Abstract
Introduction
Pestalotiopsis-like taxa represent a ubiquitous group of fungi that form significant associations with various plants. They are functioning as pathogens, endophytes, or saprobes, and widely distributed in tropical and temperate regions (Maharachchikumbura et al. 2014a; 2014b; Jayawardena et al. 2019; Hyde et al. 2020c; Dong et al. 2023; Sun et al. 2023; Razaghi et al. 2024). Pestalotiopsis-like species have gained significant attention due to their capacity to produce novel chemical metabolites (Wu et al. 2022), their high species diversity and their role as phytopathogens (Darapanit et al. 2021). As phytopathogens, pestalotiopsis-like fungi are responsible for a variety of economically important plant diseases, including flower blight (Akinsanmi et al. 2017; Daengsuwan et al. 2021), leaf blight (Hsu et al. 2022; Rajashekara et al. 2023), twig blight (Qi et al. 2021), leaf spot (Tsai et al. 2021; Nozawa et al. 2022; Xia et al. 2022), root rot (Sun et al. 2021), fruit rot (Nozawa et al. 2020; Qin et al. 2023), and postharvest diseases (Abbas et al. 2022; Li et al. 2023b).
Pestalotiopsis-like taxa comprise three closely related genera—Pestalotiopsis, Neopestalotiopsis, and Pseudopestalotiopsis—belonging to Sporocadaceae, Amphisphaeriales, Xylariomycetidae, Sordariomycetes (Wijayawardene et al. 2022). Over the past decade, the number of species in this group has grown rapidly (Maharachchikumbura et al. 2014b; Razaghi et al. 2024; Wang et al. 2025). As of October 2025, Index Fungorum (www.indexfungorum.org) and MycoBank (www.mycobank.org) list more than 450 taxa under Pestalotiopsis, nearly 140 species in Neopestalotiopsis, and about 35 species in Pseudopestalotiopsis. However, the expanding spectrum of species highlights ongoing challenges in understanding their taxonomy. For intergeneric, observing that the three median cells of conidia are concolourous in Neopestalotiopsis ageratinae, N. amomi, N. hyperici and N. olivaceous (Liu et al. 2017; Sun et al. 2023; Cui et al. 2024; Razaghi et al. 2024), versicoloured within Pestalotiopsis biappendiculata, Pe. multicolor and Pe. taxicola (Razaghi et al. 2024; Wang et al. 2024), and Pseudopestalotiopsis camelliae-sinensis produces distinct conidiophores (Liu et al. 2017), which do not conform to the morphological characteristics at the genus level. For interspecific comparisons, there are considerable overlapping phenotypic traits that complicate the segregation of morphologically ambiguous taxa (Maharachchikumbura et al. 2014b; Li et al. 2021b; Cui et al. 2024).
Although phylogenetic analyses based on the combined ITS, tef1, and tub2 loci have greatly improved the delineation of major clades both among and within genera (Maharachchikumbura et al. 2014b), several challenges remain unresolved. In particular, several studies observed that certain branches in the multi-locus phylogenetic trees of Neopestalotiopsis and Pestalotiopsis exhibited notably short branch lengths and relatively low support values (Liu et al. 2017; Tsai et al. 2021; Hsu et al. 2024). Moreover, phylogenetic reconstructions based on ITS-tef1-tub2 sequence data for Neopestalotiopsis by Sun et al. (2023) and Razaghi et al. (2024) yielded discordant topologies. This incongruence suggests that the topology of the three-gene is unstable in Neopestalotiopsis, raising concerns about whether previously established backbone trees accurately represent the evolutionary relationships within this genus. Such instability, coupled with the short internal branches and low statistical support, makes it difficult to clearly distinguish between species and infraspecific lineages. One plausible explanation is taxonomic over-splitting, wherein minimal morphological or molecular differences are overinterpreted, leading to the recognition of multiple species that may, in fact, represent a single species (Dissanayake et al. 2024). Over-splitting not only contributes to systematic confusion but also has broader implications (Stengel et al. 2022). In plant pathology, for instance, it may result in the overestimation of pathogen diversity, misinterpretation of disease emergence, and inaccurate inference of pathogen evolution and transmission, ultimately complicating disease management and quarantine strategies.
The advent of integrative taxonomy has provided a new robust framework for resolving the aforementioned taxonomic inconsistencies (Maharachchikumbura et al. 2021; Stengel et al. 2022). By combining morphological, molecular, and ecological evidence, this approach not only enhances the accuracy of species delimitation but also provides a solid foundation for evolutionary reconstruction. Recent studies have demonstrated the effectiveness of integrative taxonomy across various fungal genera. For example, Sklenář et al. (2022) employed combined multi-gene phylogenetic analyses, species delimitation methods (ABGD, bPTP, PTP, bGMYC, GMYC, and STACEY), and morphological as well as physiological traits to reduce 17 species in the Aspergillus versicolores series to four. Similarly, Dissanayake et al. (2024) integrated single- and multi-gene phylogenies (ITS, tef, tub, cal, and his) with GCPSR, PTP, and mPTP analyses to reconstruct the genus Diaporthe, resulting in its subdivision into seven sections and the clarification of 13 species and 15 species complexes, along with 31 synonymies. Moreover, the rapid advancement of high-throughput sequencing technologies has further expanded the application of integrative taxonomy to phenotypically conserved and non-model organisms. In recent years, genome-scale phylogenetic analyses have significantly improved taxonomic resolution across fungi, animals, and plants. For instance, Steenwyk et al. (2024) performed whole-genome sequencing and phylogenomic analyses of multiple Aspergillus strains, including type species, revealing numerous misidentifications in morphology-based taxonomy and highlighting the superior precision of genome-scale data. Likewise, Liu et al. (2024) redefined generic boundaries within the Saccharomycetaceae using a genome-based classification framework.
Building upon this background, the current study aims to 1) resolve the species boundaries of pestalotiopsis-like taxa by employing the three loci genes (ITS, tef1, and tub2) in conjunction with whole-genome data, and 2) construct a robust and reliable backbone phylogenomic tree for pestalotiopsis-like fungi to assist in resolving species boundaries using ITS, tef1, and tub2 sequences, 19 publicly available genomes obtained from the NCBI Genome database, and 70 genome sequences generated in this research.
Materials and methods
Isolates
During field trips conducted from 2021 to 2023, specimens were collected from various host plants in China and Thailand to identify pestalotiopsis-like fungi. Sampling included diseased tissues showing leaf spots or other symptoms, healthy plant tissues, and dead twigs (Table S1). Relevant data, including location and date, were documented. Samples were transported to the laboratory in envelopes or ziplock bags under sterile conditions and stored in a refrigerator at 4°C until fungal isolation and examination were conducted. Some type strains were provided by the Centre of Excellence in Fungal Research at Mae Fah Luang University, Thailand (Table S2). The names of the new taxa were registered in Index Fungorum (2025).
Morphological observation and characterisation
VHX-7000 (Keyence, Osaka, Japan), Fully-Integrated Head VHX-7100 (Keyence, Osaka, Japan) and High-Performance Camera VHX-7020 (Keyence, Osaka, Japan) dissecting microscopes were used as vehicles for observing the conidiomata. Morphological characters were examined and photographed by a ZEISS Axioscope 5 Camera (Zeiss, Oberkochen, Germany) compound microscope fitted with a ZEISS Axiocam 208 Color Microscope Camera (Zeiss, Oberkochen, Germany). Tarosoft Image Frame Work software was used for measurement. Photoplates were prepared with Adobe Photoshop CS6 Extended (Adobe, USA).
Morphological Comparison of Existing Species
To evaluate the correlation between phylogenetic relationships and morphological characteristics, 15 micromorphological traits were collected from published taxonomic descriptions of all pestalotiopsis-like species. The collected morphological data are provided in Tables S4, S12, and S14. These traits included conidial length and width, basal cell length, the length and coloration of the three median cells, second, third, and fourth cell lengths, apical cell length, apical appendage length, and basal appendage length, following the morphological classification criteria outlined by Maharachchikumbura et al. (2014b). The compiled data were visualized as bar charts using R v. 4.3.3 (R core team 2024). To further assess morphological variation among species, a distance matrix was constructed based on the collected micromorphological measurements. Multidimensional Scaling (MDS) was then applied to reduce the dimensionality of the dataset while preserving the relative morphological relationships among species. The analysis was conducted using the Euclidean distance metric, and the results were visualized as scatter plots in R v. 4.3.3 (R core team 2024).
PCR and Sequence Assembly
The genomic DNA was extracted from mycelium grown for seven days on PDA using the BIOMIGA Fungus Genomic DNA Extraction Kit (Biomiga GD2416, USA). This DNA served as the template for Polymerase Chain Reaction (PCR). The PCR reaction mixture included 1 μL of DNA template, 1 μL each of forward and reverse primers, 12.5 μL of Taq PCR Master Mix (2X, with Blue Dye; Sangon Biotech, Shanghai, P.R. China), and 9.5 μL of ddH2O. DNA amplification was performed for three loci, including the 5.8S nuclear ribosomal DNA gene with the two flanking internally transcribed spacer regions (ITS rDNA), as well as partial sequences of the translation elongation factor 1-alpha (tef1) and β-tubulin (tub2) genes. The primer pairs used were ITS4/ITS5 (White et al. 1990) for ITS, EF1-526F/EF1-1567R (Maharachchikumbura et al. 2013a) for tef1, and T1/Bt2b (Glass and Donaldson 1995; O'Donnell and Cigelnik 1997) for tub2. The amplification conditions for ITS, tef1, and tub2 included an initial denaturation at 94°C for 4 minutes, followed by 35 cycles of 94°C for 30 seconds, 55°C for 30 seconds, and 72°C for 60 seconds, with a final extension at 72°C for 10 minutes. The amplified PCR products were purified and sequenced at Sangon Biotech (Shanghai) Co., Ltd. Consensus sequences were obtained using Geneious Prime v. 2023.2.1 (https://www.geneious.com/) from sequences generated with forward and reverse primers. Newly generated sequences were deposited in GenBank (Table S1).
Phylogenetic analysis
All type isolates and other representative strains of pestalotiopsis-like fungi, published prior to December 2024 (Table S1), were retrieved from GenBank based on existing literature. The downloaded sequences were aligned with those obtained from this study using the MAFFT online service (Katoh et al. 2019) available at (https://mafft.cbrc.jp/alignment/server/), and manually refined in AliView v. 1.28 (Larsson 2014) for maximum alignment. Phylogenetic networks were constructed using the Log Det distances with the Neighbor Net algorithm in SplitsTree v. 4.19.2 (Huson and Bryant 2006). For each genus, a Maximum Likelihood (ML) tree of the three-gene sequence data was estimated using RAxML-NG v. 1.2.2 (Kozlov et al. 2019), with GTR model and rapid bootstrap analysis involving 1000 replicates. The Bayesian analyses were conducted using MrBayes (Huelsenbeck and Ronquist 2001) with six simultaneous Markov Chain Monte Carlo (MCMC) chains across two independent runs, totaling 50,000,000 generations. The sampling frequency was set to every 1,000 generations, and the run was automatically terminated when the standard deviation of split frequencies fell below 0.01. The first 25% of the trees were discarded as burn-in.
Selection of fungal strains for whole-genome sequencing
A total of 35 strains (including 15 type strains) of Neopestalotiopsis, 28 strains (including five type strains) of Pestalotiopsis, and seven strains (including one type strain) of Pseudopestalotiopsis were chosen from different clades of the three-gene phylogenetic tree (ITS, tef1, and tub2) for whole-genome sequencing. These strains were sourced from earlier sampling conducted by our team and Mae Fah Luang University. We initiated the process by searching the NCBI Genome Browser (https://www.ncbi.nlm.nih.gov/) on August 1, 2024, using the terms "Neopestalotiopsis", "Pestalotiopsis", and "Pseudopestalotiopsis" to identify publicly available genome data relevant to this study. Consequently, we obtained six Neopestalotiopsis genomes, 11 Pestalotiopsis genomes, and two Pseudopestalotiopsis genomes. Additionally, we extracted the ITS, tef1, and tub2 gene sequences from these 19 publicly available genomes for a three-gene phylogenetic analyses. Detailed taxonomic information and the sources of the 89 pestalotiopsis-like fungal genomes used in this study are provided in Table S2.
Genome sequencing and assembly
In this study, 70 new genomes were sequenced. Mycelia from 3-day-old colonies growing on PDA were transferred to a 500 mL conical flask containing 300 mL of potato dextrose broth (PDB) culture medium and incubated at 25°C for 3 to 5 days in a Laboratory Shaking Incubator (ZHICHENG ZWYR-D2401, China) set to 120 rpm. After incubation, fresh mycelia were harvested by centrifugation at 4000 g for 10 minutes (at 4°C), the supernatant was discarded, and the fresh mycelia were stored at −80 °C for DNA extraction. The genomic DNA was extracted from mycelium using the BIOMIGA Fungus Genomic DNA Extraction Kit (Biomiga GD2416, USA). DNA concentration and purity were assessed using a Qubit 3.0 Fluorometer (Invitrogen, USA), while DNA integrity was confirmed on a 1% agarose gel. For samples that met the established criteria, a 150 bp paired-end reads sequencing library was constructed utilizing the VAHTS® Universal Plus DNA Library Prep Kit for Illumina (ND617-02, Vazyme Biotech Co., Ltd, Nanjing, China). The prepared libraries were subsequently sequenced on an Illumina NovaSeq X Plus (Illumina Inc., San Diego, CA USA).
Fastp v. 0.22.0 was used to remove adapter sequences and trim low-quality reads (Chen et al. 2018a). The quality-filtered sequences were assembled using the SPAdes assembler v. 3.6.2 (Bankevich et al. 2012). The genome assemblies generated in this study have been deposited in the National Microbiology Data Centre (NMDC) under BioProject NMDC10019240 (https://nmdc.cn/; Table S2).
Genome assessment
Understanding the quality of genomic resources is essential before undertaking downstream analyses to guarantee an impartial interpretation of results (Manni et al. 2021b). Benchmarking Universal Single-Copy Orthologs (BUSCO) provides a metric for the quantitative assessment of genome assembly and annotation completeness, based on evolutionarily informed expectations of gene content (Simão et al. 2015). Here, we used BUSCO v. 5.7.1 (Manni et al. 2021a) with the sordariomycetes_odb10 database to assess the completeness of all genome assemblies.
Phylogenomic data matrix construction
Whole-genome protein-coding genes of pestalotiopsis-like fungi (Table S2) were clustered into orthogroups with Orthofinder v. 2.5.5 (Emms & Kelly 2015). Single-copy ortholog sequences were aligned individually using MAFFT v. 7.520 (Katoh & Standley 2013) with default parameters, then trimmed using trimAL v. 1.4.rev22 (Capella-Gutiérrez et al. 2009) with options “-automated1” flag. All single-locus alignments were concatenated into a supermatrix using a Python script. The resulting supermatrix and partition file were feed into IQ-TREE v. 2.13 (Minh et al. 2020) with parameters “-m TEST” to construct individual gene trees and to select the best-fit evolutionary model for each gene based on the Bayesian Information Criterion (BIC) using ModelFinder (Kalyaanamoorthy et al. 2017). The concatenated supermatrix and corresponding evolutionary models were employed to reconstruct the species tree using a concatenation-based method.
Species delimitation
To define species boundaries, we employed a comprehensive framework that integrates various species delimitation methods to analyze concatenated ITS, tef1, and tub2 gene datasets. This approach included: (1) genealogical concordance phylogenetic species recognition (GCPSR) (Taylor et al. 2000); (2) two heuristic methods, i.e., Poisson Tree Processes (PTP) (Zhang et al. 2013a) and multi-rate PTP (mPTP) (Kapli et al. 2017); and (3) two genetic distance-based approaches, i.e., Automatic Barcode Gap Discovery (ABGD) (Puillandre et al. 2012) and Assemble Species by Automatic Partitioning (ASAP) (Puillandre et al. 2021). Furthermore, to further evaluate species boundaries, we conducted an average nucleotide identity (ANI) analysis and calculated the core genome percentages. The taxa delimited by the DNA-based species delimitation analyses were considered as molecular operational taxonomic units (MOTU).
The GCPSR principle involves evaluating individual gene trees to compare highly supported evolutionary branches, thereby detecting phylogenetic conflicts between these branches. Based on these assessments, the GCPSR principle is applied to determine species boundaries. Subclades are regarded as distinct independent evolutionary lineages (IEL) if they meet the following criteria: (a) they are clearly separated from other lineages, forming bifurcating branches with associated relative lengths; (b) they receive strong support in single-gene trees (ML ≥ 70% and PP ≥ 0.90), and no conflicting branches with comparable or higher support levels appear in multiple other single-gene trees. For these evaluations, ML and BI analyses were conducted on single-gene sequence alignments as described above. Species lacking specific gene data were excluded from the analysis for that gene region. Strains with only a single gene were similarly excluded from the analysis. IELs were ultimately confirmed if they demonstrated strong support (ML ≥ 70% and PP ≥ 0.90) in phylogenetic analyses of the majority of concatenated datasets.
The PTP and mPTP analyses are statistical methods employed for species delimitation within molecular phylogenetic trees (Zhang et al. 2013a; Kapli et al. 2017). In this study, we employed ML trees generated in RAxML-NG as input for both PTP and mPTP analyses. The PTP analysis was conducted using the implementation integrated within iTaxoTools v. 0.1.1 alpha (Vences et al. 2021), with 1,000,000 Markov Chain Monte Carlo (MCMC) generations, a thinning interval of 100, and a burn-in fraction of 10%. The convergence of MCMC iterations was evaluated by examining log-likelihood trace plots. The mPTP analysis was conducted on a local Linux server with 5,000,000 MCMC generations, employing two independent MCMC runs to ensure robustness.
The ABGD and ASAP analyses were carried out using the Kimura two-parameter (K2P) nucleotide substitution model to calculate pairwise nucleotide distance matrices, with all other parameters set to their defaults. The ABGD analysis was conducted on the specific web server (https://bioinfo. mnhn.fr/abi/public/abgd/abgdweb.html); to minimize subjectivity, the median number of partitions (i.e., the partition closest to P = 0.01) was chosen as the basis for hypothetical species delimitation (Puillandre et al. 2012; Pereira & Phillips 2024). ASAP analysis was carried out on its respective web server (https://bioinfo.mnhn.fr/abi/public/asap/), with species delimitation determined based on the partition with the best ASAP score.
The ANI analyses have been widely used for species boundary delineation and taxonomic identification in bacteriology (Varghese et al. 2015; Jain et al. 2018). In recent years, its application has been expanded to include the classification of eukaryotic microbes (Wibberg et al. 2021; Gotting et al. 2022). In this study, ANI values for all genomes were calculated using FastANI v. 1.33 (Jain et al. 2018) with default parameters. Additionally, pangenome and core genome analyses were conducted using Orthofinder v. 2.5.5 (Emms & Kelly 2015).
The combined species names adhere to the priority rules set forth by the International Code of Nomenclature for algae, fungi, and plants (Turland et al. 2018).
Repetitive annotation
We utilized four software tools — LTR_FINDER v. 1.06 (Ou and Jiang 2019), MITE-Hunter (Han & Wessler 2010), RepeatScout v. 1.0.6 (Price et al. 2005), and RepeatModeler v. 4.0.7 (Flynn et al. 2020) — to construct a repeat sequence database for the fungal genome based on structural prediction and de novo prediction principles. The database was classified using PASTEClassifier (Hoede et al. 2014) and then merged with the Repbase (20181026) (Bao et al. 2015) to form the final repeat sequence database. Subsequently, RepeatMasker v. 4.1.5 (http://www.repeatmasker.org/) was employed to predict the repeat sequences in the fungal genome based on the constructed repeat sequence database.
Gene prediction and protein annotation
Gene prediction was carried out using a combination of ab initio and homology-based methods. Ab initio prediction of coding regions was performed with Augustus v. 2.4 (Stanke et al. 2006), GeneID v. 1.4 (Blanco et al. 2007), GlimmerHMM v. 3.0.4 (Majoros et al. 2004) and SNAP v. 2006-07-28 (Korf 2004). For homology-based gene prediction, GeMoMa v. 1.3.1 (Keilwagen et al. 2016) was be use. Details of the reference genomes used are presented in Table S2. Gene models created from all the methods were integrated by EvidenceModeler (EVM) (Haas et al. 2008). Weights for each type of evidence were set as follows: GeMoMa > Augustus > GenelD = GlimmerHMM = SNAP.
