia, Vanderwaltozyma and Zygotorulaspora. FEMS Yeast Research 4: 233–245.
https://doi.org/10.1016/S1567-1356(03)00175-2 - Kurtzman CP (2006) New species and new combinations in the yeast genera Kregervanrija gen. nov., Saturnispora and Candida. FEMS Yeast Research 6: 288–297.
https://doi.org/10.1111/j.1567-1364.2005.00019.x - Kurtzman CP (2011a) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Discussion of teleomorphic and anamorphic ascomycetous yeasts and yeast-like taxa, 5th edn. Elsevier, The Netherland, pp 293–307.
https://doi.org/10.1016/B978-0-444-52149-1.00013-6 - Kurtzman CP (2011b) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Babjeviella Kurtzman & Suzuki M (2010), 5th edn. Elsevier, The Netherlands, pp 329–331.
https://doi.org/10.1016/B978-0-444-52149-1.00017-3 - Kurtzman CP (2011c) Phylogeny of the ascomycetous yeasts and the renaming of Pichia anomala to Wickerhamomyces anomalus. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 99: 13–23.
https://doi.org/10.1007/s10482-010-9505-6 - Kurtzman CP (2011d) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Ogataea Y. Yamada, K. Maeda & Mikata (1994), 5th edn. Elsevier, The Netherland, pp 645–671.
https://doi.org/10.1016/B978-0-444-52149-1.00070-7 - Kurtzman CP (2016) Description of Groenewaldozyma gen. nov. for placement of Candida auringiensis, Candida salmanticensis and Candida tartarivorans. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 109: 1041–1045.
https://doi.org/10.1007/s10482-016-0703-8 - Kurtzman CP, Albertyn J, Basehoar-Powers E (2007) Multigene phylogenetic analysis of the Lipomycetaceae and the proposed transfer of Zygozyma species to Lipomyces and Babjevia anomala to Dipodascopsis. FEMS Yeast Research 7: 1027–1034.
https://doi.org/10.1111/j.1567-1364.2007.00246.x - Kurtzman CP, Fell JW (1998) The yeasts, a taxonomic study. Elsevier, The Netherlands.
- Kurtzman CP, Fell JW, Boekhout T (2011) In: The Yeasts, A Taxonomic Study (Kurtzman CP, Fell JW, Boekhout T, eds), 5th edn. Elsevier, The Netherlands.
- Kurtzman CP, Mateo RQ, Kolecka A, Theelen B, Robert V, Boekhout T (2015) Advances in yeast systematics and phylogeny and their use as predictors of biotechnologically important metabolic pathways. FEMS Yeast Research 15: fov050.
https://doi.org/10.1093/femsyr/fov050 - Kurtzman CP, Robnett CJ (1998) Identification and phylogeny of ascomycetous yeasts from analysis of nuclear large subunit (26S) ribosomal DNA partial sequences. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 73: 331–371.
https://doi.org/10.1023/a:1001761008817 - Kurtzman CP, Robnett CJ (2003) Phylogenetic relationships among yeasts of the ‘Saccharomyces complex’ determined from multigene sequence analyses. FEMS Yeast Research 3: 417–432.
https://doi.org/10.1016/S1567-1356(03)00012-6 - Kurtzman CP, Robnett CJ (2007) Multigene phylogenetic analysis of the Trichomonascus, Wickerhamiella and Zygoascus yeast clades, and the proposal of Sugiyamaella gen. nov. and 14 new species combinations. FEMS Yeast Research 7: 141–151.
https://doi.org/10.1111/j.1567-1364.2006.00157.x - Kurtzman CP, Robnett CJ (2010) Systematics of methanol assimilating yeasts and neighboring taxa from multigene sequence analysis and the proposal of Peterozyma gen. nov., a new member of the Saccharomycetales. FEMS Yeast Research 10: 353–361.
https://doi.org/10.1111/j.1567-1364.2010.00625.x - Kurtzman CP, Robnett CJ, Basehoar-Powers E (2008) Phylogenetic relationships among species of Pichia, Issatchenkia and Williopsis determined from multigene sequence analysis, and the proposal of Barnettozyma gen. nov., Lindnera gen. nov. and Wickerhamomyces gen. nov. FEMS Yeast Research 8: 939–954.