The predicted protein-coding genes were functionally annotated by comparing their protein sequences with the KOG and the Kyoto Encyclopedia of Genes and Genomes (KEGG) (Kanehisa et al. 2017), TrEMBL/Swiss-Prot (Boeckmann et al. 2003), and NCBI non-redundant (nr) databases. Gene ontology (GO) terms were assigned to predicted genes using Blast2GO (Conesa et al. 2005). Pfam were assigned to predicted genes using hmmer (Finn et al. 2011).
The predictions of secreted proteins and effectors were based on the methods described by Ayukawa et al. (2021) and Amezrou et al. (2024). We analyzed all predicted gene protein sequences using SignalP 4.0 (Petersen et al. 2011) to detect signal peptides, and TMHMM (Krogh et al. 2001) to identify proteins with transmembrane helices. Proteins with signal peptides but without transmembrane helices were classified as secretory proteins. We further analyzed the secretory proteins using EffectorP (Sperschneider et al. 2016) to predict fungal effector proteins. To annotate the functions of carbohydrate enzyme genes, we utilized three tools within dbCAN3 (Zheng et al. 2023a): HMMER with an E-value threshold of less than 1e-15 and a coverage greater than 0.35, dbCAN_sub with an E-value threshold of less than 1e-102, and DIAMOND with an E-value of less than 1e-15 and coverage greater than 0.35. The results were determined by taking the intersection of the outputs from HMMER, dbCAN_sub, and DIAMOND.
Statistical analyses
All statistical analyses were performed in R v. 4.3.3 (R core team 2024). Figures were created using R and FigTree, with final adjustments made in Adobe Illustrator CC 2019 to enhance their quality visualization.
Results
To minimize analytical errors, we excluded the sequences of certain strains from the analyses in this study due to their significant genetic divergence from other strains within the same genus in the alignment results. Furthermore, these sequences showed low similarity to other species within the same genus in the NCBI database. The excluded sequences include tef1 (MH388389) of Neopestalotiopsis pandanicola KUMCC 17-0175, which, although identified as Neopestalotiopsis based on BLAST results, showed substantial divergence from other Neopestalotiopsis taxa in the alignment; tef1 (KU844185) of Pestalotiopsis lijiangensis CFCC 50738, which BLAST results indicated was more closely related to Neopestalotiopsis with a similarity exceeding 99% and tub2 (KU844184) of Pe. lijiangensis CFCC 50738, which was mostly similar to Lophodermium and Phlyctema, but the similarity was only around 80%.
Considering that certain species within pestalotiopsis-like taxa have shown unusually long-branch attraction in both earlier studies and our preliminary phylogenetic analyses, we re-sequenced the available type materials to mitigate possible sequencing errors. Furthermore, some previously published strains lacked complete sequencing data and required additional sequencing. Consequently, we rectified erroneous sequences for six markers across five species: tef1 (MH388404) and tub2 (MH412725) of N. chiangmaiensis MFLUCC 18-0113; tef1 (KX789689) of N. cocoes MFLUCC 15-0152; tef1 (MZ683389) of N. zingiberis HGUP 10001; tef1 (MZ868328) of Pe. ficicrescens GUCC 21556, and tub2 (JX399027) of Pe. linearis MFLUCC 12-0271_. We supplemented the following sequence for three markers across three species: ITS of _N. chiangmaiensis MFLUCC 18-0113, tub2 of N. cocoes MFLUCC 15-0152, and tub2 of Pe. endophytica MFLUCC 18-0932. All corrected and newly generated supplementary sequences have been deposited in GenBank (Table S1).
Neopestalotiopsis
Phylogenetic analyses and phylogenetic species recognition of Neopestalotiopsis
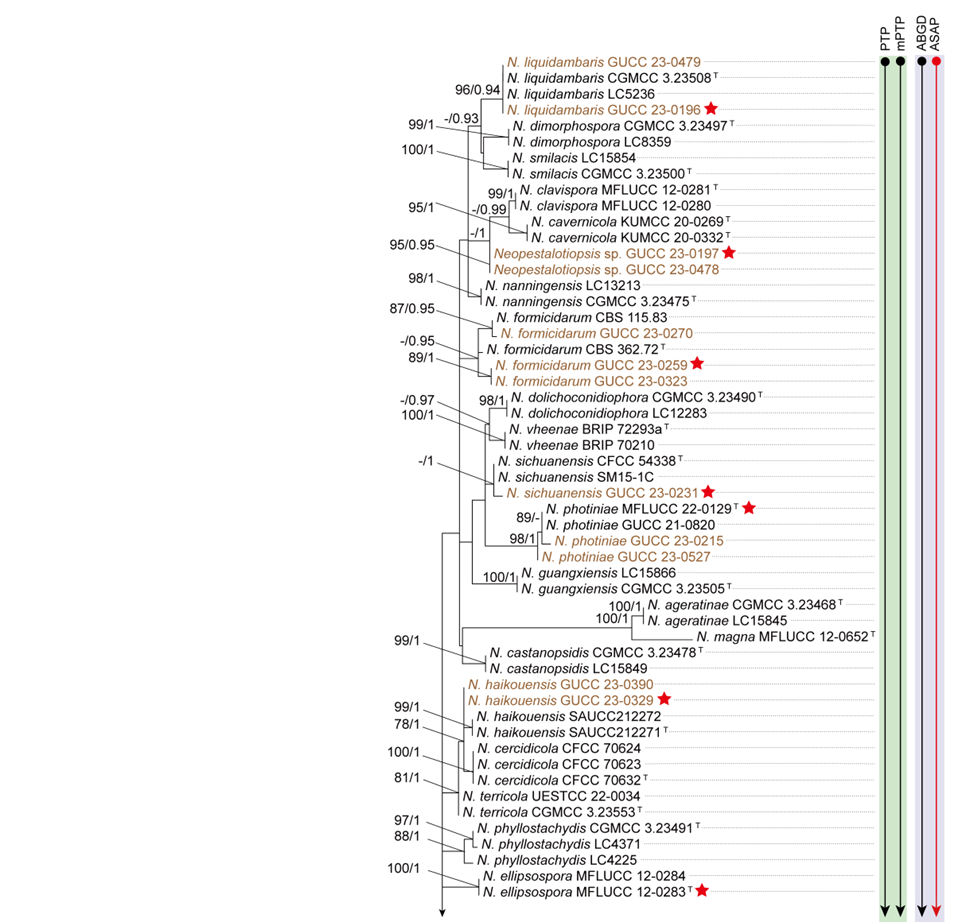
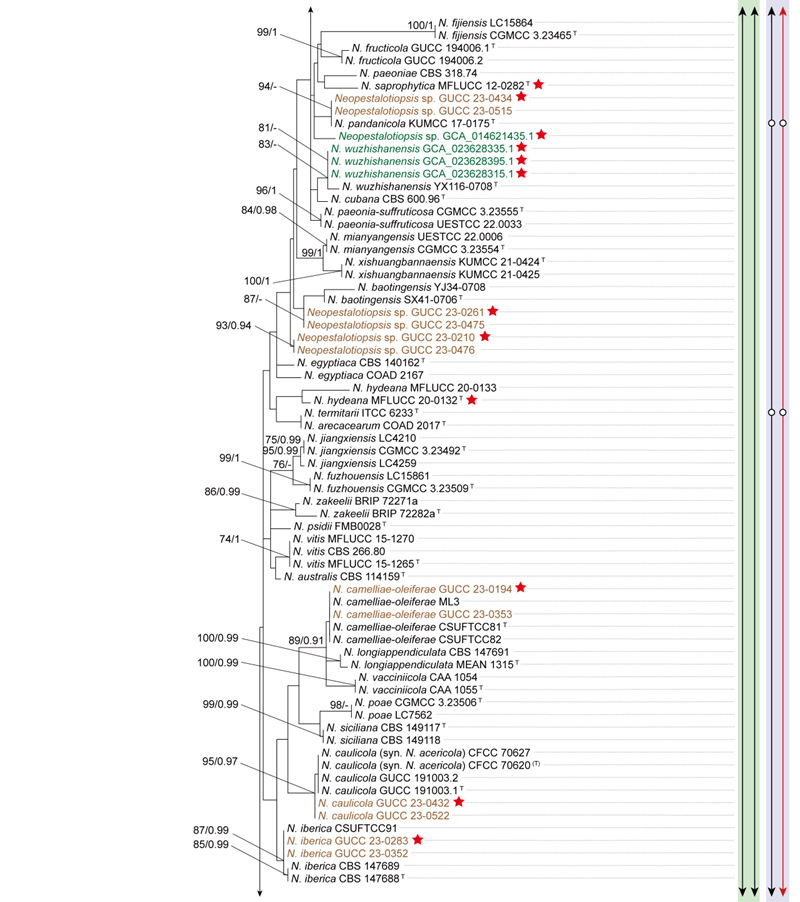
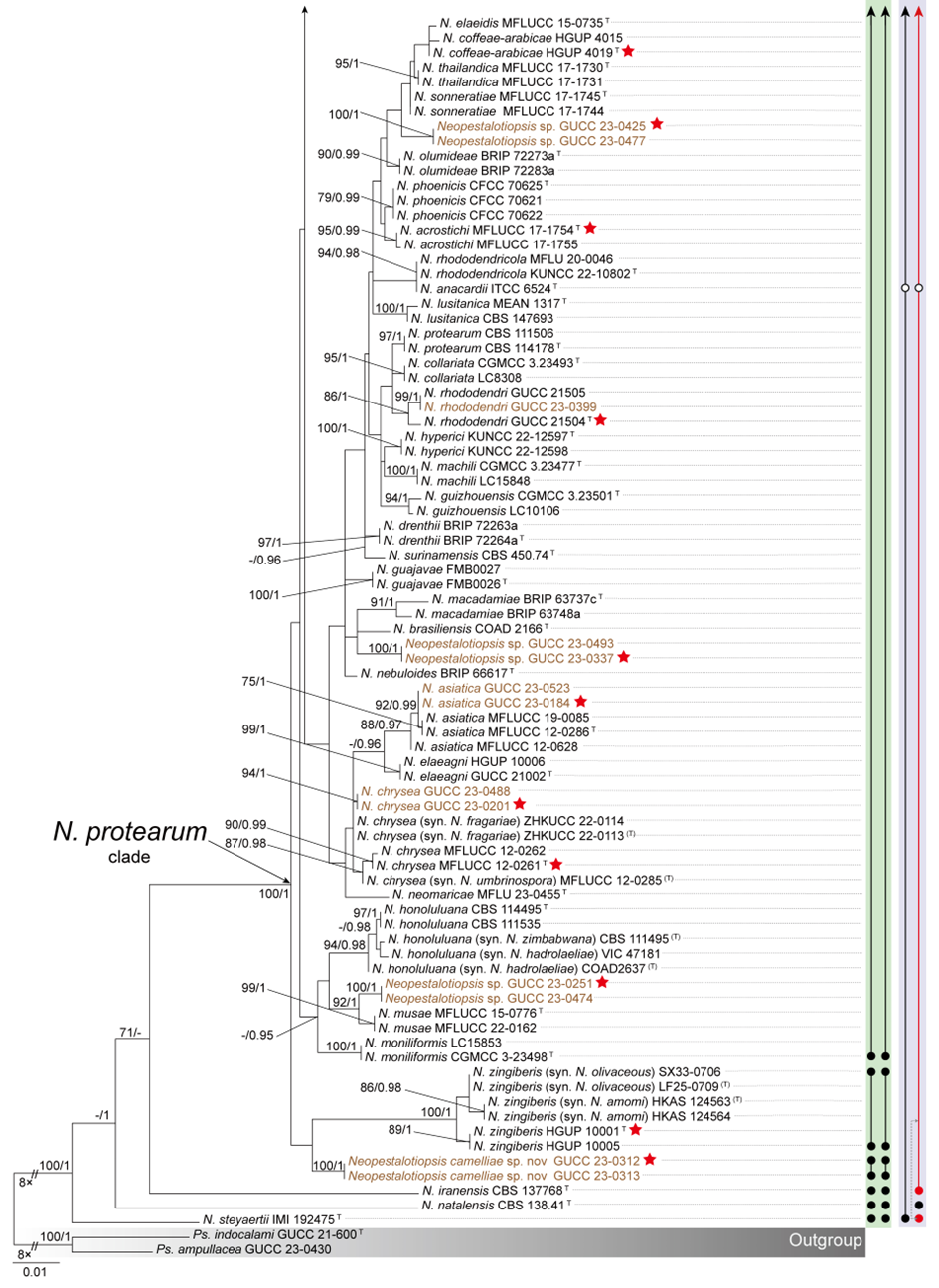
To evaluate the genus Neopestalotiopsis, phylogenetic trees were constructed based on individual loci (ITS, tef1, and tub2) as well as a concatenated three-locus dataset, incorporating all available type strains (Fig. S1 and Fig. 1). The combined phylogenetic tree (Fig. 1) was generated using alignments of the ITS (511 bp), tef1 (528 bp), and tub2 (741 bp) regions, including alignment gaps. This comprehensive analysis included 276 isolates, with Pseudopestalotiopsis indocalami GUCC 21600 and Ps. ampullacea GUCC 23-0430 designated as outgroup taxa. Among these, 45 isolates were newly generated in this study, and six strains were retrieved from the NCBI Genome database. The ML tree was inferred using RAxML-NG, resulting in a best-scoring tree with a final likelihood value of -10467.982263 (Fig. 1). The overall topologies of the ML and Bayesian trees were consistent for N. iranensis, N. natalensis, N. steyaertii, and the terminal nodes of the N. protearum clade. However, discrepancies were observed at the internal nodes of the N. protearum clade. In the ML tree, this clade was subdivided into several poorly supported sub-branches, whereas in the BI tree, the N. protearum clade appeared as a polytomous node. This is in line with the observation that the majority of the terminal nodes within the N. protearum clade show high support values, whereas most internal nodes lack support. For clarity and conciseness, only the ML tree is presented (Fig. 1). Bootstrap values and posterior probabilities are provided for clades that exhibit strong support.
Applying the GCPSR principle, N. iranensis, N. natalensis, and N. steyaertii each form distinct monophyletic lineages in both single-gene and concatenated three-gene phylogenies, supporting their recognition as three independent evolutionary lineages. In contrast, the remaining Neopestalotiopsis species, which cluster together with high support values (BS/PP = 100%/1), are collectively referred to here as the N. protearum clade. Applying the GCPSR principle to the N. protearum clade is challenging due to several factors. The ITS gene exhibits limited resolution in distinguishing taxa within this clade (Fig. S1a), while the tef1 and tub2 genes display substantial conflicts (Fig. S1b, c). For instance, taxa that form distinct monophyletic lineages in the tub2 phylogeny appear distantly related in the tef1 tree. Under the original classification framework of Neopestalotiopsis, most species cannot be clearly distinguished from other lineages based on tef1 or tub2 gene sequences, as they fail to form well-supported bifurcating branches of appropriate relative length. This limitation is particularly evident in species such as N. fructicola, N. jiangxiensis, and N. poae. Despite these challenges, there is inadequate evidence to support the hypothesis that all taxa within the N. protearum clade represent a single species. This is partly attributed to the presence of long branches in individual gene trees for certain taxa (Fig. S1), such as N. ageratinae, N. amomi, N. fijiensis, N. magna, and N. zingiberis. The inconsistencies in topological structures, low support values across the three-gene phylogenetic trees, and significant gene conflicts within the N. protearum clade (Figs. 1, S1) further underscore the unresolved species boundaries. These findings underscore the difficulties in confidently classifying certain isolates included in this study.
Both PTP and mPTP analyses yielded consistent results, delineating N. iranensis, N. natalensis, and N. steyaertii as three MOTUs, while subdividing the N. protearum clade into three MOTUs (Fig. 1). Specifically, isolates GUCC 23-0312 and GUCC 23-0313 were grouped into one MOTU; N. amomi, N. olivaceous, and N. zingiberis formed a second MOTU, and the remaining species of the N. protearum clade constituted a third MOTU. In contrast, the ABGD analysis classified all taxa within Neopestalotiopsis as a single MOTU, whereas ASAP identified two MOTUs: N. natalensis as one MOTU, with all remaining taxa grouped into a second (Fig. 1)
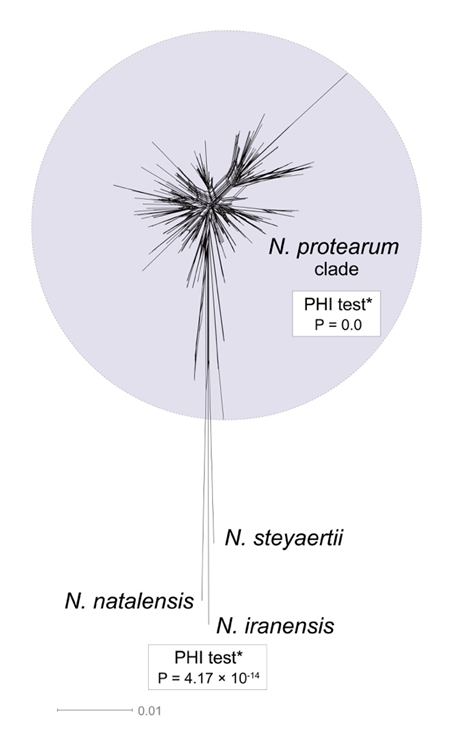


Further analysis using a phylogenetic network revealed that N. iranensis, N. natalensis, and N. steyaertii each formed distinct long branches. However, the network analysis of the N. protearum clade indicated multiple conflicting evolutionary signals, suggesting significant recombination events (P = 0). These conflicts were evident in the network’s structure, which exhibited boxlike polygons and irregular branch lengths. Such patterns obscure clear species boundaries and complicate interpretations of evolutionary relationships among the taxa (Fig. 2).
Whole-genome data and phylogenomic assessment of Neopestalotiopsis
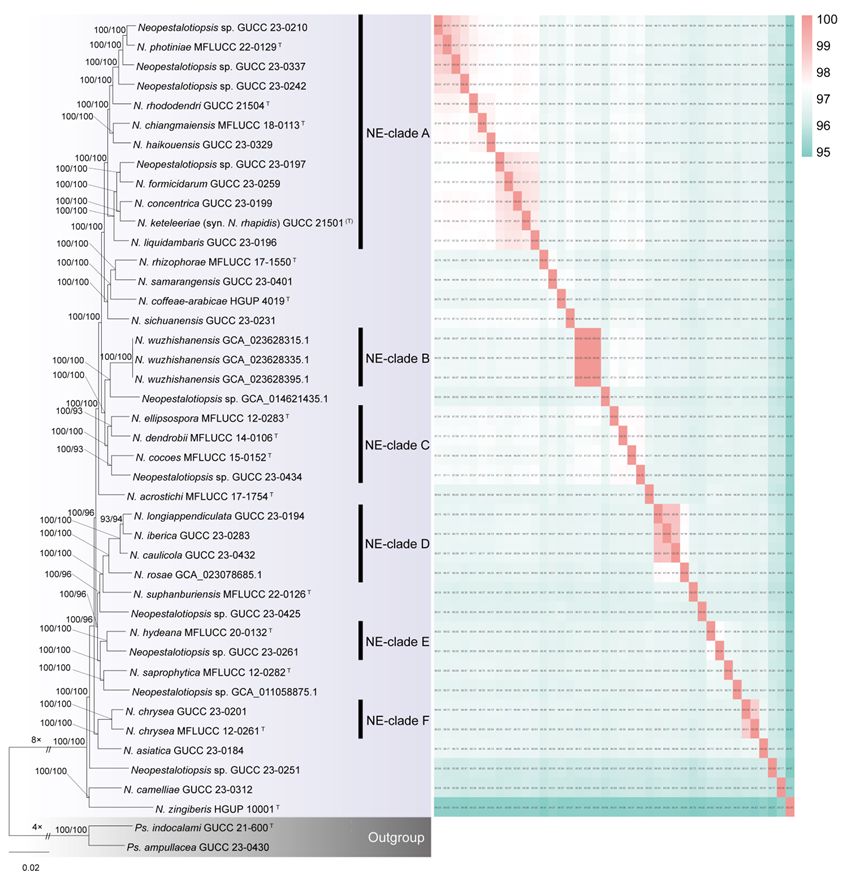
This study analyzed 41 Neopestalotiopsis genomes, including six publicly available genomes retrieved from the NCBI database and 35 newly sequenced genomes. These genomes belong to the N. protearum clade and represent 28 known species (including 15 type strains), one novel species, and ten unidentified taxa. Phylogenomic analysis based on 9,046 single-copy orthologous genes, incorporating 41 Neopestalotiopsis strains and two outgroups (Pseudopestalotiopsis indocalami GUCC 21-600 and Ps. ampullacea GUCC 23-0430), revealed well-resolved topologies (Fig. 3)
To assess nucleotide-level genomic similarity among different genomes, an average nucleotide identity (ANI) analysis was performed (Fig. 3; Table S5). The clustering pattern based on ANI was largely consistent with the whole-genome phylogenetic tree (Fig. 3). Based on a threshold of ANI > 97.30%, six clades (NE-clade A to NE-clade F) could be identified in the whole-genome phylogenetic tree. Within NE-clade A, ANI values ranged from 97.22% to 98.70% (mostly > 97.30%); NE-clade B (three genomes) ranged ANI values from 99.99–100%; NE-clade C (four genomes) from 97.34–97.46%; NE-clade D (four genomes) from 97.38–98.69%; NE-clade E (two genomes) was 97.37%; and NE-clade F (two genomes) was 98.41%.