https://doi.org/10.1111/j.1567-1364.2008.00419.x - Kurtzman CP, Robnett CJ, Blackwell M (2016) Description of Teunomyces gen. nov. for the Candida kruisii clade, Suhomyces gen. nov. for the Candida tanzawaensis clade and Suhomyces kilbournensis sp. nov. FEMS Yeast Research 16: fow041.
https://doi.org/10.1093/femsyr/fow041 - Kurtzman CP, Suzuki M (2010) Phylogenetic analysis of ascomycete yeasts that form coenzyme Q-9 and the proposal of the new genera Babjeviella, Meyerozyma, Millerozyma, Priceomyces, and Scheffersomyces. Mycoscience 51: 2–14.
https://doi.org/10.1007/S10267-009-0011-5 - Kuzmanović N, Fagorzi C, Mengoni A, Lassalle F, diCenzo GC (2022) Taxonomy of Rhizobiaceae revisited: proposal of a new framework for genus delimitation. International Journal of Systematic and Evolutionary Microbiology 72: 005243.
https://doi.org/10.1099/ijsem.0.005243 - Lachance MA (2018) CP Kurtzman's evolving concepts of species, genus and higher categories. FEMS Yeast Research 18: foy103.
https://doi.org/10.1093/femsyr/foy103 - Lachance MA, Boekhout T, Scorzetti G, Fell JW Kurtzman CP (2011) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Candida Berkhout, 5th edn. Elsevier, The Netherlands, pp 987–1278.
https://doi.org/10.1016/B978-0-444-52149-1.00164-6 - Lee BK, Kim JK (2001) Production of Candida utilis biomass on molasses in different culture types. Aquacultural Engineering 25: 111–124.
https://doi.org/10.1016/S0144-8609(01)00075-9 - Li AH, Yuan FX, Groenewald M, Bensch K, Yurkov AM, Li K, Han PJ, Guo LD, Aime MC, Sampaio JP, Jindamorakot S, Turchetti B, Inacio J, Fungsin B, Wang QM et al. (2020) Diversity and phylogeny of basidiomycetous yeasts from plant leaves and soil: proposal of two new orders, three new families, eight new genera and one hundred and seven new species. Studies in Mycology 96: 17–140.
https://doi.org/10.1016/j.simyco.2020.01.002 - Limtong S, Koowadjanakul N, Jindamorakot S, Yongmanitchai W, Nakase T (2012) Candida sirachaensis sp. nov. and Candida sakaeoensis sp. nov. two anamorphic yeast species from phylloplane in Thailand. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 102: 221–229.
https://doi.org/10.1007/s10482-012-9728-9 - Liu F, Hu ZD, Yurkov A, Chen XH, Bao WJ, Ma Q, Zhao WN, Pan S, Zhao XM, Liu JH, Wang QM, Boekhout T (2024a) Saccharomycetaceae: delineation of fungal genera based on phylogenomic analyses, genomic relatedness indices and genomics-based synapomorphies. Persoonia 52: 1–21.
https://doi.org/10.3767/persoonia.2024.52.02 - Liu F, Hu ZD, Zhao XM, Zhao WN, Feng ZX, Yurkov A, Alwasel S, Boekhout T, Bensch K, Hui FL, Bai FY, Wang QM (2024b) Phylogenomic analysis of the Candida auris-Candida haemuli clade and related taxa in the Metschnikowiaceae, and proposal of thirteen new genera, fifty-five new combinations and nine new species. Persoonia 52: 22–43.
https://doi.org/10.3767/persoonia.2024.52.01 - Liu XJ, Yi ZH, Ren YC, Li Y, Hui FL (2016) Five novel species in the Lodderomyces clade associated with insects. International Journal of Systematic and Evolutionary Microbiology 66: 4881–4889.
https://doi.org/10.1099/ijsem.0.001446 - Liu Z, Kurtzman CP (1991) Phylogenetic relationships among species of Williopsis and Saturnospora gen. nov. as determined from partial rRNA sequences. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 60: 21–30.
https://doi.org/10.1007/BF00580437 - Li Y, Steenwyk JL, Chang Y, Wang Y, James TY, Stajich JE, Spatafora JW, Groenewald M, Dunn CW, Hittinger CT, Shen XX, Rokas A (2021) A genome-scale phylogeny of the kingdom fungi. Current Biology 31: 1653–1665.
https://doi.org/10.1016/j.cub.2021.01.074 - Lopes MR, Santos ARO, Moreira JD, Santa-Brígida R, Martins MB, Pinto FO, Valente P, Morais PB, Jacques N, Grondin C, Casaregola S, Lachance MA, Rosa CA (2019) Kurtzmaniella hittingeri f.a. sp. nov., isolated from rotting wood and fruits, and transfer of three Candida species to the genus Kurtzmaniella as new combinations. International Journal of Systematic and Evolutionary Microbiology 69: 1504–1508.
https://doi.org/10.1099/ijsem.0.003337 - Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD, Lopez R (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Research 268: 1–6.
https://doi.org/10.1093/nar/gkz268 - Malimas T, Huong VTL, Yukphan P, Tanasupawat S, Yamada Y (2023) The genus Hansenula Sydow et Sydow (1919) nom. rev. and the genus Wickerhamomyces Kurtzman et al. (2008). International Journal Novel Research Life Science 10: 42–47.
https://doi.org/10.13140/rg.2.2.11568.12804 - Manni M, Berkeley MR, Seppey M, Simão FA, Zdobnov EM (2021) BUSCO update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Molecular Biology and Evolution 38: 4647–4654.
https://doi.org/10.1093/molbev/msab199 - Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution 37: 1530–1534.
https://doi.org/10.1093/molbev/msaa015 - Minter DW (2009) Cyberlindnera, a replacement name for Lindnera Kurtzman et al., nom. illegit. Mycotaxon 110: 473–476.
https://doi.org/10.5248/110.473 - Miura Y, Kondo K, Saito T, Shimada H, Fraser PD, Misawa N (1998) Production of the carotenoids lycopene, beta-carotene, and astaxanthin in the food yeast Candida utilis. Applied and Environmental Microbiology 64: 1226–1229.
https://doi.org/10.1128/AEM.64.4.1226-1229.1998 - Monerawela C, Bond U (2018) The hybrid genomes of Saccharomyces pastorianus: A current perspective. Yeast 35: 39–50.
https://doi.org/10.1002/yea.3250 - Montecillo JAV (2023) Phylogenomics and comparative genomic analyses support the creation of the novel family Ignatzschineriaceae fam. nov. comprising the genera Ignatzschineria and Wohlfahrtiimonas within the order Cardiobacteriales. Research in Microbiology 174: 103988.
https://doi.org/10.1016/j.resmic.2022.103988 - Morais CG, Batista TM, Kominek J, Borelli BM, Furtado C, Moreira RG, Franco GR, Rosa LH, Fonseca C, Hittinger CT, Lachance MA, Rosa CA (2017) Spathaspora boniae sp. nov., a D-xylose-fermenting species in the Candida albicans/Lodderomyces clade. International Journal of Systematic and Evolutionary Microbiology 67: 3798–3805.
https://doi.org/10.1099/ijsem.0.002186 - Muller HB, Kock JLF (1986) Waltiozyma gen. nov. (Saccharomycetaceae), a new genus of the Endomycetales. South African Journal of Science 82: 491–492.
- Nakase T, Jindamorakot S, Ninomiya S, Imanishi Y, Kawasaki H. (2009) Candida wancherniae sp. nov. and Candida morakotiae sp. nov., two novel ascomycetous anamorphic yeast species found in Thailand. Journal of General and Applied Microbiology 55: 93–100.
https://doi.org/10.2323/jgam.55.93 - Naumova ES, Naumov GI, Nosek J, Tomáska L (2004) Differentiation of the yeasts Williopsis, Zygowilliopsis and Komagataea by karyotypic and PCR analyses. Systematic and Applied Microbiology 27: 192–197.
https://doi.org/10.1078/072320204322881817 - Nguyen NH, Suh SO, Blackwell M (2007) Five novel Candida species in insect-associated yeast clades isolated from Neuroptera and other insects. Mycologia 99: 842–858.
https://doi.org/10.3852/mycologia.99.6.842 - Novák EK, Zsolt J (1961) A new system proposed for yeasts. Acta Biologica Academiae Scientiarum Hungaricae 7: 93–145.