However, the clustering pattern inferred from whole-genome data exhibited marked discrepancies compared to the concatenated three-gene phylogenetic tree (Fig. 1). For example, MFLUCC 22-0129, GUCC 23-0210, and GUCC 23-0337 clustered together in the whole-genome phylogenetic tree and exhibited ANI values exceeding 98.35%, despite being positioned in separate subclades in the three-gene phylogeny (Figs. 1, 3). Likewise, strains GUCC 23-0432, GUCC 23-0283, and GUCC 23-0194, forming NE-clade D in the genome-wide tree (ANI > 98.6%), and strains MFLUCC 12-0261 and GUCC 23-0201, representing NE-clade F (ANI > 98.4%), were placed in distinct subclades in the multi-locus phylogeny. These results highlight substantial incongruence between phylogenetic relationships inferred from multi-locus sequence data and those derived from whole-genome comparisons.
Gene family analyses of the six clades (NE-clade A–F) revealed that core gene clusters accounted for 88.35–99.42% of the pangenome, variable gene clusters for 0.27–9.37%, and unique gene clusters for 0.31–3.42% (Fig. 4). This indicates that members within each clade share the majority of their fundamental biological functions.
Integrating evidence from the whole-genome phylogeny, ANI, and pangenome composition, an ANI threshold of > 97.30% combined with a core-genome proportion exceeding 88% appears to be a reasonable boundary for species delimitation within Neopestalotiopsis.
Morphology of Neopestalotiopsis
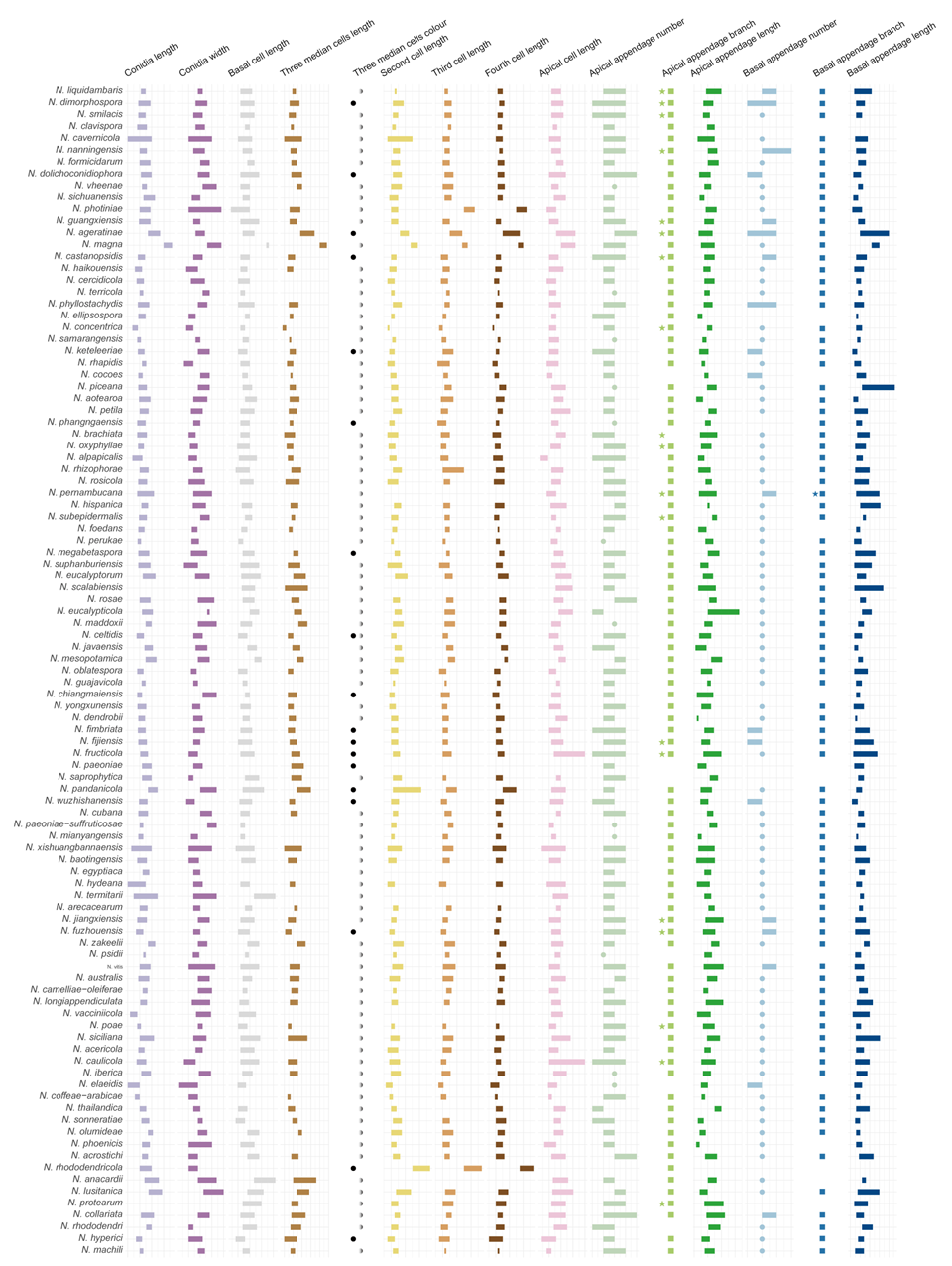
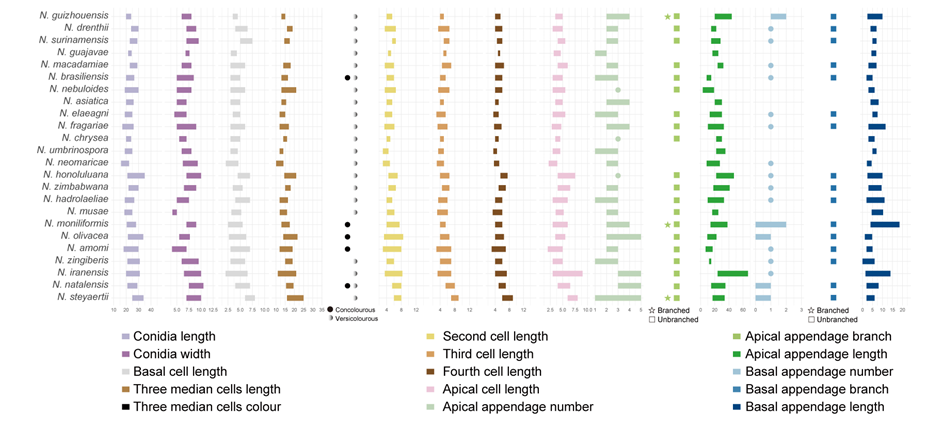
The morphological characteristics of conidia, as detailed in published descriptions of type strains of Neopestalotiopsis species, were graphically represented (Fig. 5). Key traits analyzed included conidial length, basal cell length, the length and coloration of the three median cells, the lengths of the second, third, and fourth cells, apical cell length, as well as the number, length, and branching patterns of both apical and basal appendages. A comprehensive summary of these morphological data is provided in Table S4.
Figure 5 provides a comprehensive overview of the micromorphological dimensions of each species, aligned with their positions in the three-gene concatenated phylogenetic tree. Notably, the analysis reveals significant discrepancies between molecular and morphological data. For instance, N. keteleeriae and N. rhapidis are closely related phylogenetically, they exhibit distinct morphological differences. Specifically, N. keteleeriae is characterized by significantly wider conidia and the presence of both concolourous and versicolourous three median cells, whereas N. rhapidis displays narrower conidia with exclusively versicolourous three median cells. Conversely, some species with highly similar morphological traits, such as N. ellipsospora and N. mianyangensis, as well as N. cercidicola and N. machili, are phylogenetically distant. Across the genus Neopestalotiopsis, while most species feature versicolourous three median cells, 23 species, including N. ageratinae, N. amomi, N. brasiliensis, N. castanopsidis, N. celtidis, N. chiangmaiensis, N. dimorphospora, N. dolichoconidiophora, N. fijiensis, N. fimbriata, N. fructicola, N. fuzhouensis, N. hyperici, N. keteleeriae, N. megabetaspora, N. moniliformis, N. natalensis, N. olivaceous, N. paeoniae, N. pandanicola, N. phangngaensis, N. rhododendricola, and N. wuzhishanensis, exhibit concolourous three median cells. Conidial dimensions vary considerably, with lengths ranging from 10 to 47 μm and widths from 3 to 12.5 μm. Furthermore, the morphology of apical appendages exhibits significant variation, with lengths varying from 2 to 67 μm. Despite these observed morphological variations, no clear correlation is evident between morphological traits and phylogenetic relationships. Furthermore, the morphological spectra of most taxa exhibit significant overlap, further complicating species delimitation based solely on conidial morphology of the genus.
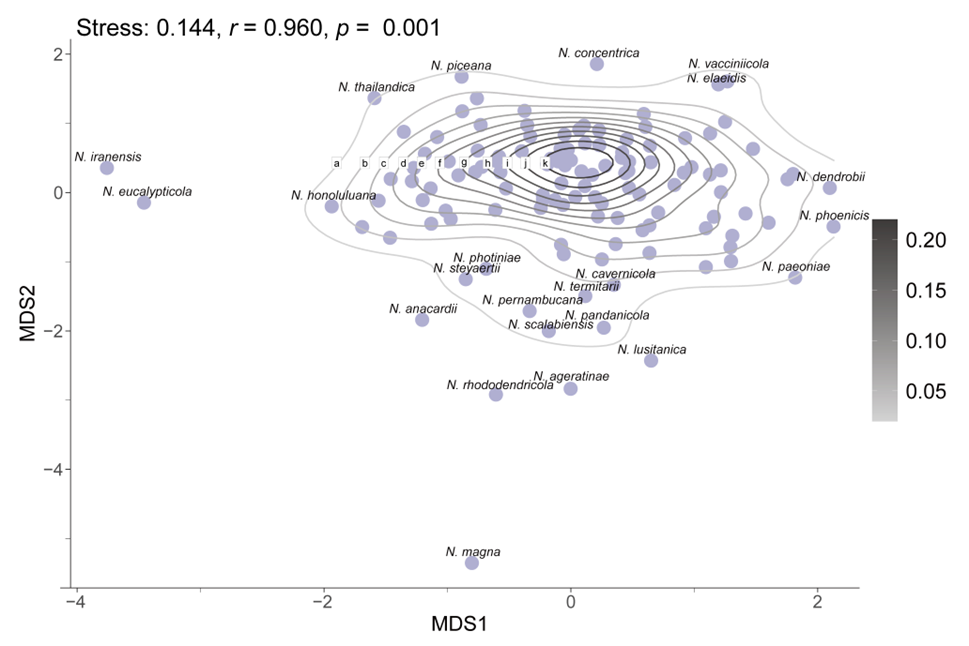
To assess morphological variation, MDS was applied using a distance matrix to reduce dimensionality while preserving the relative relationships among species. The resulting MDS plot reflects the original morphological similarities and differences through the spatial arrangement of data points (Fig. 6). The analysis produced a stress value of 0.144, indicating that the dimensionality reduction effectively maintains the structural integrity of the dataset. Additionally, a high r-value of 0.96 demonstrates a strong positive correlation between the Euclidean distances in the original dataset and those in the reduced-dimensional space, confirming the accuracy of the MDS configuration. The statistical significance of these findings is further supported by a p-value of 0.001 (p < 0.05). The MDS plot indicates that while certain species, such as N. eucalypticola, N. iranensis, and N. magna, exhibit distinct morphological traits, the majority display close relative distances or overlapping morphological distributions.
Taxonomy of Neopestalotiopsis
Neopestalotiopsis Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 135 (2014)
Notes: Neopestalotiopsis was introduced and typified with Neopestalotiopsis protearum by Maharachchikumbura et al. (2014b). The genus is distinguished from Pestalotiopsi_s by its versicolourous median cells (Maharachchikumbura et al. 2014b). Members of _Neopestalotiopsis are predominantly distributed in tropical and subtropical ecosystems, functioning mainly as plant pathogens (Santos et al. 2020; Shi et al. 2022; Qi et al. 2023a; Rajashekara et al. 2023), but they also occur as endophytes (Freitas et al. 2019; Ma et al. 2019; Zhang et al. 2024b) or saprobes (Maharachchikumbura et al. 2014b).
Over the past few years, the number of newly described Neopestalotiopsis species has increased rapidly, with most species’ delimitations relying primarily on the concatenated ITS-tef1-tub2 phylogenetic trees (Sun et al. 2023; Cui et al. 2024; Razaghi et al. 2024). However, our analyses revealed that in these three-locus trees, the overall branch lengths were notably short and often supported by low bootstrap values. Moreover, the topologies of these trees were largely incongruent with the whole-genome phylogeny, and many described taxa were indistinguishable. Genomic analyses further demonstrated that Neopestalotiopsis exhibits a pattern of over-splitting. Strains with ANI values exceeding 97.30% and sharing more than 88% of their core gene clusters likely represent conspecifics, despite having been described as distinct species based on three-locus data. Considering the large number of named taxa and the limited availability of genomic data, we adopted a conservative approach by synonymizing only those taxa that exhibited nearly identical sequences and closely related phylogenetic positions. Accordingly, seven previously described taxa were synonymized based on integrated evidence from whole-genome analyses, phylogenetic inference, species delimitation results, and sequence similarity. Specifically, N. acericola was synonymized with N. caulicola; N. amomi and N. olivaceous were synonymized with N. zingiberis; N. fragariae and N. umbrinospora were synonymized with N. chrysea; and N. hadrolaeliae and N. zimbabwana were synonymized with N. honoluluana. In addition, a new species, N. camelliae, was introduced based on combined evidence from multi-locus and genome-scale phylogenetic analyses.
Neopestalotiopsis camelliae Q. Zhang & Yong Wang bis, sp. nov.
Index Fungorum number: IF904599; Fig. 7
Etymology: named after the host genus, Camellia.
Holotype: HGUP 23-0290
Associated with diseased leaves of Camellia japonica.
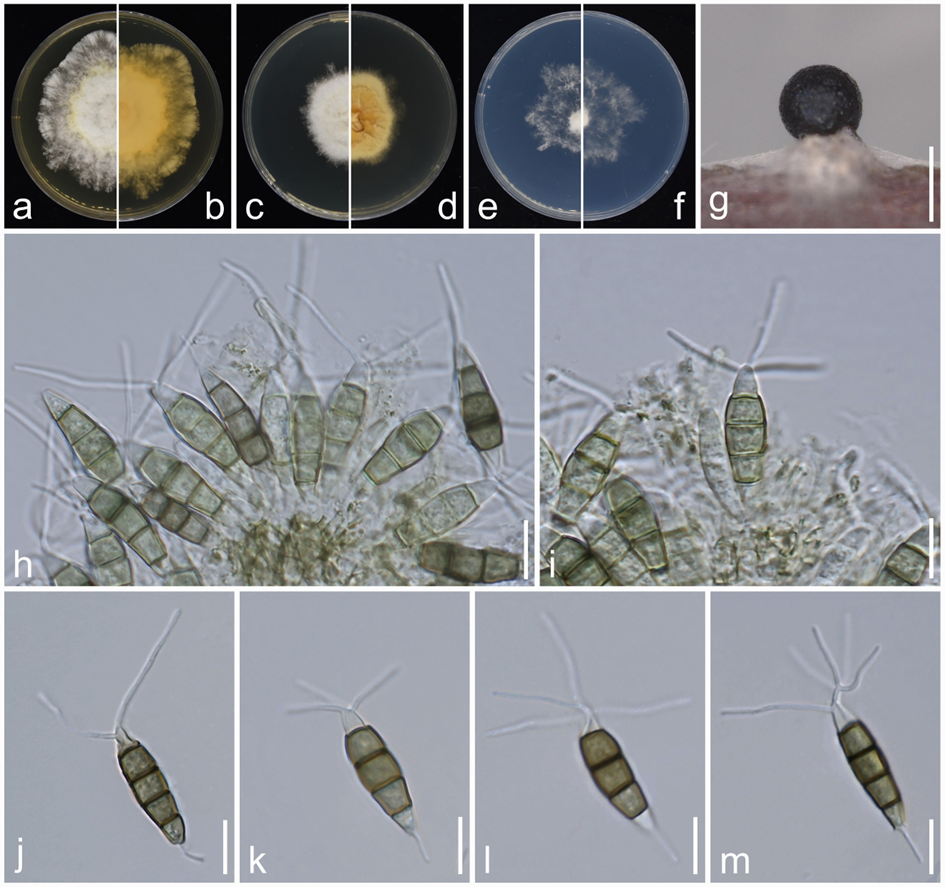
Sexual morph: Undetermined. Asexual morph: Conidiophores when present 1–2 septate, hyaline, thin-walled, often reduced to conidiogenous cells. Conidiogenous cells ampulliform, cylindrical, hyaline, thin walled, 2.5–15 × 1.5–2.5 μm. Conidia fusoid, ellipsoid, straight to slightly curved, 4-septate, not constricted at septa, 20.5–28.5 × 5–7 μm (av. ± SD = 23.50 ± 1.79 × 5.88 ± 0.45 μm); basal cell obconic to obtuse, with a truncate base, hyaline to pale olivaceous, rugose and thin-walled, 3–6 μm (av. ± SD = 4.45 ± 0.66 μm) long; three median cells doliiform to subcylindrical, 13.5–18.5 μm (av. ± SD = 15.58 ± 1.02 μm) long, concolourous, but occasionally the second cell lighter than the other cells, olivaceous to pale brown, (second cell from base 4–6 μm long; third cell 4.5–6 μm long; fourth cell 4.5–7 μm long), septa darker than the rest of cells; apical cell 3–6 μm (av. ± SD = 4.38 ± 0.73 μm) long, hyaline, conic to broad or long conic, rugose and thin-walled; with 2–4 tubular apical appendages (mostly 3), arising from the apical crest, with some branched appendages, filiform, flexuous, 7.5–28.5 μm (av. ± SD = 17.6 ± 5.19 μm) long; single basal appendage, tubular, centric, unbranched, 2–9 μm (av. ± SD = 5.18 ± 1.57 μm) long.
Culture characteristics: Colonies on MEA raised with erose or dentate edge, smooth, colony from above white to buff-white, from below initially white, pale yellowish in centre, aerial mycelia flocculent, reaching 60–64 mm diam after 7 d at 25°C; on PDA convex or dome-shaped, with erose or dentate edge, smooth to somewhat radiated, dense, colony from above white to buff-white, from below yellow, reaching 40–43 mm diam after 7 d at 25°C; on SNA flat with undulate edge or with concave edge, colony from above and from below white, aerial mycelia flocculent, reaching 44–47 mm diam after 7 d at 25°C.
Material examined: China, Guizhou Province, Shuicheng, Yushe National Forest Park, on leaf spot of Camellia japonica, 29 Apr. 2023, Q. Zhang, Q126 (HGUP 23-0290, holotype), ex-type GUCC 23-0312; ibid., GUCC 23-0313.