- Nowell RW, Almeida P, Wilson CG, Smith TP, Fontaneto D, Crisp A, Micklem G, Tunnacliffe A, Boschetti C, Barraclough TG (2018) Comparative genomics of bdelloid rotifers: Insights from desiccating and nondesiccating species. PLoS Biology 16: e2004830.
https://doi.org/10.1371/journal.pbio.2004830 - Opulente DA, LaBella AL, Harrison MC, Wolters JF, Liu C, Li Y, Kominek J, Steenwyk JL, Stoneman HR, VanDenAvond J, Miller CR, Langdon QK, Silva M, Gonçalves C, Ubbelohde EJ et al. (2024) Genomic factors shape carbon and nitrogen metabolic niche breadth across Saccharomycotina yeasts. Science 384: eadj4503.
https://doi.org/10.1126/science.adj4503 - Pereira GH, Müller PR, Szeszs MW, Levin AS, Melhem MS (2010) Five-year evaluation of bloodstream yeast infections in a tertiary hospital: the predominance of non-C. albicans Candida species. Medical Mycology 48: 839–842.
https://doi.org/10.3109/13693780903580121 - Pérez-Través L, Lopes CA, Querol A, Barrio E (2014) On the complexity of the Saccharomyces bayanus taxon: hybridization and potential hybrid speciation. PLoS One 9: e93729.
https://doi.org/10.1371/journal.pone.0093729 - Peris D, Lopes CA, Belloch C, Querol A, Barrio E (2012) Comparative genomics among Saccharomyces cerevisiae × Saccharomyces kudriavzevii natural hybrid strains isolated from wine and beer reveals different origins. BMC Genomics 13: 407.
https://doi.org/10.1186/1471-2164-13-407 - Pontes A, Paraíso F, Silva M, Lagoas C, Aires A, Brito PH, Rosa CA, Lachance MA, Sampaio JP, Gonçalves C, Gonçalves P (2024) Extensive remodeling of sugar metabolism through gene loss and horizontal gene transfer in a eukaryotic lineage. BMC Biology 22: 128.
https://doi.org/10.1186/s12915-024-01929-7 - Qin QL, Xie BB, Zhang XY, Chen XL, Zhou BC, Zhou J, Oren A, Zhang YZ (2014) A proposed genus boundary for the prokaryotes based on genomic insights. Journal of Bacteriology 196: 2210–2215.
https://doi.org/10.1128/JB.01688-14 - Qiu YJ, Zhu HY, Zhu QY, Yang LX, Mao YC, Wen Z, Chen SX, Qiu JZ, Bai FY, Han PJ (2025) High yeast diversity in primeval forest of Shennongjia, including 21 new species characterized by morphological, phylogenetic, and genomic analyses. Mycosphere 16: 4723–4782.
https://doi.org/10.5943/mycosphere/16/1/35 - Ramírez C, J Boidin (1953) Nuevas especies de levaduras aisladas de liquidos curtientes. Microbiology 4: 405–414.
- R Core Team (2025) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna.
- Ren YC, Liu XJ, Yi ZH, Hui FL (2016) Nematodospora anomalae sp. nov., a novel and D-xylose-fermenting yeast species in the Lodderomyces clade. International Journal of Systematic and Evolutionary Microbiology 66: 4046–4050.
https://doi.org/10.1099/ijsem.0.001308 - Riesco R, Trujillo ME (2024) Update on the proposed minimal standards for the use of genome data for the taxonomy of prokaryotes. International Journal of Systematic and Evolutionary Microbiology 74: 006300.
https://doi.org/10.1099/ijsem.0.006300 - Rosa CA, Lachance MA (1998) The yeast genus Starmerella gen. nov. and Starmerella bombicola sp. nov., the teleomorph of Candida bombicola (Spencer, Gorin & Tulloch) Meyer & Yarrow. International Journal of Systematic Bacteriology 48: 1413–1417.
https://doi.org/10.1099/00207713-48-4-1413 - Sakpuntoon V, Angchuan J, Boontham W, Khunnamwong P, Boonmak C, Srisuk N (2020) Grease waste as a reservoir of lipase-producing yeast and description of Limtongella siamensis gen. nov., sp. nov. Microorganisms 8: 27.
https://doi.org/10.3390/microorganisms8010027 - Sampaio JS (2011) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Cyrenella Gochenaur (1981), 5th edn. Elsevier, The Netherlands, pp 1743–1745.
https://doi.org/10.1016/B978-0-444-52149-1.00140-3 - Sampaio JS, Oberwinkler F (2011) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Kriegeria Bresadola (1891), 5th edn. Elsevier, The Netherlands, pp 1477–1479.
https://doi.org/10.1016/B978-0-444-52149-1.00118-X - Santos ARO, Leon MP, Barros KO, Freitas LFD, Hughes AFS, Morais PB, Lachance MA, Rosa CA (2018) Starmerella camargoi f.a., sp. nov., Starmerella ilheusensis f.a., sp. nov., Starmerella litoralis f.a., sp. nov., Starmerella opuntiae f.a., sp. nov., Starmerella roubikii f.a., sp. nov. and Starmerella vitae f.a., sp. nov., isolated from flowers and bees, and transfer of related Candida species to the genus Starmerella as new combinations. International Journal of Systematic and Evolutionary Microbiology 68: 1333–1343.
https://doi.org/10.1099/ijsem.0.002675 - Schmalreck AF, Lackner M, Becker K, Fegeler W, Czaika V, Ulmer H, Lass-Flörl C (2014) Phylogenetic relationships matter: antifungal susceptibility among clinically relevant yeasts. Antimicrobial Agents and Chemotherapy 58: 1575–1585.
https://doi.org/10.1128/AAC.01799-13 - Shen W, Sipos B, Zhao L (2024) SeqKit2: A Swiss army knife for sequence and alignment processing. Imeta 3: e191.
https://doi.org/10.1002/imt2.191 - Shen XX, Opulente DA, Kominek J, Zhou X, Steenwyk JL, Buh KV, Haase MAB, Wisecaver JH, Wang M, Doering DT, Boudouris JT, Schneider RM, Langdon QK, Ohkuma M et al. (2018) Tempo and mode of genome evolution in the budding yeast subphylum. Cell 175: 1533–1545.
https://doi.org/10.1016/j.cell.2018.10.023 - Sipiczki M (2013) Detection of yeast species also occurring in substrates associated with animals and identification of a novel dimorphic species in Verbascum flowers from Georgia. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 103: 567–576.
https://doi.org/10.1007/s10482-012-9841-9 - Sipiczki M, Baghela A (2025) Identification of Starmerella aleppica f.a., sp. nov. and large indels in the rRNA cistron that split the Starmerella genus. International Journal of Systematic and Evolutionary Microbiology 75.
https://doi.org/10.1099/ijsem.0.006629 - Smith MT (2011) The yeasts, a taxonomic study. In: Kurtzman CP, Fell JW, Boekhout T (ed) Aciculoconidium King DS & Jong SC (1976), 5th edn. Elsevier, The Netherlands, pp 955–957.
https://doi.org/10.1016/B978-0-444-52149-1.00086-0 - Smith MT (1986) Zygoascus hellenicus gen. nov., sp. nov., the teleomorph of Candida hellenica (= C. inositophila = C. steatolytica). Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 52: 25–37.
https://doi.org/10.1007/BF00402684 - Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30: 1312–1313.
https://doi.org/10.1093/bioinformatics/btu033 - Stavrou AA, Lackner M, Lass-Flörl C, Boekhout T (2019) The changing spectrum of Saccharomycotina yeasts causing candidemia: phylogeny mirrors antifungal susceptibility patterns for azole drugs and amphothericin B. FEMS Yeast Research 19: foz037.
https://doi.org/10.1093/femsyr/foz037 - Stelling-Dekker NM (1931) Die sporogenen Hefen. Verhandelingen der Koninklijke Akademie van Wetenschappen. Afdeeling Natuurkunde, Tweede Sectie 28: 1–547.