Notes: Two isolates of Neopestalotiopsis camelliae (GUCC 23-0312 and GUCC 23-0313) formed a sister clade with N. zingiberis in the three-gene concatenated phylogenetic tree (Fig. 1). In the ITS and tub2 phylogenies, these isolates clustered together with a distinct branch length (Fig. S1a, c), which is consistent with the GCPSR principle, thereby supporting their designation as a separate evolutionary lineage. Both PTP and mPTP analyses produced congruent results, grouping isolates GUCC 23-0312 and GUCC 23-0313 into a single MOTU. The ex-type culture of N. camelliae (GUCC 23-0312) exhibited the following nucleotide similarities with the ex-type culture of N. zingiberis (HGUP 10001): 96.18% (478/497, including six gaps) in the ITS region, 94.38% (420/445, no gaps) in the tef1 region, and 98.34% (771/784, including eight gaps) in the tub2 region. Additionally, in the whole-genome phylogenetic tree, N. camelliae formed a well-supported sister clade with N. zingiberis (HGUP 10001) (Fig. 3), with SH-aLRT = 100% and UFBoot = 100%. The ANI between N. camelliae GUCC 23-0312 and N. zingiberis HGUP 10001 was 95.07% (Fig. 3, Table S5). Morphologically, N. camelliae differs from N. zingiberis by having longer apical appendages (7.5–28.5 μm vs. 12–15 μm) (He et al. 2022). Therefore, based on both multi-locus phylogenetic analyses and whole-genome evidence, we describe Neopestalotiopsis camelliae as a novel species.

Neopestalotiopsis caulicola H. Zhang & Y.L. Jiang, Journal of Systematics and Evolution 62 (4): 643 (2024).
= Neopestalotiopsis acericola W.S. Zhang & X.L. Fan, Journal of Fungi 10 (7, no. 475): 6 (2024).
See Zhang et al. (2024b) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Guizhou Province, Guiyang City, from healthy stems of Rosa roxburghii, 22 April 2020, H. Zhang (HGUP 191003, holotype); ex-type GUCC 191003.1.
Host range: Acer palmatum (Zhang et al. 2024d), Rosa roxburghii (Zhang et al. 2024b).
Known distribution: China (Zhang et al. 2024b; Zhang et al. 2024d).
Notes: Neopestalotiopsis caulicola was identified from Rosa roxburghii in China (Zhang et al. 2024b). Neopestalotiopsis acericola was introduced from Acer palmatum in China (Zhang et al. 2024d). Although the phylogenetic relationships within Neopestalotiopsis remain unstable, N. caulicola and N. acericola consistently cluster together. Phylogenetic analyses based on the concatenated dataset of three genes (Fig. 1; BS/PP = 95%/0.97) and the tef1 phylogenetic tree (Fig. S1b; BS/PP = 85%/0.99) revealed that these two species form a strongly supported clade. In the ITS and tub2 phylogenies, these two species also cluster together, though with the inclusion of other species such as N. siciliana (Fig. S1a, c). The ex-type culture of N. acericola (CFCC 70620) exhibited high nucleotide similarity with the ex-type culture of N. caulicola (GUCC 191003.1): 99.34% for ITS (449/452, including three gaps), 99.86% for tef1 (696/697, no gaps), and 100% for tub2 (702/702). Morphologically, their primary difference in conidial width, with N. acericola measuring 6.5–8.0 μm and N. caulicola ranging from 4.0–6.5 μm (Zhang et al. 2024b; 2024d). However, we do not consider such a minor morphological difference to be a reliable criterion for species delimitation (see discussion). Furthermore, since N. caulicola was published online approximately six months earlier than N. acericola, it was absent from the dataset when N. acericola was introduced as a new species. Given the strong phylogenetic and molecular evidence, we formally synonymize N. acericola under N. caulicola.
Neopestalotiopsis chrysea (Maharachch. & K.D. Hyde) Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 138 (2014)
= Neopestalotiopsis fragariae Prematunga & Jayaward., Asian Journal of Mycology 5 (10): 230 (2022)
= Neopestalotiopsis umbrinospora (Maharachch. & K.D. Hyde) Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 149 (2014)
See Maharachchikumbura et al. (2012) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Guangxi Province, Shangsi, Shiwandashan, Wangle, dead leaves of unidentified plant, 2 January 1997, Wenping Wu WUFH1303a (HMAS042855, holotype; MFLU 12-0411, isotype); ex-type NN042855 = MFLUCC 12-0261.
Host range: Fragaria × ananassa (Prematunga et al. 2022), Liquidambar formosana (Fan et al. 2022), Unidentified host (Maharachchikumbura et al. 2012).
Known distribution: China (Maharachchikumbura et al. 2012; Fan et al. 2022; Prematunga et al. 2022).
Notes: Neopestalotiopsis chrysea was introduced by Maharachchikumbura et al. (2012) as Pestalotiopsis chrysea on dead plant material in China and was later accommodated in Neopestalotiopsis by Maharachchikumbura et al. (2014b). Similarly, Neopestalotiopsis umbrinospora was first described as Pestalotiopsis umbrinospora by Maharachchikumbura et al. (2012) and subsequently reclassified within Neopestalotiopsis by Maharachchikumbura et al (2014b). Phylogenetic analyses based on the concatenated dataset of three-gene (Fig. 1; BS/PP = 87%/0.98) and the tub2 gene (Fig. S1c; BS/PP = 83%/0.99) strongly support the clustering of these two species, as confirmed by both ML and BI analyses. Moreover, they cluster together and cannot be distinctly differentiated from other lineages based on ITS and tef1 sequences. The ex-type culture of N. chrysea (MFLUCC 12-0261) exhibits high nucleotide similarity with the ex-type culture of N. umbrinospora (MFLUCC 12-0285): 100% for ITS (482/482), 99.79% for tef1 (949/951, including 0 gaps), and 99.56% for tub2 (450/452, including 0 gaps). In the three-gene phylogenetic tree, N. fragariae clustered close to N. chrysea. In the whole-genome phylogeny, N. fragariae GUCC 23-0201 grouped with the ex-type strain of N. chrysea (MFLUCC 12-0261), with an ANI value of 98.41% and a core-gene proportion of 97.48% (Fig. 3). The ex-type culture of N. chrysea (MFLUCC 12-0261) and N. fragariae (ZHKUCC 22-0113) also exhibit high sequence similarity: 99.79% for ITS (481/482), 99.32% for tef1 (291/293, including 0 gaps), and 99.78% for tub2 (446/447, including 0 gaps). Morphologically, N. chrysea, N. fragariae and N. umbrinospora exhibit nearly overlapping phenotypic characteristics (Maharachchikumbura et al. 2012; Prematunga et al. 2022). Based on these findings, we formally synonymize N. fragariae and N. umbrinospora under N. chrysea, and the corresponding synonymy is provided.
Neopestalotiopsis honoluluana Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 141 (2014).
= Neopestalotiopsis hadrolaeliae E.F.S. Freitas, Meir. Silva & M.C.M. Kasuya, Phytotaxa 416 (3): 215 (2019)
= Neopestalotiopsis zimbabwana Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 149 (2014)
See Maharachchikumbura et al. (2014b) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: USA, Hawaii, Honolulu, from Telopea sp., 8 December 1998, P.W. Crous & M.E. Palm (CBS H-21771, holotype); ex-type CBS 114495 = STE-U 2076.
Host range: Telopea sp. (Maharachchikumbura et al. 2014b), Hadrolaelia jongheana (Freitas et al. 2019), Leucospermum cunciforme (Maharachchikumbura et al. 2014b), Xylaria sp. (Hermawan et al. 2021).
Known distribution: America (Maharachchikumbura et al. 2014b), Brazil (Freitas et al. 2019), Zimbabwe (Maharachchikumbura et al. 2014b).
Notes: Neopestalotiopsis honoluluana was introduced by Maharachchikumbura et al. (2014b) from Telopea sp. in America, while N. zimbabwana was described by the same authors from Leucospermum cuneiforme in Zimbabwe. Neopestalotiopsis hadrolaeliae was later introduced by Freitas et al. (2019) from Hadrolaelia jongheana in Brazil. Phylogenetic analyses of the concatenated three-gene dataset (Fig. 1; BS/PP = 94%/0.98) and the tub2 gene (Fig. S1c; BS/PP = 86%/0.97) strongly support these three species forming a distinct cluster, with both ML and BI trees consistently supporting this relationship in both datasets.
In the ITS phylogenetic tree (Fig. S1a), they could not be clearly distinguished from other lineages, whereas in the tef1 phylogenetic tree (Fig. S1b), they form a distinct clade together with N. eucalypticola (CBS 264.37). Sequence comparisons between the type strains of N. hadrolaeliae, N. honoluluana, and N. zimbabwana showed 99.44–99.75% nucleotide similarity for ITS, 99.15–99.58% for tef1, and 98.84–99.87% for tub2. Morphologically, their phenotypic spectra largely overlap, with differences mainly in conidial size: N. hadrolaeliae (19–26.5 × 5–7.5 μm) has smaller conidia compared to N. honoluluana (21–35 × 7–10 μm) and N. zimbabwana (22–30 × 6.5–9 μm) (Maharachchikumbura et al. 2014b; Freitas et al. 2019). However, due to the morphological variability within Neopestalotiopsis, relying solely on morphological traits is inadequate for species delimitation, rendering molecular data essential for classification (see discussion). Based on these findings, we formally synonymize N. hadrolaeliae and N. zimbabwana under N. honoluluana.
Neopestalotiopsis zingiberis Y.K. He & Yong Wang bis, Biodivers. Data J. 10 (e90709): 10 (2022)
= Neopestalotiopsis amomi Y.R. Sun & Yong Wang bis, Microbiol. Spectrum 11 (1): e03987-22, 2 (2023)
= Neopestalotiopsis olivaceous X.F. Cui & Z.G. Hao, J. Fungi 10 (6, no. 371): 15 (2024)
See He et al. (2022) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Hainan Province, Haikou City, Wuzhishan Nature Reserve, from leaf blight of Zingiber officinale, 2020, Y.K. He (HGUP 10001, holotype); ex-type GUCC 21001.
Host range: Alpinia oxyphylla (Cui et al. 2024), Amomum villosum (Sun et al. 2023), Zingiber officinale (He et al. 2022).
Known distribution: China (He et al. 2022; Sun et al. 2023; Cui et al. 2024).