- Sugita T, Nakase T (1999) Non-universal usage of the leucine CUG codon and the molecular phylogeny of the genus Candida. Systematic and Applied Microbiology 22: 79–86.
https://doi.org/10.1016/S0723-2020(99)80030-7 - Suh SO, McHugh JV, Pollock DD, Blackwell M (2005) The beetle gut: a hyperdiverse source of novel yeasts. Mycological Research 109: 261–265.
https://doi.org/10.1017/s0953756205002388 - Suzuki M, Nakase T (2002) A phylogenetic study of ubiquinone-7 species of the genus Candida based on 18S ribosomal DNA sequence divergence. International Journal of Systematic and Evolutionary Microbiology 48: 55–65.
https://doi.org/10.2323/jgam.48.55 - Sylvester K, Wang QM, James B, Mendez R, Hulfachor AB, Hittinger CT (2015) Temperature and host preferences drive the diversification of Saccharomyces and other yeasts: a survey and the discovery of eight new yeast species. FEMS Yeast Research 15: fov002.
https://doi.org/10.1093/femsyr/fov002 - Takashima M, Manabe RI, Nishimura Y, Endoh R, Ohkuma M, Sriswasdi S, Sugita T, Iwasaki W (2019) Recognition and delineation of yeast genera based on genomic data: lessons from Trichosporonales. Fungal Genetics and Biology 130: 31–42.
https://doi.org/10.1016/j.fgb.2019.04.013 - Takashima M, Sugita T (2022) Taxonomy of pathogenic yeasts Candida, Cryptococcus, Malassezia, and Trichosporon. Medical Mycology Journal 63: 119–132.
https://doi.org/10.3314/mmj.22.004 - Tatusova TA, Madden TL (1999) BLAST 2 Sequences, a new tool for comparing protein and nucleotide sequences. FEMS Microbiology Letter 174: 247–250.
https://doi.org/10.1111/j.1574-6968.1999.tb13575.x - Teixeira ACP, Marini MM, Nicoli JR, Antonini Y, Martins RP, Lachance MA, Rosa CA (2003) Starmerella meliponinorum, a novel ascomycetous yeast species associated with stingless bees. International Journal of Systematic and Evolutionary Microbiology 53: 339–343.
https://doi.org/10.1099/ijs.0.02262-0 - Ter-Hovhannisyan V, Lomsadze A, Chernoff YO, Borodovsky M (2008) Gene prediction in novel fungal genomes using an ab initio algorithm with unsupervised training. Genome Research 18: 1979–1990.
https://doi.org/10.1101/gr.081612.108 - van der Walt JP (1966) Lodderomyces, a new genus of the Saccharomycetaceae. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 32: 1–5.
https://doi.org/10.1007/BF02097439 - van der Walt JP, Johannsen E (1973) The perfect state of Torulopsis magnoliae. Antonie Van Leeuwenhoek International Journal of General and Molecular Microbiology 39: 635–647.
https://doi.org/10.1007/BF02578907 - Villas-Bôas SG, Esposoito E, de Mendonça MM (2002) Novel lignocellulolytic ability of Candida utilis during solid-substrate cultivation on apple pomace. World Journal of Microbiology Biotechnology 18: 541–545.
https://doi.org/10.1023/A:1016350612380 - Wang QM, Bai FY (2008) Molecular phylogeny of basidiomycetous yeasts in the Cryptococcus luteolus lineage (Tremellales) based on nuclear rDNA and mitochondrial cytochrome b gene sequence analyses: proposal of Derxomyces gen. nov. and Hannaella gen. nov., and eight novel Derxomyces species. FEMS Yeast Research 8: 799–814.
https://doi.org/10.1111/j.1567-1364.2008.00403.x - Wickerham LJ, Kurtzman CP (1971) Two new Saturn-spored species of Pichia. Mycologia 63: 1013–1018.
https://doi.org/10.1080/00275514.1971.12019197 - Wickham H (2016) ggplot2: elegant graphics for data analysis. Springer, New York.
- Wu YQ, Ying CM (2016) Drug resistance genes of Candida albicans to azoles and their regulation mechanisms. Laboratory Medicine 14: 1848–1853.
https://doi.org/10.3969/j.issn.1673-8640.2016.09.002 - Yamada Y, Higashi T, Ando S, Mikata K (1998) The phylogeny of strains of species of the genus Pichia Hansen (Saccharomycetaceae) based on the partial sequences of 18S ribosomal RNA: the proposals of Phaffomyces and Starmera, the new genera. Bulletin of Faculty of Agriculture 47: 23–35.
- Yamada Y, Kawasaki H, Nagatsuka Y, Mikata K, Seki T (1999) The phylogeny of the cactophilic yeasts based on the 18S ribosomal RNA gene sequences: the proposals of Phaffomyces antillensis and Starmera caribaea, new combinations. Bioscience, Biotechnology, and Biochemistry 63: 827–832.
https://doi.org/10.1271/bbb.63.827 - Yamada Y, Maeda K, Mikata K (1994a) The phylogenetic relationships of the hat-shaped ascospore-forming, nitrate-assimilating Pichia species, formerly classified in the genus Hansenula Sydow et Sydow, based on the partial sequences of 18S and 26S ribosomal RNAs (Saccharomycetaceae): the proposals of three new genera, Ogataea, Kuraishia, and Nakazawaea. Bioscience, Biotechnology, and Biochemistry 58: 1245–1257.
https://doi.org/10.1271/bbb.58.1245 - Yamada Y, Matsuda M, Maeda K, Mikata K (1995a) The phylogenetic relationships of methanol-assimilating yeasts based on the partial sequences of 18S and 26S ribosomal RNAs: the proposal of Komagataella gen. nov. (Saccharomycetaceae). Bioscience, Biotechnology, and Biochemistry 59: 439–444.
https://doi.org/10.1271/bbb.58.1245 - Yamada Y, Matsuda M, Maeda K, Sakakibara C, Mikata K (1994b) The phylogenetic relationships of the saturn-shaped ascospore-forming species of the genus Williopsis Zender and related genera based on the partial sequences of 18S and 26S ribosomal RNAs (Saccharomycetaceae): the proposal of Komagataea gen. nov. Bioscience, Biotechnology, and Biochemistry 58: 1236–1244.
https://doi.org/10.1271/bbb.58.1236 - Yamada Y, Suzuki T, Matsuda M, Mikata K (1995b) The phylogeny of Yamadazyma ohmeri (Etchells et Bell) Billon Grand based on partial sequences of 18S and 26S ribosomal RNAs: the proposal of Kodamaea gen. nov. (Saccharomycetaceae). Bioscience, Biotechnology, and Biochemistry 59: 1172–1174.
https://doi.org/10.1271/bbb.59.117 - Yamazaki A, Lorliam W, Kawasaki H, Uchino M, Suzuki KI (2020) Fourteen novel lipomycetaceous yeast species isolated from soil in Japan and transfer of Dipodascopsis anomala to the genus Babjevia based on ascospore production phenotype. International Journal of Systematic and Evolutionary Microbiology 70: 1372–1397.
https://doi.org/10.1099/ijsem.0.003924 - Yarrow D, Meyer SA (1978) Proposal for amendment of the diagnosis of the genus Candida Berkhout nom. cons. International Journal of Systematic Bacteriology 28: 611–615.
https://doi.org/10.1099/00207713-28-4-611 - Yu HT, Shang YJ, Zhu HY, Han PJ, Wang QM, Santos ARO, Barros KO, Souza GFL, Alvarenga FBM, Abegg MA, Rosa CA, Bai FY (2023) Yueomyces silvicola sp. nov., a novel ascomycetous yeast species unable to utilize ammonium, glutamate, and glutamine as sole nitrogen sources. Yeast 40: 540–549.
https://doi.org/10.1002/yea.3901 - Yurkov A, Wehde T, Kahl T, Begerow D (2012) Aboveground deadwood deposition supports development of soil yeasts. Diversity 4: 453–474.
https://doi.org/10.3390/d4040453 - Zheng J, Liu KF, Liu XJ, Zhang L, Hui FL (2017) Deakozyma yunnanensis sp. nov., a novel yeast species isolated from rotten wood. International Journal of Systematic and Evolutionary Microbiology 67: 2436–2439.
https://doi.org/10.1099/ijsem.0.001978 - Copyright: The Author(s) 2026. Published by BioAcademic Press on behalf of Kunming Institute of Botany, Chinese Academy of Sciences (CAS) and Mushroom Research Foundation. This is an open access article under the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0), which permits use, distribution and reproduction in any medium, provided the original work is properly cited.