Notes: Neopestalotiopsis zingiberis was first described from Zingiber officinale in China (He et al. 2022), while N. amomi was isolated from diseased leaves of Amomum villosum in China (Sun et al. 2023). More recently, N. olivaceous was introduced from Alpinia oxyphylla in China (Cui et al. 2024). In the concatenated three-gene phylogenetic analysis (Fig. 1), N. zingiberis, N. amomi, and N. olivaceous form a strongly supported clade (BS/PP = 100%/1). Additionally, they cluster together in both the ITS and tef1 phylogenies with high support (BS ≥ 99%, PP = 1; Fig. S1a, b), fulfilling the GCPSR principle for recognition as a single evolutionary lineage. Species delimitation analyses using PTP and mPTP further support their classification as a single species (Fig. 1). Morphologically, these taxa display some variation, primarily in the coloration of the three median cells and their length conidiophores. Neopestalotiopsis amomi has concolourous three median cells and short conidiophores (3–5 μm) (Sun et al. 2023), whereas N. zingiberis has versicolorous three median cells and longer conidiophores (12–25 μm) (He et al. 2022). Neopestalotiopsis olivaceous also possesses concolourous three median cells (Cui et al. 2024). Sequence comparisons reveal high similarity among these taxa, with ITS sequence identity ranging from 99.17% to 99.79%, tef1 from 98.48% to 99.73%, and tub2 from 98.05% to 100%. Given that N. zingiberis was described prior to N. amomi and N. olivaceous, we propose synonymizing N. amomi and N. olivaceous under N. zingiberis.
Pestalotiopsis
Phylogenetic analyses of Pestalotiopsis
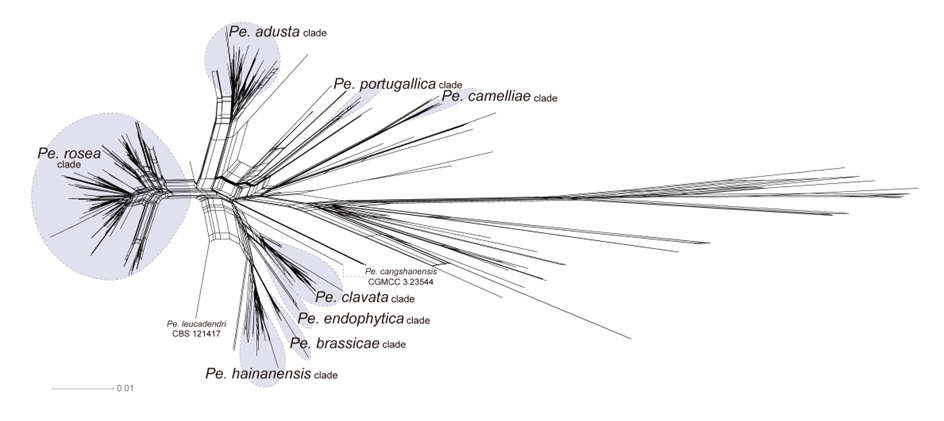
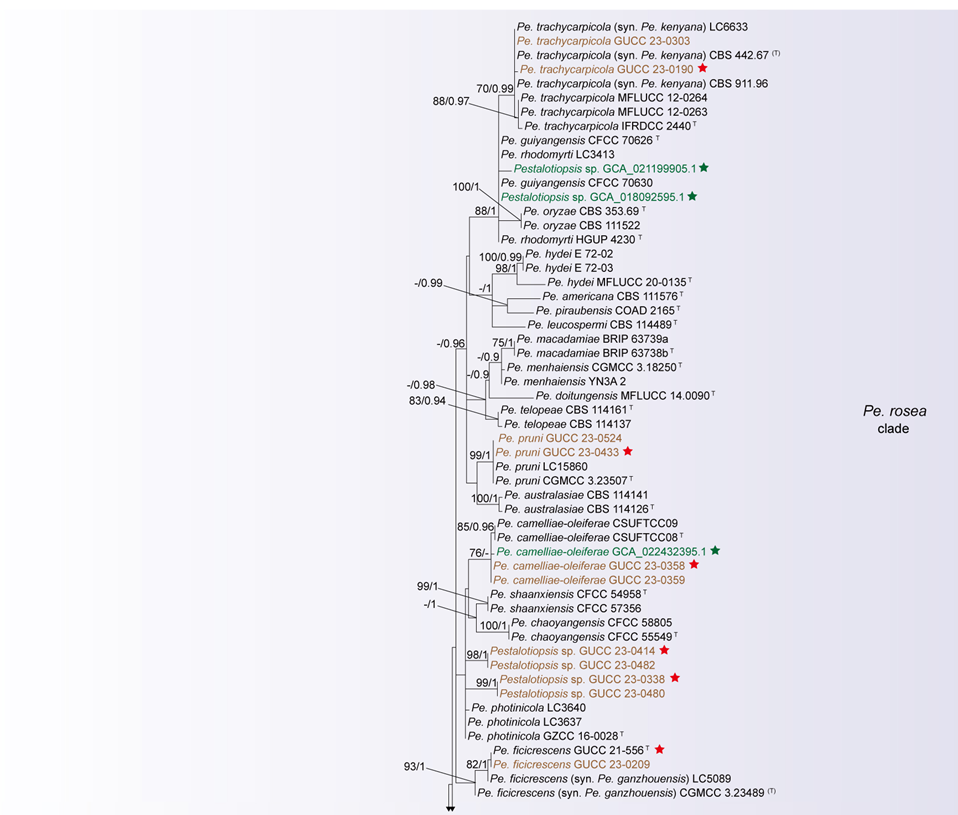
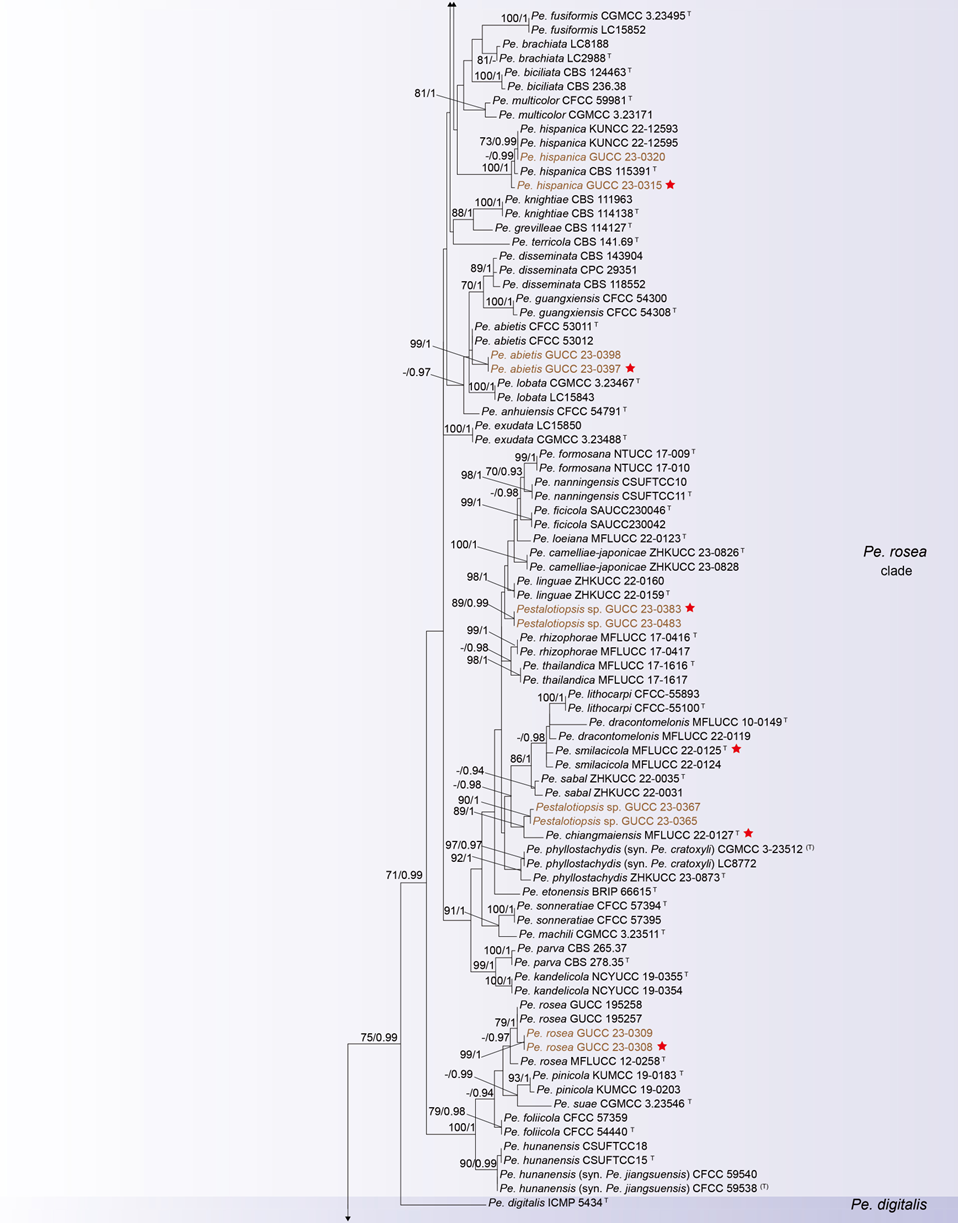
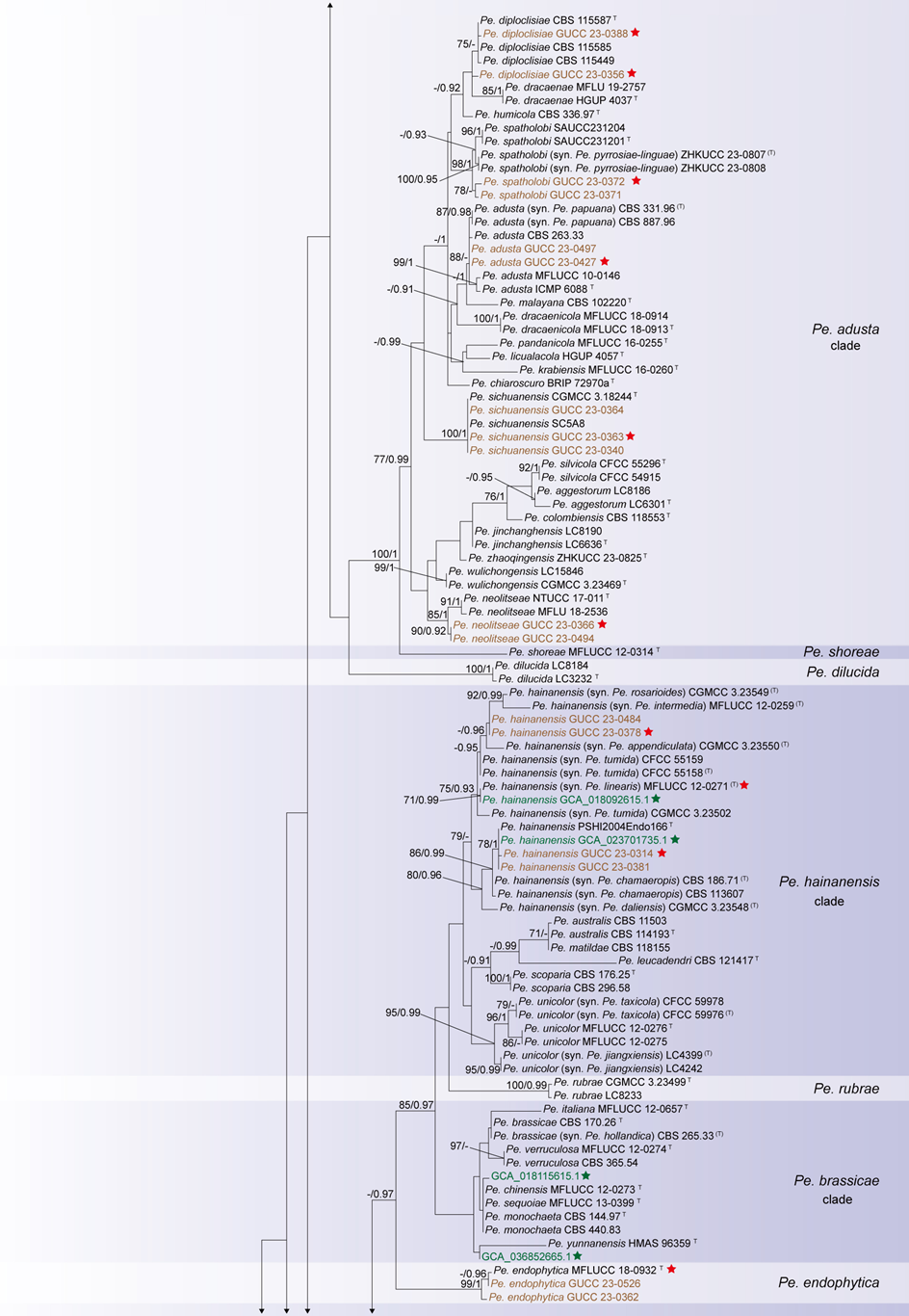
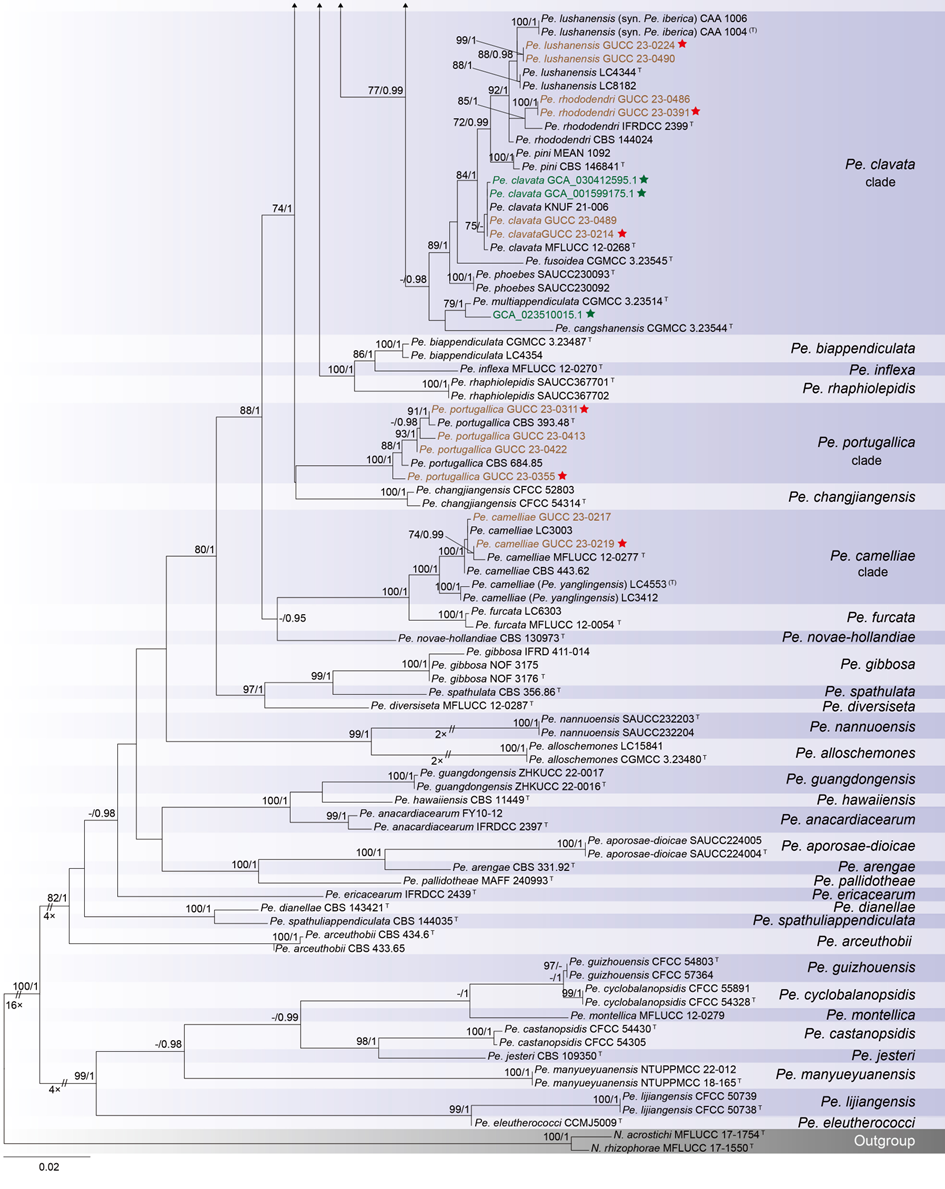
The concatenated ITS-tef1-tub2 dataset for the genus Pestalotiopsis included 319 isolates, comprising 50 newly collected in this study and ten retrieved from the NCBI genome database. Initially, 11 Pestalotiopsis genomes were obtained from NCBI. However, phylogenetic analyses based on both multi-gene (Fig. 26) and whole-genome (Fig. 28) datasets revealed that GCA_000516985.1 is positioned within Pseudopestalotiopsis. Phylogenetic network analyses (Fig. 8) based on these isolates revealed that the 60 isolates clustered into eight distinct clades (Pe. adusta, Pe. brassicae, Pe. camelliae, Pe. clavata, Pe. endophytica, Pe. hainanensis, Pe. portugallica, and Pe. rosea clades), which are labeled according to the earliest typified species for ease of reference and visualization, rather than representing formal taxonomic designations. When two outgroup taxa (Neopestalotiopsis acrostichi MFLUCC 17-1754 and N. rhizophorae MFLUCC 17-1550) were included, ML and BI analyses produced similar tree topologies, each showing well-resolved clades for all analyzed species, generally supported by high bootstrap and posterior probability values. The ML tree is presented to illustrate these findings (Fig. 9). Phylogenetic analyses employing both the concatenated dataset and single-locus datasets revealed that most of the eight major clades constituted distinct, independent clusters in both the combined and individual gene trees (Fig. 9 and Fig. S2a–c). However, exceptions were observed. For instance, in the ITS phylogenetic tree (Fig. S2a), a few taxa from the Pe. adusta, Pe. brassicae, Pe. hainanensis, and Pe. rosea clades did not cluster with the majority of isolates within their respective clades. Similarly, in the tub2 tree (Fig. S2c), certain taxa from the Pe. clavata and Pe. hainanensis clades were separated from the majority of isolates within their respective branches. For example, in the tub2 phylogenetic tree, Pe. leucadendri CBS 121417 (assigned to the P_e. hainanensis_ clade) and Pe. cangshanensis CGMCC 3.23544 (assigned to the Pe. clavata clade) were clustered within the Pe. rosea clade. We recommend a careful re-examination of their tub2 sequences to rule out possible sequencing or data processing errors. If no errors are detected, this incongruent placement may be attributed to factors such as incomplete lineage sorting, or gene introgression events. Subsequently, all eight Pestalotiopsis clades, along with the closely related outgroup taxa, were analyzed in greater detail to refine species boundaries and ensure the accurate assignment of isolates to their respective Pestalotiopsis species. Due to the limited species diversity within the Pe. endophytica and Pe. portugallica clades, the corresponding results for these two lineages were incorporated into the analyses of the Pe. brassicae and Pe. camelliae clades, respectively.
Whole-genome data and phylogenomic assessment of Pestalotiopsis
This study analyzed 38 genomes of Pestalotiopsis, including ten publicly available genomes from the NCBI database and 28 new sequences. These genomes represent eight distinct clades within Pestalotiopsis, comprising four single species and four species complexes. A total of 7,936 single-copy orthologous genes were identified and used for phylogenomic reconstruction, incorporating 38 Pestalotiopsis strains together with two outgroups, Neopestalotiopsis acrostichi MFLUCC 17-1754 and N. rhizophorae MFLUCC 17-1550. The resulting phylogenomic tree recovered the same eight well-defined clades as the three-locus phylogeny based on ITS, tef1, and tub2 (Figs. 9–10). However, minor discrepancies were observed in the placement of certain strains within the Pe. rosea and Pe. clavata clades when comparing the whole-genome and three-gene phylogenies (Figs. 9–10). For instance, strains GUCC 23-0363 and GUCC 23-0366 were positioned distantly from each other in the three-gene phylogenetic tree but formed a sister lineage with extremely short branch lengths in the whole-genome phylogeny, exhibiting a high ANI value of 99.31%.
Across Pestalotiopsis, ANI values ranged from 89.50% to 99.48% (Fig. 10 and Table S5). Genomic similarity among the eight major clades largely corresponded with their positions in the three-gene and network analyses, but some variation was evident. Within the Pe. adusta clade, six genomes showed ANI values of 96.29–99.31%, with the highest similarity (99.31%) between Pe. sichuanensis GUCC 23-0363 and Pe. neolitseae GUCC 23-0366. The Pe. brassicae clade, comprising two genomes (GCA_018115615.1 and GCA_036852665.1), exhibited an ANI of 98.97%. The Pe. camelliae clade contained a single genome, which shared less than 89.94% similarity with other members of the genus. In the Pe. clavata clade, ANI values among six genomes ranged from 97.47% to 99.48%. The Pe. endophytica clade included only the type strain MFLUCC 18-0932, which showed less than 94.66% similarity to other species. The Pe. hainanensis clade, comprising five genomes, exhibited ANI values of 97.73–99.45%, while the Pe. portugallica clade (two genomes) showed 97.83% similarity. The Pe. rosea clade, with 15 genomes, displayed ANI values ranging from 94.36% to 99.22%.
The proportion of the core genome within the pan-genome was further assessed across six major clades of Pestalotiopsis. Specifically, six genomes from the Pe. adusta clade, two from Pe. brassicae, six from Pe. clavata, five from Pe. hainanensis, two from Pe. portugallica, and 15 from Pe. rosea were analyzed. The core genome accounted for 90.88% (Fig. 11a), 91.49% (Fig. 11b), 90.29% (Fig. 11c), 91.12% (Fig. 11d), 96.33% (Fig. 11e), and 83.83% (Fig. 11f) of the total pan-genome orthogroups, respectively, indicating that members within each clade share a high degree of genomic conservation.
Morphology of Pestalotiopsis
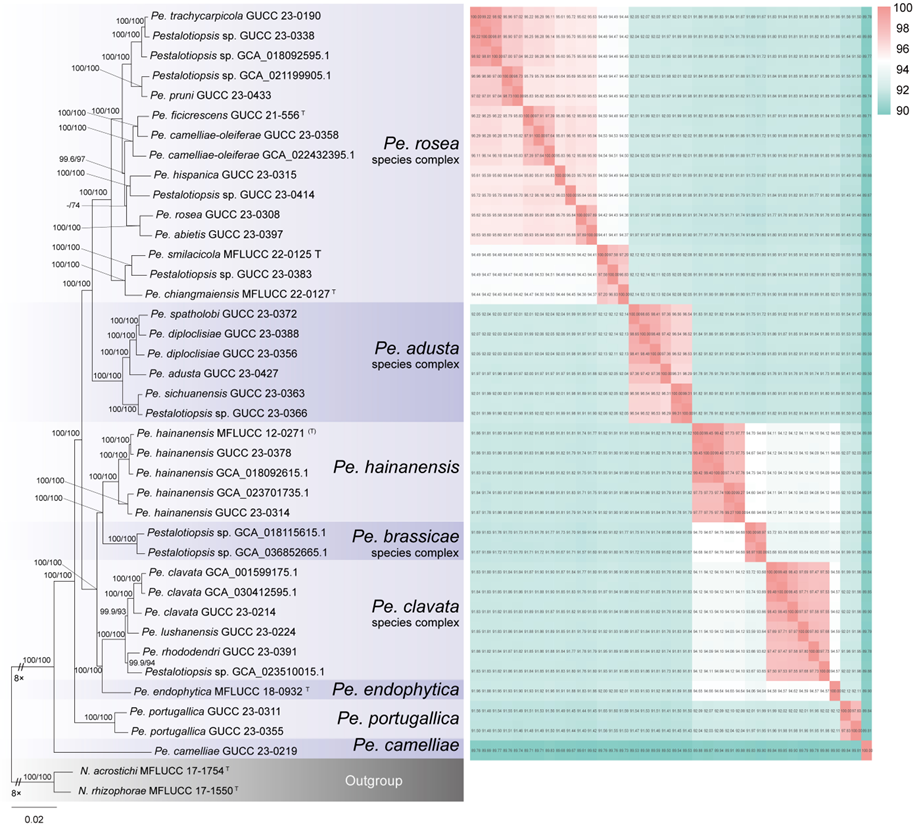
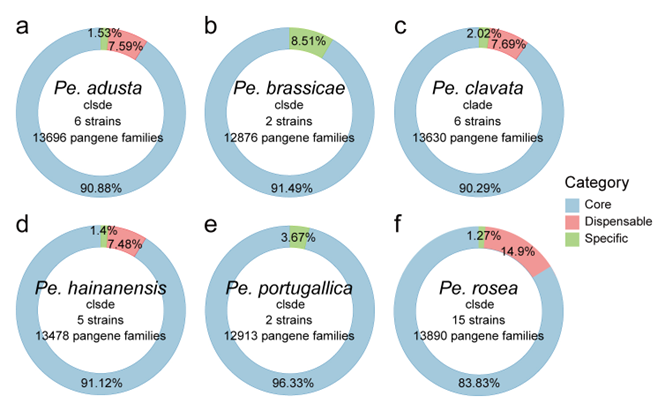
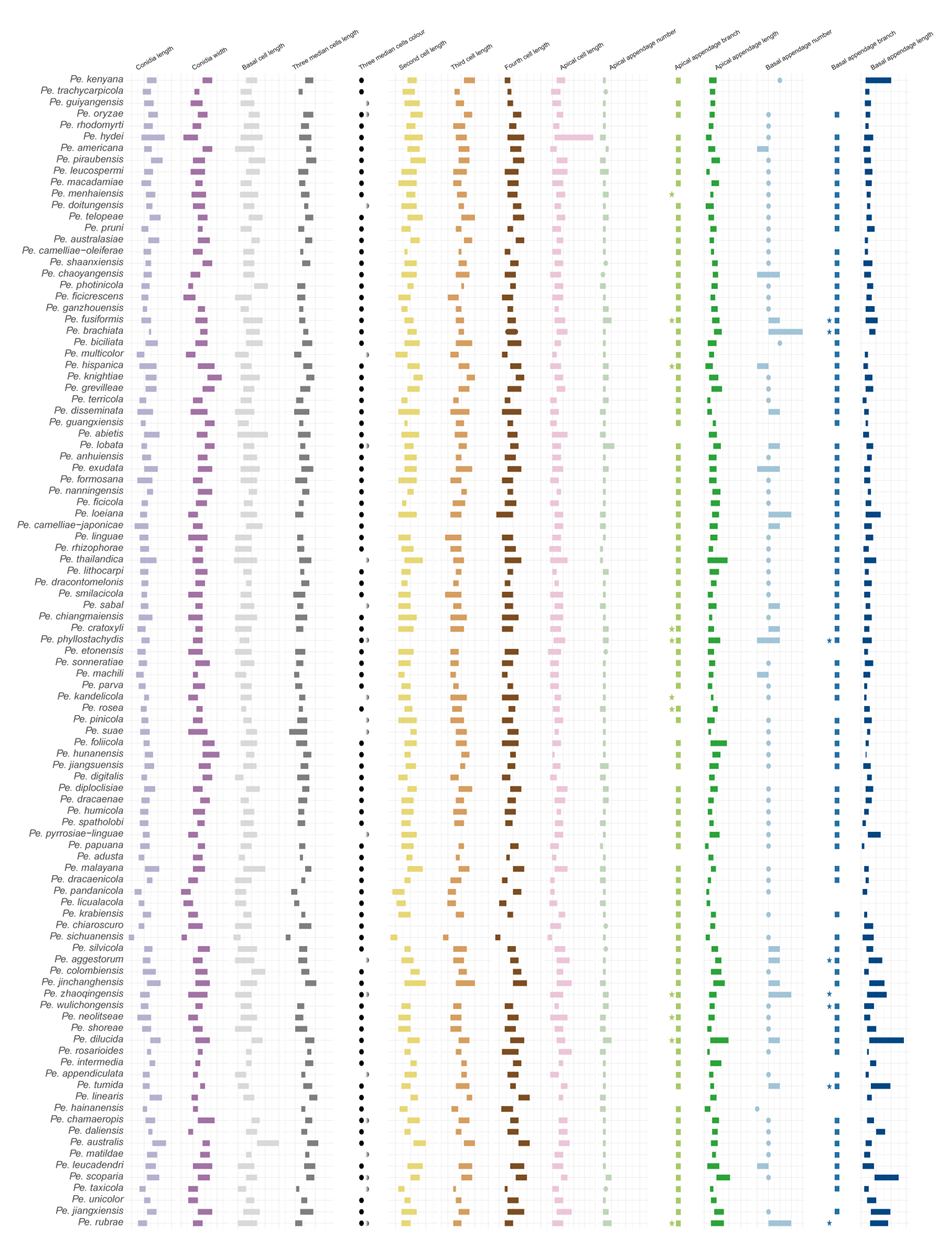
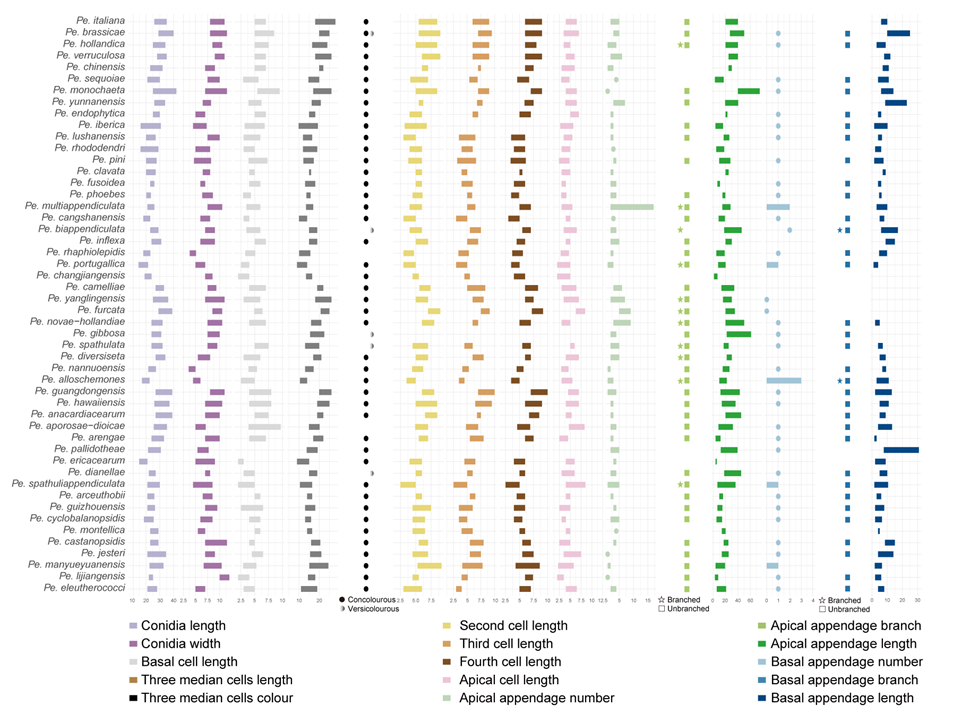
Figure 12 provides an overview of the micromorphological dimensions for each Pestalotiopsis species, arranged according to their positions in the three-gene concatenated phylogenetic tree. Notably, discrepancies between molecular and morphological observations are noted. Some species that are phylogenetically closely related display morphological differences. For instance, Pe. kenyana and Pe. trachycarpicola are positioned near each other in the three-gene phylogenetic tree (Fig. 9), yet Pe. kenyana has significantly wider conidia and longer three median cells and basal appendages compared to Pe. trachycarpicola (Fig. 12). Similar patterns are observed in Pe. ficicrescens and Pe. ganzhouensis, Pe. spatholobi and Pe. pyrrosiae-linguae, Pe. brassicae and Pe. hollandica, as well as Pe. chinensis, Pe. sequoiae, and Pe. monochaeta. Conversely, species exhibiting highly similar morphological traits may sometimes be phylogenetically distant (Figs. 9, 12), such as Pe. americana and Pe. arceuthobii, Pe. kandelicola and Pe. appendiculata, and Pe. rhizophorae and Pe. spatholobi. Across the genus Pestalotiopsis, there is considerable morphological overlap among species (Fig. 12, Table S12). Conidial length ranges from 8.6 to 42 μm, while width varies from 2.5 to 9.5 μm. Most species possess concolourous three median cells; however, 25 species, including Pe. aggestorum, Pe. appendiculata, Pe. biappendiculata, Pe. brassicae, Pe. chamaeropis, Pe. dianellae, Pe. doitungensis, Pe. gibbosa, Pe. guiyangensis, Pe. kandelicola, Pe. lobata, Pe. matildae, Pe. multicolor, Pe. oryzae, Pe. phyllostachydis, Pe. pinicola, Pe. pyrrosiae-linguae, Pe. rubrae, Pe. sabal, Pe. scoparia, Pe. spathulata, Pe. suae, Pe. taxicola, Pe. thailandica, and Pe. zhaoqingensis, exhibit versicolorous three median cells. Variations in apical and basal appendage characteristics are evident among certain species. Apical appendage length ranges from 1 to 75 μm, with the number of appendages varying from 1 to 17. Basal appendage length spans from 0.5 to 30.8 μm. Despite these morphological differences, there is no clear correlation observed between morphological traits and phylogenetic relationships.
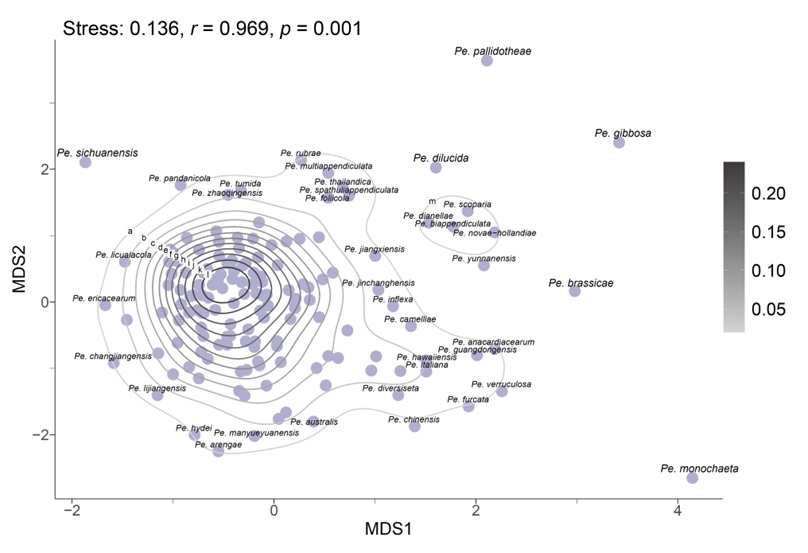
In the resulting MDS plot (Fig. 13), the spatial arrangement of data points reflects the original morphological similarities and differences. The analysis produced a stress value of 0.14, indicating that the dimensionality reduction effectively maintains the structural integrity of the dataset. Moreover, the high r-value of 0.97 indicates a strong positive correlation between the Euclidean distances in the original dataset and those in the reduced-dimensional space, thereby confirming the accuracy of the MDS configuration. The statistical significance of the results is further supported by a p-value of 0.001 (p < 0.05), indicating that the observed morphological patterns are not attributable to random variation. The MDS plot reveals that while some species, such as Pe. brassicae, Pe. gibbosa, Pe. monochaeta, Pe. pallidotheae, and Pe. sichuanensis, exhibit distinct morphological traits, the majority show close relative distances or overlapping morphological distributions.
Taxonomy of Pestalotiopsis
Pestalotiopsis Steyaert, Bull. Jard. Bot. État Bruxelles 19 (3): 300 (1949)
Notes: Pestalotiopsis was established by Steyaert (1949). Species of Pestalotiopsis are cosmopolitan, occurring as saprobes, endophytes, or opportunistic pathogens on a wide range of economically important and ornamental plants (Hyde et al. 2020b). Several studies have examined the diversity of this genus based on ITS, t_ub2_, and tef1 gene regions (Liu et al. 2019; Peng et al. 2022; Sun et al. 2023). However, our analyses revealed several evolutionary lineages characterized by short branch lengths and low terminal bootstrap and posterior probability values, indicating limited phylogenetic resolution. Integrating evidence from both multi-locus and genome-scale phylogenetic analyses suggests that Pestalotiopsis is likely affected by taxonomic over-splitting. To accommodate lineages that form well-supported monophyletic groups in both multi-locus and whole-genome phylogenies but exhibit short internal branches, low support values, and indistinct species boundaries—and for which there is insufficient evidence for formal synonymization—we herein recognize four species complexes: the Pe. adusta species complex, Pe. brassicae species complex, Pe. clavata species complex, and Pe. rosea species complex. Furthermore, based on combined phylogenetic topology, sequence similarity, and genomic evidence, eighteen species with highly similar sequences and overlapping phylogenetic positions are synonymized as follows: Pe. appendiculata, Pe. chamaeropis, Pe. daliensis, Pe. intermedia, Pe. linearis, Pe. rosarioides, and Pe. tumida are synonymized under Pe. hainanensis; Pe. cratoxyli under Pe. phyllostachydis; Pe. ganzhouensis under Pe. ficicrescens; Pe. hollandica under Pe. brassicae; Pe. iberica under Pe. lushanensis; Pe. jiangsuensis under Pe. hunanensis; Pe. jiangxiensis and Pe. taxicola under Pe. unicolor; Pe. kenyana under Pe. trachycarpicola; Pe. papuana under Pe. adusta; Pe. pyrrosiae-linguae under Pe. spatholobi; and Pe. yanglingensis under Pe. camelliae.
The Pestalotiopsis adusta clade
According to the phylogenetic analyses of the concatenated alignment, the 11 isolates obtained in this study clustered within a strongly supported clade (77% BS / 0.99 PP; Fig. 9), here designated as the Pe. adusta clade. To provide a clear comparison, a phylogenetic tree for the Pe. adusta clade, as well as individual gene trees, was constructed (Fig. S3a–c). These trees include 47 ingroup taxa and one closely related outgroup taxa (Table S6). The concatenated alignment consisted of 1,777 characters, derived from three loci: 531 characters from ITS, 496 from tef1, and 750 from tub2 (including alignment gaps). Detailed alignment characteristics for the ML and BI analyses of the Pe. adusta clade are summarized in Table S6.
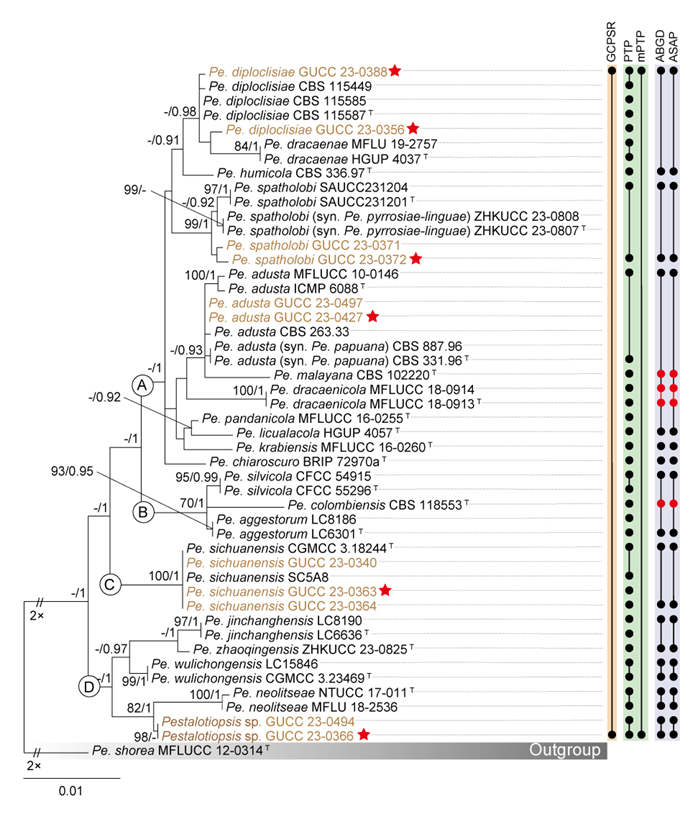
The phylogenetic trees constructed using maximum likelihood (ML) and Bayesian inference (BI) from the concatenated dataset display similar topologies. Therefore, only the ML trees are presented here, with well-supported nodes clearly indicated (Fig. 14). The phylogenetic tree derived from the combined dataset reveals that the Pe. adusta clade comprises four distinct subclades. Subclade A includes Pe. adusta, Pe. chiaroscuro, Pe. diploclisiae, Pe. dracaenae, Pe. dracaenicola, Pe. humicola, Pe. krabiensis, Pe. licualacola, Pe. malayana, Pe. pandanicola, Pe. papuana, Pe. pyrrosiae-linguae, and Pe. spatholobi. Subclade B consists of Pe. aggestorum, Pe. colombiensis, and Pe. silvicola. Subclade C is represented by Pe. sichuanensis, while subclade D includes Pe. jinchanghensis, Pe. neolitseae, Pe. wulichongensis, and Pe. zhaoqingensis. This detailed subclade structure underscores the genetic diversity within the Pe. adusta clade.
The individual ML and BI gene trees were analyzed to identify congruent branches and to apply the GCPSR principle. Although all individual ML and BI gene trees displayed topological similarity and formed well-delimited clades (Fig. S3a–c), conflicts were observed among the phylogenies of different loci. Some variation in tree topologies was observed across individual gene genealogies, with a few branches showing incongruence and several internal nodes receiving low or no bootstrap support values (Fig. S3a–c). In the tef1 gene phylogeny, subclades A and B clustered together, and the longer branch lengths observed suggested potential genetic divergence within the AB, C, and D subclades (Fig. S3b). In contrast, the ITS (Fig. S3a) and the tub2 (Fig. S3c) gene phylogenetic analyses failed to resolve A–D as four independent and well-supported monophyletic lineages.
Although the tub2 gene displayed limited effectiveness in resolving subclades within the Pe. adusta clade, it was observed that all taxa in subclade A consistently clustered near the top of the tub2 phylogram (BS/PP = -/0.96, Fig. S3c), whereas taxa from subclades B, C, and D were interspersed. In contrast, multilocus phylogenetic analysis offered a more robust framework for delineating the Pe. adusta clade than single-gene trees (Figs. 14, S3a–c). Based on the combined dataset (Fig. 14) and single-gene phylogenies (Fig. S3a–c) analyzed under the GCPSR principle, the Pe. adusta clade was identified as a distinct and independent lineage. The mPTP analysis yielded consistent results with GCPSR, identifying the Pe. adusta clade as one MOTU. In contrast, PTP over-split the group, delimiting 28 MOTUs, even partitioning nearly identical strains into separate units—for example, the two isolates of Pe. dracaenicola (MFLUCC 18-0913 and MFLUCC 18-0914) and five isolates of Pe. sichuanensis (CGMCC 3.18244, GUCC 23-0340, GUCC 23-0363, GUCC 23-0364, and SC5A8). The ABGD and ASAP analyses produced congruent results, recognizing 14 MOTUs within the Pe. adusta clade (Fig. 14).
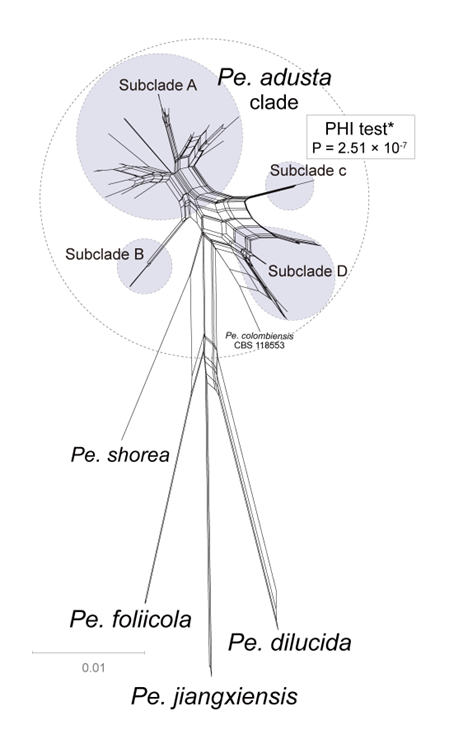
In contrast, phylogenetic network analysis revealed distinct differences between the outgroup and the Pe. adusta clade (Fig. 15). The parallel edges and box-like polygons connecting nearly all taxa within the Pe. adusta clade suggest a high likelihood of recombination. This inference is consistent with the results of the PHI test, which detected statistically significant evidence of recombination (P = 2.51 × 10⁻7; Fig. 15). Subclade B showed variability in its position within both the phylogenetic network and the three-locus phylogenetic tree. While various lines of evidence indicate that the 21 species in the Pe. adusta clade might be merged into one species, the considerable genetic distance noted between subclade A and both subclades C and D in the three-locus phylogenetic tree persists unexplained. This study therefore reinforces the acknowledgment of the Pe. adusta clade as a species complex instead of a singular species. Determining whether the Pe. adusta clade should be classified as a single species will necessitate further, in-depth investigations, utilizing more data and sophisticated analytical methods to explore the identified genetic variations divergence.
Species residing in the Pestalotiopsis adusta species complex
Pestalotiopsis adusta (Ellis & Everh.) Steyaert, Transactions of the British Mycological Society 36 (2): 82, 236 (1953)
Pestalotiopsis aggestorum F. Liu & L. Cai, Scientific Reports 7 (no. 866): 4 (2017)
Pestalotiopsis chiaroscuro Rapley, Steinrucken, Vitelli, Holdom & Y.P. Tan, Persoonia 48: 355 (2022)
Pestalotiopsis colombiensis Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 158 (2014)
Pestalotiopsis diploclisiae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 160 (2014)
Pestalotiopsis dracaenae Yong Wang bis, Y. Song, K. Geng & K.D. Hyde, Fungal Diversity 75: 164 (2015)
Pestalotiopsis dracaenicola Chaiwan & K.D. Hyde, Mycology 11 (4): 311 (2020)
Pestalotiopsis humicola Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 165 (2014)
Pestalotiopsis jinchanghensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 8 (2017)
Pestalotiopsis krabiensis Tibpromma & K.D. Hyde, Fungal Diversity 93: 143 (2018)
Pestalotiopsis licualacola K. Geng, Y. Song, K.D. Hyde & Yong Wang bis, Phytotaxa 88 (3): 51 (2013)
Pestalotiopsis malayana Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 169 (2014)
Pestalotiopsis neolitseae Ariyaw. & K.D. Hyde, Mycosphere 9 (5): 1005 (2018)
Pestalotiopsis pandanicola Tibpromma & K.D. Hyde, Fungal Diversity 93: 145 (2018)
Pestalotiopsis sichuanensis Y.C. Wang, X.C. Wang & Y.J. Yang, Plant Disease 103 (10): 2554 (2019)
Pestalotiopsis silvicola Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 22 (2022)
Pestalotiopsis spatholobi Z.X. Zhang, J.W. Xia and X.G. Zhang, Microorganisms 11 (7, no. 1627): 9 (2023)
Pestalotiopsis wulichongensis P. Razaghi, F. Liu, M. Raza & L. Cai, Studies in Mycology 109: 254 (2024)
Pestalotiopsis zhaoqingensis H.J. Zhao & W. Dong, Mycosphere 14 (1): 2238 (2023)
Notes: Phylogenetic analyses based on a three-gene dataset indicate that, Pe. adusta forms a well-supported clade (BS/PP = 77%/0.99) with 20 other species, including Pe. aggestorum, Pe. chiaroscuro, Pe. colombiensis, Pe. diploclisiae, Pe. dracaenae, Pe. dracaenicola, Pe. humicola, Pe. jinchanghensis, Pe. krabiensis, Pe. licualacola, Pe. malayana, Pe. neolitseae, Pe. pandanicola, Pe. papuana, Pe. pyrrosiae-linguae, Pe. sichuanensis, Pe. silvicola, Pe. spatholobi, Pe. wulichongensis, and Pe. zhaoqingensis (Fig. 9). The phylogenetic network analysis (Fig. 15) revealed numerous reticulations and parallel edges within this lineage, indicating extensive phylogenetic conflicts and poorly defined subclade boundaries. Both GCPSR and mPTP analyses yielded consistent results, supporting the treatment of this lineage as a single MOTU (Fig. 14). Genomic evidence also demonstrated a high degree of genetic coherence among the six available genomes, with core genomes accounting for 90.88% of the total pangenome and ANI values ranging from 96.29% to 99.31%. However, because this lineage encompasses numerous taxa and the genomic data of all type strains are not yet available, the current evidence is insufficient to justify formal species merger. Therefore, based on the available phylogenetic and genomic data, this lineage is herein regarded as the Pestalotiopsis adusta species complex.
Synonymies in the Pestalotiopsis adusta species complex
Pestalotiopsis adusta (Ellis & Everh.) Steyaert, Transactions of the British Mycological Society 36 (2): 82, 236 (1953)
= Pestalotiopsis papuana Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 174 (2014)
See Maharachchikumbura et al. (2012) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: USA, Newfield, New Jersey, on leaves of Prunus cerasus L., cultivated plum, 20 July 1887 (NY 00937391, holotype); FIJI, on refrigerator door PVC gasket, 1 June 1978, E.H.C. McKenzie (MFLU 12-0425, epitype; ex-epitype living culture ICMP 6088 = PDDCC 6088).
Host range: Celtis formosana (Tennakoon et al. 2021), Clerodendrum canescens (Xu et al. 2016), Cocos nucifera (Maharachchikumbura et al. 2014b; Rosado et al. 2015; Tian et al. 2024), Dictyosperma album (Zhu et al. 2015), Ericaceae (Kohout and Tedersoo 2017), Hevea brasiliensis (de Oliveira Amaral et al. 2022), Immature coconut (Rosado et al. 2015), Mangifera indica (Shu et al. 2020; Adikaram et al. 2023), Nectandra lineatifolia (Nelson et al. 2020), Oryza sp. (Pak et al. 2017), Palm (Zhang et al. 2024c), Podocarpus macrophyllus (Wei et al. 2007), Prunus cerasus (Maharachchikumbura et al. 2012), Refrigerator door PVC gasket (Maharachchikumbura et al. 2012), Rhizophora mucronata (Apurillo et al. 2019), Rubus idaeus (Yan et al. 2019), Sinopodophyllum hexandrum (Xiao et al. 2017), Smilax nipponica (Watanabe et al. 2010), Soil along the coast (Maharachchikumbura et al. 2014b), Syagrus oleracea (Cardoso et al. 2017), Syzygium sp. (Maharachchikumbura et al. 2012), Unknown grass species (Pak et al. 2017), Vaccinium corymbosum (Zheng et al. 2023b).
Known distribution: America (Maharachchikumbura et al. 2012; Pak et al. 2017), Brazil (Rosado et al. 2015; Cardoso et al. 2017; de Oliveira Amaral et al. 2022), China (Zhu et al. 2015; Xu et al. 2016; Shu et al. 2020; Tennakoon et al. 2021), Ecuador (Nelson et al. 2020), Fiji (Maharachchikumbura et al. 2012), Japan (Watanabe et al. 2010), Papua New Guinea (Maharachchikumbura et al. 2014b), Philippines (Apurillo et al. 2019), South Africa (Kohout and Tedersoo 2017), Sri Lanka (Adikaram et al. 2023), Thailand (Maharachchikumbura et al. 2012; Tian et al. 2024; Zhang et al. 2024c).
Notes: Pestalotiopsis adusta was originally described by Steyaert (1949) from cultivated plum in America. This species has been reported from a wide range of hosts and exhibits a cosmopolitan distribution. In this study, we only synonymize species that exhibit minimal genetic divergence in ITS, tef1, and tub2 and form a well-supported monophyletic lineage. Specifically, Pe. adusta and Pe. papuana cluster together with strong statistical support (BS/PP = 88%/-, Fig. 9) and exhibit a short branch length. Sequence comparisons between the type specimens of Pe. adusta and Pe. papuana reveal 100% identity for ITS (539/539 bp), 99.79% identity for tef1 (473/474 bp, no gaps), and 99.55% identity for tub2 (443/445 bp, no gaps). Morphologically, the primary distinctions between Pe. adusta and Pe. papuana in conidial length and the number and length of apical appendages. Pestalotiopsis adusta produces shorter conidia (16–20 μm) than Pe. papuana (17–24 μm), and its apical appendages (2–3, 7–15 μm) are more numerous and longer than those of Pe. papuana (1–2, 1.5–7 μm) (Maharachchikumbura et al. 2012; 2014b). However, such minor morphological differences present limitations for species delimitation in pestalotiopsis-like taxa (see discussion). Given the strong molecular evidence, we formally synonymize Pe. papuana under Pe. adusta.
Pestalotiopsis spatholobi Z.X. Zhang, J.W. Xia and X.G. Zhang, Microorganisms 11 (7, no. 1627): 9 (2023)
= Pestalotiopsis pyrrosiae-linguae H. Li, Mycosphere 14 (1): 2238 (2023)
See Zhang et al. (2023) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China. Hainan Province, East Harbour National Nature Reserve, on diseased leaves of Spatholobus suberectus, 23 May 2021, Z.X. Zhang, (HMAS 352479, holotype); ex-type SAUCC231201.
Host range: Pyrrosia lingua (Dong et al. 2023), Spatholobus suberectus (Zhang et al. 2023).
Known distribution: China (Dong et al. 2023; Zhang et al. 2023).
Notes: The three-gene phylogenetic tree (Fig. 9) revealed that the two isolates (GUCC 23-0371 and GUCC 23-0372) clustered together with Pe. pyrrosiae-linguae and Pe. spatholobi with strong statistical support (BS = 98%, PP = 1). Species delimitation analyses using PTP, ABGD, and ASAP consistently identified these taxa as a single MOTU (Fig. 14). The ex-type cultures of Pe. pyrrosiae-linguae and Pe. spatholobi showed high sequence similarity: 99.66% for ITS (595/597 bp; no gaps), 100% for tef1 (231/231 bp), and 100% for tub2 (757/757 bp). Morphologically, Pe. pyrrosiae-linguae and Pe. spatholobi exhibit highly similar conidial characteristics (Dong et al. 2023; Zhang et al. 2023). Based on congruent molecular and morphological evidence, Pe. pyrrosiae-linguae is herein regarded as a synonym of Pe. spatholobi.
The Pestalotiopsis brassicae clade
The three-locus gene tree (Fig. 9) and the individual gene trees (Fig. S2a–c) of Pestalotiopsis demonstrated that the Pe. brassicae clade (Pe. brassicae, Pe. chinensis, Pe. hollandica, Pe. italiana, Pe. sequoiae, Pe. verruculosa) and the Pe. endophytica clade (including the type strain of Pe. endophytica and two isolates recovered in this study) each formed monophyletic groups. Moreover, the placements of these two clades were closely linked in both the three-locus gene tree (Fig. 9) and the separate gene trees (Fig. S2a–c). To facilitate further analysis, we constructed three-locus and single-locus phylogenetic trees, which included the Pe. brassicae clade, the Pe. endophytica clade, and their closely related species, Pe. unicolor and Pe. iberica. The concatenated alignment comprised 1,791 characters derived from three loci: 541 characters from ITS, 496 from tef1, and 754 from tub2, including alignment gaps. A summary of the detailed alignment characteristics used in the ML and BI analyses of the Pe. brassicae clade is provided in Table S7.
The multilocus ML and BI phylogenetic trees for the Pe. brassicae clade and the Pe. endophytica clade displayed similar topologies, with both forming well-supported branches. These findings strongly reinforce the monophyly of both the Pe. brassicae clade and the Pe. endophytica clade (Fig. S4a).
Gene trees from individual ML and BI analyses were compared to identify common branches while applying the GCPSR principle. All individual ML and BI gene trees exhibited similar topologies, showing the same clearly defined clades. Furthermore, there were almost no discrepancies among the individual phylogenies of different loci, aligning well with the analysis of the concatenated dataset (Fig. S4a-c). The only exception was the ITS-phylogram, in which Pe. yunnanensis HMAS 96359 formed a long branch (Fig. S4a). Given that Pe. yunnanensis HMAS 96359 lacked tef1 and tub2 gene sequences, it was excluded from subsequent analyses to ensure robust interpretations. Similarly, while Pe. sequoiae MFLUCC 13-0399 and Pe. chinensis MFLUCC 12-0273 formed a highly supported monophyletic group (BS/PP = 90%/0.95) with other members of the Pe. brassicae clade in the ITS-phylogram (Fig. S4a), the absence of tef1 and tub2 gene data also led to their exclusion. Excluding these strains, all remaining Pestalotiopsis species displayed consistent resolution across individual phylogenetic trees (Fig. S4a-c). Every dataset showed similar resolution, revealing the same well-defined groupings found in the combined dataset phylogenies. Notably, the Pe. brassicae clade and Pe. endophytica clade were consistently recovered with high support values (BS ≥ 85%; PP ≥ 0.95) across nearly all individual gene trees (Fig. S4a-c). According to the GCPSR criterion, this lineage was recognized as an independent evolutionary lineage. Species delimitation analyses using PTP, ABGD, and ASAP produced results consistent with GCPSR, each identifying the Pe. brassicae clade as a single MOTU (Fig. 16). However, the mPTP analysis divided the Pe. brassicae clade into three MOTUs, assigning Pe. monochaeta to one MOTU, GCA 018115615.1 to another, and the remaining strains to a third (Fig. 16). The Pe. endophytica clade comprised the ex-type strain of Pe. endophytica together with two isolates obtained in this study. All five species delimitation methods (GCPSR, PTP, mPTP, ABGD, and ASAP) consistently supported the recognition of this clade as a single MOTU (Fig. 16).
This phylogenetic network structure was consistent with the results of the PHI test, which found no statistically significant evidence of recombination within Pe. brassicae, the Pe. endophytica, Pe. iberica and Pe. jiangxiensis (P = 0.92; Fig. 17b). For the Pe. brassicae clade, the PHI test also indicated no recombination (P = 0.32; Fig. 17a). Nevertheless, this calculation relied on just ten informative sites, suggesting the result may be a false negative. The small number of informative sites also indicates the low genetic divergence among strains of Pe. brassicae clade.
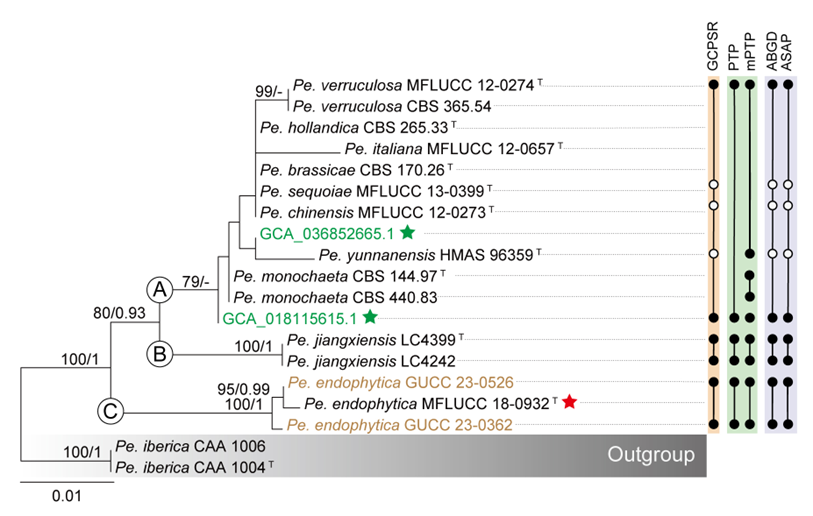
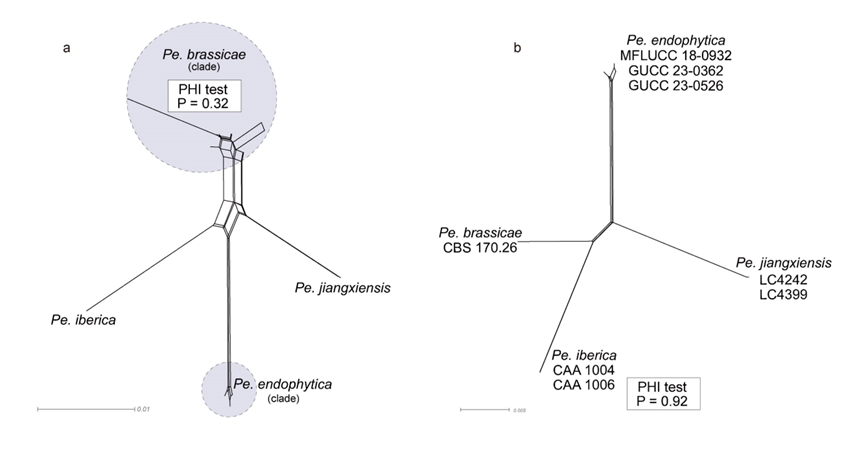
Species residing in the Pestalotiopsis brassicae species complex
Pestalotiopsis brassicae (Guba) Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 157 (2014)
Pestalotiopsis chinensis Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 107 (2012)
Pestalotiopsis italiana Maharachch., Camporesi & K.D. Hyde, Fungal Diversity 72: 16 (2015)
Pestalotiopsis monochaeta Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 170 (2014)
Pestalotiopsis sequoiae W.J. Li, Camporesi & K.D. Hyde, Fungal Diversity 80: 210 (2016)
Pestalotiopsis verruculosa Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 123 (2012)
Notes: Phylogenetic analyses based on a three-gene dataset revealed that, Pe. brassicae forms a distinct clade together with seven other species—Pe. chinensis, Pe. hollandica, Pe. italiana, Pe. monochaeta, Pe. sequoiae, Pe. verruculosa, and Pe. yunnanensis—indicating a close evolutionary relationship among them (Fig. 9). This clustering is further supported by phylogenetic network analyses (Fig. 8). Species delimitation analyses using both the GCPSR and PTP methods consistently support the recognition of these eight species as independent evolutionary lineages (Fig. 16). However, Pe. yunnanensis was excluded from the analysis because only ITS sequences are currently available, and its ITS sequence differs significantly from those of other taxa within the Pe. brassicae clade. In ITS-based phylogenetic trees (Fig. S2a), Pe. yunnanensis does not cluster within the Pe. brassicae clade, further supporting its exclusion. In contrast, the remaining seven species consistently group together in single-gene phylogenies based on ITS, tef1, and tub2 (Fig. S2a–c). However, we currently lack sufficient evidence to confirm that, Pe. brassicae, Pe. chinensis, Pe. hollandica, Pe. italiana, Pe. monochaeta, Pe. sequoiae, and Pe. verruculosa represent a single species. Several factors contribute to this uncertainty. Pestalotiopsis monochaeta exhibits a unique 9-bp insertion in the tef1 gene and highly distinctive morphological characteristics compared to other members of the genus. Additionally, no whole-genome data are currently available for Pe. monochaeta. Furthermore, Pe. chinensis and Pe. sequoiae are only represented by ITS sequences, lacking tef1 and tub2 data, which limits the resolution of their phylogenetic placement. Due to these limitations, we propose that, Pe. brassicae, Pe. chinensis, Pe. hollandica, Pe. italiana, Pe. monochaeta, Pe. sequoiae, and Pe. verruculosa should not be treated as a single species but rather as the Pestalotiopsis brassicae species complex, pending further morphological and molecular studies.
Synonymies in the Pestalotiopsis brassicae species complex
Pestalotiopsis brassicae (Guba) Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 157 (2014)
= Pestalotiopsis hollandica Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 164 (2014)
See Maharachchikumbura et al. (2014b) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: New Zealand, from seeds of Brassica napus, May 1926, G.H. Cunningham (CBS H-7542, isotype); ex-isotype CBS 170.26.
Host range: Brassica napus (Maharachchikumbura et al. 2014b), Cupressus sempervirens (Crous et al. 2018), Pinus pinea (Silva et al. 2020), Sciadopitys verticillate (Maharachchikumbura et al. 2014b).
Known distribution: Netherlands (Maharachchikumbura et al. 2014b), New Zealand (Maharachchikumbura et al. 2014b), Portugal (Silva et al. 2020), Spain (Crous et al. 2018).
Notes: Pestalotiopsis brassicae was originally described by Guba (1961) as Pestalotia brassicae from Brassica napus seeds in New Zealand and was subsequently reassigned to Pestalotiopsis by Maharachchikumbura et al. (2014b). Sequence similarity analysis between the type strains of Pe. brassicae and Pe. hollandica reveals high genetic similarity, with both ITS and tef1 sequences showing 100% identity (540/540 bp and 447/447 bp, respectively). The primary morphological differences between Pe. brassicae and Pe. hollandica in conidial length and the length of the basal appendages. Pe. brassicae has longer conidia (29–40 μm vs. 25–34 μm) and basal appendages (10–25 μm vs. 3–9 μm) than Pe. hollandica (Maharachchikumbura et al. 2014b; Figs. 12–13). However, we emphasize the priority of molecular evidence in species delimitation (see discussion). Given the molecular data, we formally propose synonymizing Pe. hollandica under Pestalotiopsis brassicae.
The Pestalotiopsis camelliae clade
The three-locus gene tree (Fig. 9) and individual gene trees (Fig. S2a–c) of Pestalotiopsis demonstrated that the Pe. camelliae clade, consisting of seven strains (including three Pe. camelliae strains, two Pe. yanglingensis strains, and two isolates collected in this study), constituted a distinct and well-supported monophyletic lineage. In a similar vein, the Pe. portugallica clade, consisting of six strains (two previously described Pe. portugallica strains and four additional isolates collected during this study), also demonstrated a strongly supported monophyletic structure. Given the small number of strains in these two clades and their close evolutionary relationship, they were analyzed together. We constructed three-locus and single-locus phylogenetic trees that included the Pe. camelliae clade, the Pe. portugallica clade, and their closely related species: Pe. changjiangensis, Pe. furcata, Pe. novae-hollandiae, and Pe. gibbosa (Figs. 18, S5a–c). The concatenated alignment consisted of 1,823 characters derived from three loci: 547 characters from ITS, 521 from tef1, and 755 from tub2, including alignment gaps. A detailed summary of the alignment characteristics used in ML and BI analyses of the Pe. camelliae and Pe. portugallica clades is provided in Table S8.
The ML and BI phylogenetic analyses of the three-locus gene tree for this clade displayed highly similar topologies, delineating five subclades (Fig. 18). Subclade A corresponded to the Pe. camelliae clade, encompassing Pe. camelliae and Pe. yanglingensis. Subclades B, C, and D were identified as Pe. furcata, Pe. novae-hollandiae, and Pe. changjiangensis, respectively. Subclade E represented the Pe. portugallica clade, comprising Pe. portugallica.
The individual ML and BI gene trees were compared to identify shared branches and evaluate species boundaries based on the GCPSR principle. In general, the topologies of all individual ML and BI gene trees were very similar, consistently showcasing clearly defined clades. Nevertheless, certain discrepancies were noted among the individual phylogenies of the different loci. In the ITS- (Fig. S5a) and tub2-phylograms (Fig. S5c), Pe. yanglingensis clustered with Pe. camelliae with strong support (BS ≥ 88, PP ≥ 0.99). In contrast, the position of Pe. yanglingensis in the tef1-phylogram (Fig. S5b) was unresolved and lacked robust support. In the Pe. portugallica clade, all strains consistently grouped into one monophyletic branch in every individual gene tree, revealing strongly congruent evolutionary relationships (Fig. S5a–c). Based on the phylogenetic analyses of the combined dataset and the application of the GCPSR principle, Pe. yanglingensis is proposed as a potential synonym of Pe. camelliae. Additionally, all strains within the Pe. portugallica clade were identified as belonging to a single species. The results of ABGD and ASAP were consistent with the GCPSR principle, recognizing Pe. yanglingensis and Pe. camelliae as a single MOTU. In contrast, PTP and mPTP analyses separated Pe. yanglingensis and Pe. camelliae into distinct MOTUs (Fig. 18)
Based on the relative genetic distances and the topology of the phylogenetic network (Fig. 19), the Pe. camelliae clade and Pe. furcata displayed distinct bifurcated, tree-like evolutionary patterns. This observation is consistent with prior analyses that identified these groups as two separate species (Fig. 19). Additionally, the phylogenetic network for Pe. camelliae, Pe. yanglingensis, and Pe. furcata showed no evidence of recombination (P = 0.22), further supporting their classification as distinct species. For the Pe. camelliae clade, the PHI test similarly revealed no evidence of recombination (P = 0.16). However, this result was based on only 13 informative sites, raising concerns about its reliability due to insufficient sequence variation. The limited number of informative sites also indicates high sequence similarity within the Pe. camelliae clade. Conversely, networks within the Pe. portugallica clade tested positive for recombination (P = 0.03), providing evidence that these strains should be regarded as a single species.
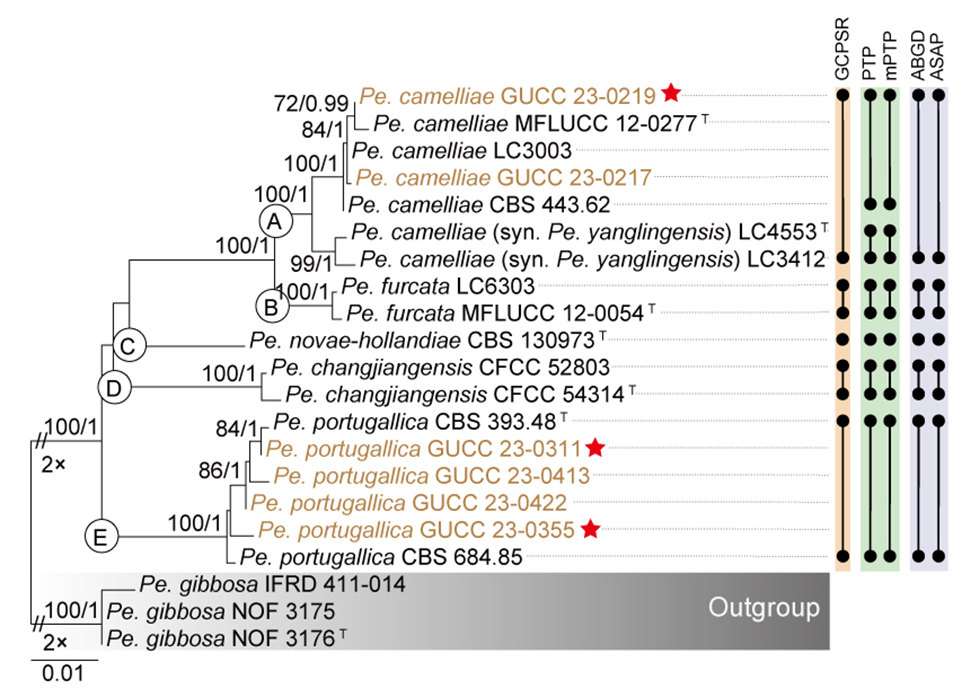

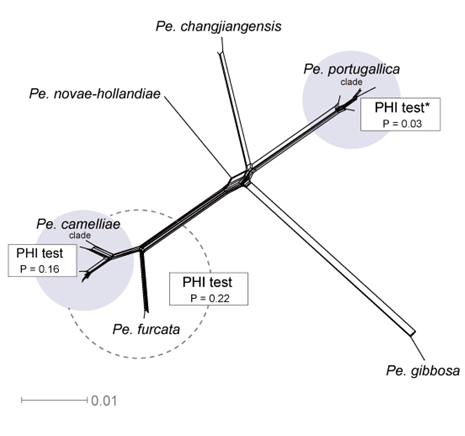
Synonymies in the Pestalotiopsis camelliae clade
Pestalotiopsis camelliae Y.M. Zhang, Maharachch. & K.D. Hyde, Sydowia 64 (2): 337 (2012)
= Pestalotiopsis yanglingensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 11 (2017)
See Zhang et al. (2012b) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Yunnan Province, Chuxiong, Shuangbai, on living leaves of Camellia japonica, July 2011, Y.M. Zhang (IFRD OP111, holotype); ex-type MFLUCC 12-0277.
Host range: Camellia japonica (Zhang et al. 2012b), Camellia sinensis (Liu et al. 2017; Chen et al. 2018c; Win et al. 2018; Tsai et al. 2021; Win et al. 2021).
Known distribution: China (Zhang et al. 2012b; Liu et al. 2017; Chen et al. 2018c; Tsai et al. 2021), Japan (Win et al. 2018; Win et al. 2021), Turkey (Vu et al. 2019).
Notes: Pestalotiopsis camelliae and Pe. yanglingensis, both originally described from Camellia in China (Zhang et al. 2012b; Liu et al. 2017), form a well-supported cluster in both the three-gene phylogenetic tree and single-gene phylogenies (Fig. 9 and Fig. S2). Species delimitation using the GCPSR approach categorizes them as a single species. Furthermore, molecular comparisons reveal high sequence similarity between type strain of Pe. yanglingensis and Pe. camelliae, with ITS, tef1, and tub2 showing identity levels of 99.80% (503/504, including one gap), 98.62% (858/870, including four gaps), and 99.32% (439/442, no gaps), respectively. Morphological, the only notable distinction between Pe. yanglingensis and Pe. camelliae is that Pe. yanglingensis exhibits a higher proportion of branched apical appendages (Zhang et al. 2012b; Liu et al. 2017; Fig. 12). However, we consider this variation insufficient to warrant their recognition as distinct species (see discussion). Based on the strong congruence between phylogenetic, morphological, and molecular data, we synonymize Pe. yanglingensis with Pe. camelliae.
The Pestalotiopsis clavata clade
The combined gene tree (Fig. 9) and individual gene trees of Pestalotiopsis (Fig. S2) revealed that 23 isolates within the Pe. clavata clade represents nine species, collectively designated as the Pe. clavata clade. These isolates form a well-supported monophyletic group (BS/PP = -/0.98, Fig. 9). Based on this information, an additional phylogenetic tree was constructed, including 23 ingroup taxa and two outgroup taxa (Pe. biappendiculata CGMCC 3.23487 and LC4354) (Fig. 20). The combined alignment for the three loci comprised 1,792 characters, including 561 from ITS, 477 from tef1, and 754 from tub2 (including alignment gaps). Detailed alignment characteristics for ML and BI analyses of the Pe. clavata clade are provided in Table S9.
Since all ML and BI analyses yielded topologically similar trees, only the ML tree is presented, with bootstrap support values and posterior probabilities indicated for well-supported clades (Fig. 20). The phylogenetic analyses resolved the ingroup taxa into five subclades, which collectively formed a well-supported monophyletic lineage with high support values (100% BS, PP = 1; Fig. 20). Among these, subclades B (Pe. multiappendiculata), C (Pe. cangshanensis), D (Pe. phoebes), and E (Pe. fusoidea), each represent a distinct and well-supported lineage, whereas subclade A (Pe. clavata, Pe. iberica, Pe. lushanensis, Pe. pini, and Pe. rhododendri) exhibited significant internal substructure (Fig. 20).
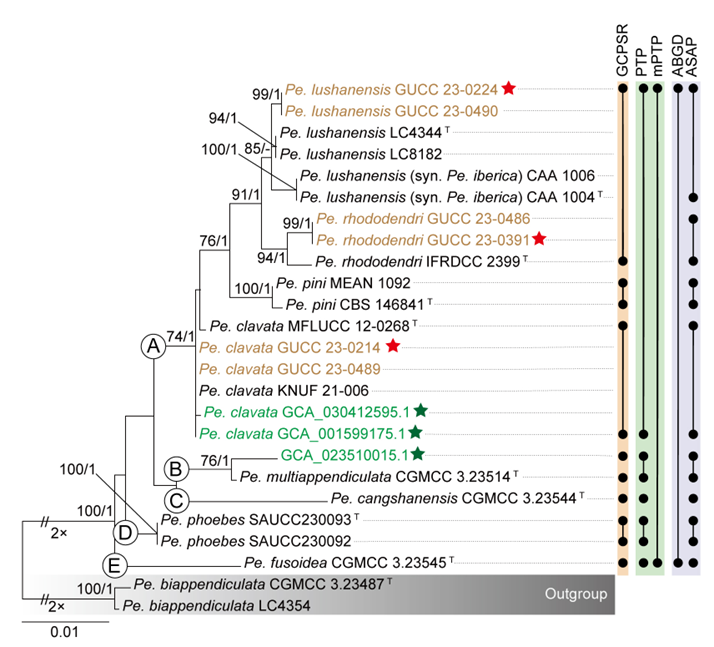
The GCPSR principle was applied, and individual ML and BI gene trees were compared to identify concordant branches. All individual ML and BI gene trees demonstrated topological similarity, delineating the same well-supported clades. However, some discrepancies were noted among the phylogenies of different loci (Fig. S6a–c). Specifically, the ITS-phylogram (Fig. S6a) presented short branches, causing all taxa to cluster closely together, making them challenging to distinguish. By contrast, tef1 and tub2 phylograms generated highly discordant branches (Fig. S6b–c), leading to unclear boundaries. Despite these inconsistencies, the tub2 phylogram displayed a topology very similar to that observed in the combined dataset analysis (Figs. 20, S6c), with subclades A–E each forming a well-defined lineage. The strains in subclade A exhibited minimal divergence in tub2-_phylogram. Conversely, the _tef1 phylogram showed greater variation in elucidating evolutionary relationships within subclade A (Fig. S6b). Specifically, the subclade A was subdivided into three subclades: A1, comprising ten tef1 sequences from Pe. iberica, Pe. lushanensis, and Pe. rhododendri; A2, containing sequences from Pe. pini; and A3, consisting of six sequences of Pe. clavata. Interestingly, A3 clustered with Pe. cangshanensis CGMCC 3.23544 and Pe. multiappendiculata CGMCC 3.23514. The GCPSR analysis recognized the Pe. clavata clade as comprising eight MOTUs. The PTP analysis divided this clade into five MOTUs, corresponding to subclades A–E (Fig. 20). In contrast, both mPTP and ABGD identified the Pe. clavata clade as a single MOTU, whereas ASAP divided it into eight MOTUs (Fig. 20).
Significant genetic divergence was observed in the tub2 gene of Pe. cangshanensis and the tef1 gene of Pe. fusoidea relative to other taxa in the clade. Furthermore, the phylogenetic network (Fig. 21) analysis showed that the long branch lengths formed by Pe. fusoidea, Pe. phoebes, Pe. multiappendiculata, Pe. cangshanensis, and Pe. biappendiculata did not support the inclusion of ABGD and mPTP. Within subclade A, PHI tests among strains of Pe. clavata, Pe. iberica, Pe. lushanensis, Pe. pini, and Pe. rhododendri detected no statistically significant recombination (P = 0.22).
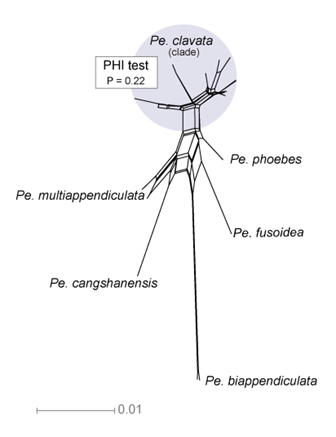
Species residing in the Pestalotiopsis clavata species complex
Pestalotiopsis cangshanensis H.W. Shen, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 7 (2022)
Pestalotiopsis clavata Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 108 (2012)
Pestalotiopsis fusoidea D.F. Bao, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 11 (2022)
Pestalotiopsis lushanensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 9 (2017)
Pestalotiopsis multiappendiculata P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 249 (2024)
Pestalotiopsis phoebes Z.X. Zhang, J.W. Xia & X.G. Zhang, Microorganisms. 11 (1627): 7 (2023)
Pestalotiopsis pini A.C. Silva, E. Diogo & H. Bragança, Forests 11 (8, no.805): 9 (2020)
Pestalotiopsis rhododendri Y.M. Zhang, Maharachch. & K.D. Hyde, Sydowia 65 (1): 123 (2013)
Notes: Pestalotiopsis clavata and eight other species—Pe. cangshanensis, Pe. fusoidea, Pe. iberica, Pe. lushanensis, Pe. multiappendiculata, Pe. phoebe_s, _Pe. pini, and Pe. rhododendri—cluster together in the three-gene phylogenetic tree with support values (BS/PP = -/0.98, Fig. 9). Species delimitation analyses using mPTP and ABGD consistently recognize them as a MOTU. Furthermore, genomic data reveal that the six available genomes within this clade share ANI values exceeding 97.73% and have a core gene content representing 90.29% of their respective pan-genomes, indicating a high degree of genetic similarity among these taxa. However, we observed that Pe. cangshanensis and Pe. fusoidea are associated with long branches in the phylogenetic tree, and the tub2 gene sequence of Pe. cangshanensis exhibits notable divergence from the other members of this clade. Moreover, genome sequences are currently unavailable for both Pe. cangshanensis and Pe. fusoidea, limiting the resolution of their phylogenetic placement. Due to these limitations, we propose that Pe. clavata, Pe. fusoidea, Pe. multiappendiculata, and Pe. phoebes should be treated as the Pestalotiopsis clavata species complex, pending further genomic and morphological analyses to clarify the taxonomic status of its constituent species.
Synonymies in the Pestalotiopsis clavata species complex
Pestalotiopsis lushanensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 9 (2017)
= Pestalotiopsis iberica P. Monteiro & M. Gonçalves, European Journal of Plant Pathology 162: 194 (2022)
See Liu et al. (2017) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Jiangxi Province, Lushan National Park, on Camellia sp., 5 September 2013, Y.H. Gao, (HMAS 247059, holotype); ex-type CGMCC 3.18160 = LC4344).
Host range: Camellia sinensis (Chen et al. 2018b; Manawasinghe et al. 2021), Camellia sp. (Liu et al. 2017), Dendrobium sp. (Ma et al. 2019), Pinus radiata (Monteiro et al. 2022), Pinus sylvestris (Monteiro et al. 2022), Podocarpus macrophyllus (Zheng et al. 2022), Quercus serrata (Jiang et al. 2022), Sarcandra glabra (Zhang et al. 2021a).
Known distribution: China (Chen et al. 2018b; Manawasinghe et al. 2021; Zhang et al. 2021a; Jiang et al. 2022; Zheng et al. 2022), Spain (Monteiro et al. 2022), Thailand (Ma et al. 2019).
Notes: The three-locus phylogenetic analysis (Fig. 9) showed that two isolates (GUCC 23-0224 and GUCC 23-0490) clustered with Pe. iberica and Pe. lushanensis with strong support (BS/PP = 88%/0.98, Fig. 9). The ASAP analysis recognized them as a single MOTU. The ex-type strains of Pe. iberica and Pe. lushanensis exhibited 96.83% similarity in ITS sequences (489/505, including 15 gaps), 99.63% similarity in tef1 (542/544, no gaps), and 99.22% similarity in tub2 (760/766, including one gap). Morphologically, Pe. iberica and Pe. lushanensis share similar conidial characteristics (Liu et al. 2017; Monteiro et al. 2022). Based on the combined molecular and morphological evidence, Pe. iberica is proposed as a synonym of Pe. lushanensis.
The Pestalotiopsis hainanensis clade
The phylogenetic tree for the entire genus revealed that the Pe. hainanensis clade includes 15 species (Fig. 9): Pe. appendiculata, Pe. australis, Pe. chamaeropis, Pe. daliensis, Pe. hainanensis, Pe. intermedia, Pe. jiangxiensis, Pe. leucadendri, Pe. linearis, Pe. matildae, Pe. rosarioides, Pe. scoparia, Pe. taxicola, Pe. tumida, and Pe. unicolor. The phylogenetic network and single-gene trees for the genus were largely consistent with the overall phylogeny, supporting the formation of a distinct Pe. hainanensis clade (Fig. 8 and Fig. S2). However, exceptions were noted: in the phylogenetic network (Fig. 8), Pe. leucadendri was positioned distantly from the Pe. hainanensis clade, which retained the remaining 14 species. This discrepancy is likely attributable to divergence in the tub2 gene, as phylogenetic analysis of the tub2 gene shows Pe. leucadendri grouping outside the Pe. hainanensis clade (Fig. S2c).
To facilitate further analysis, we constructed three-locus combined phylogenetic tree and single-gene trees for the Pe. hainanensis clade and its neighboring species, Pe. rubrae and Pe. endophytica. The combined alignment for the three loci included 1,774 characters: 532 from ITS, 476 from tef1, and 766 from tub2 (including alignment gaps). Table S10 summarizes the alignment statistics and the parameters used for ML and BI analyses. Since all ML and BI analyses produced topologically congruent trees, only the ML tree is shown, annotated with bootstrap values and posterior probabilities to highlight well-supported clades. The phylogenetic analyses resolved the ingroup taxa into five subclades, forming a well-supported monophyletic lineage with high support values (74% BS/ 0.99 PP) (Fig. 22). These subclades, designated as subclade A (Pe. appendiculata, Pe. intermedia, Pe. linearis, Pe. rosarioides, Pe. tumida, Pe. chamaeropis, Pe. daliensis, and Pe. hainanensis), subclade B (Pe. jiangxiensis, Pe. taxicola, and Pe. unicolor), subclade C (Pe. australis and Pe. matildae), subclade D (Pe. leucadendri), and subclade E (Pe. scoparia), represent distinct and strongly supported evolutionary lineages.
To assess the phylogenetic conflicts identified in the concatenated dataset, individual ML and BI gene trees were analyzed to compare concordant and discordant branches, with the aim of delineating independent evolutionary lineages within the Pe. hainanensis clade in accordance with the GCPSR principle. All individual ML and BI gene trees exhibited consistent topologies, defining well-supported and distinct clades. However, significant conflicts were observed among the phylogenies of various loci. In the ITS phylogram (Fig. S7a), ITS provided limited resolution for distinguishing taxa within the Pe. hainanensis clade. Most taxa appeared clustered together, except for strains in subclades C and D, which were positioned away from the remaining taxa in the clade. Conversely, the tef1 phylogram (Fig. S7b) showed better resolution, with subclades A, B, and E forming well-defined branches, while subclades C and D grouped together as a separate branch that was resolved as a sister group to subclade E. In contrast, the tub2 phylogram (Fig. S7c) depicted subclades A, B, and E clustering together with high support values (75% BS / 1 PP), while Subclades C and D formed independent branches. Subclade D was positioned far from the Pe. hainanensis clade, forming a long branch, which strongly suggests that the strains in subclade D represent a distinct species. Notably, none of the single-gene tree topologies fully aligned with the phylogenetic analyses derived from the concatenated dataset. The multilocus phylogenetic analyses provided greater resolution and delineation of the Pe. hainanensis clade compared to individual gene phylogram. The phylogenetic position of Pe. matildae remains uncertain, as this taxon lacks tef1 and tub2 gene sequences.
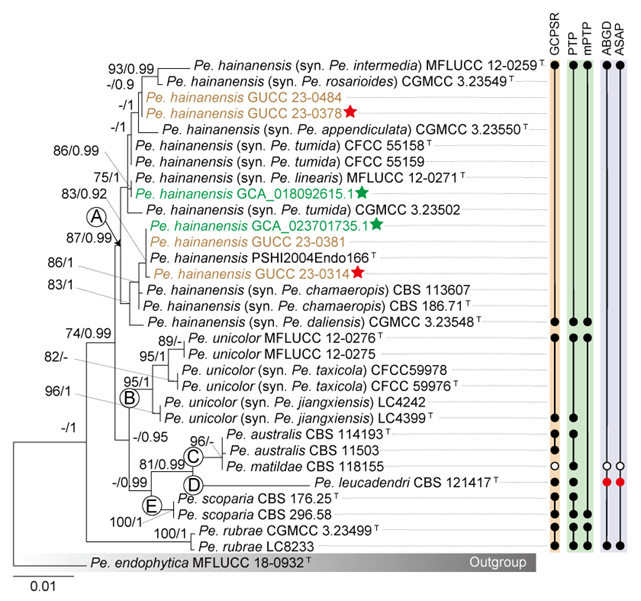
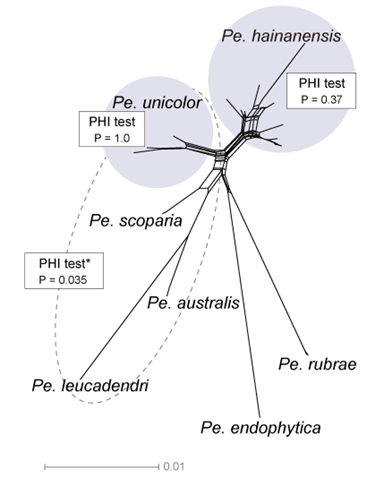
Based on the combined dataset analyses and the GCPSR principle, subclade A is confirmed as a MOTU. Within subclade A, Pe. appendiculata, Pe. chamaeropis, Pe. daliensis, Pe. intermedia, Pe. linearis, Pe. rosarioides, and Pe. tumida are proposed as potential synonyms of Pe. hainanensis. Similarly, within subclade B, Pe. jiangxiensis and Pe. taxicola are potentially synonymous with Pe. unicolor. The PTP results were largely consistent with the GCPSR principle (Fig. 22). However, Pe. matildae was excluded from the GCPSR analysis because tef1 and tub2 sequence data were unavailable. Both ABGD and ASAP yielded similar results, recognizing Pe. leucadendri as a single MOTU and grouping the remaining taxa into another MOTU. These results were inconsistent with the phylogenetic tree, likely due to the pronounced divergence of the tub2 sequence of Pe. leucadendri from other taxa within this clade.
The phylogenetic network clearly distinguished Pe. australis and Pe. leucadendri from other taxa within the Pe. hainanensis clade. Furthermore, Pe. hainanensis and Pe. unicolor, as delineated by GCPSR, each formed distinct clusters in the phylogenetic network. Although the PHI test for recombination was negative (P = 0.37; Fig. 23) for all datasets analyzed within subclade A (Pe. hainanensis), this result was based on only 14 informative sites, which suggests a false negative. The limited number of informative sites further indicates high sequence similarity within Pe. hainanensis. Similarly, for Pe. unicolor, the PHI test also returned a negative result (P = 1; Fig. 23). However, this finding is likely a false negative for the same reason, as it was based on only eight informative sites, highlighting the high sequence similarity within subclade B (Pe. unicolor).
Synonymies in the Pestalotiopsis hainanensis clade
Pestalotiopsis hainanensis A.R. Liu, T. Xu & L.D. Guo, Fungal Diversity 24: 29 (2007)
= Pestalotiopsis appendiculata D.F. Bao, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 7 (2022)
= Pestalotiopsis chamaeropis Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 158 (2014)
= Pestalotiopsis daliensis H.W. Shen, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 9 (2022)
= Pestalotiopsis intermedia Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 115 (2012)
= Pestalotiopsis linearis Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 117 (2012)
= Pestalotiopsis rosarioides H.W. Shen, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 14 (2022)
= Pestalotiopsis tumida C. Peng & C.M. Tian, Persoonia 49: 235 (2022)
See Liu et al. (2007) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Host range: Artemisia argyi (Qian et al. 2014), Camellia sp. (Liu et al. 2017), Castanopsis fissa (Jiang et al. 2022), Chamaerops humilis (Maharachchikumbura et al. 2014b), Decaying woody (Adams and Walker 2023), Erica arborea (Hlaiem et al. 2018), Eurya nitida (Qiu et al. 2022), Ophiocordyceps sp. (Hsu et al. 2024), Peristrophe japonica (Sun et al. 2023), Pieris japonica sub sp. japonica (Nozawa et al. 2019), Podocarpus macrophyllus (Liu et al. 2007), Prostanthera rotundifolia (Moslemi and Taylor 2015), Quercus acutissima, Q. aliena, Q. variabilis (Jiang et al. 2022), Rhododendron decorum (Gu et al. 2022), Rosa chinensis (Peng et al. 2022), Trachelospermum sp., Unidentified host (Maharachchikumbura et al. 2012), Vaccinium corymbosum (Santos et al. 2022; Zheng et al. 2023b), Vandopsis gigantea (Ran et al. 2017), Vitis vinifera (Jayawardena et al. 2018).
Known distribution: Australia (Moslemi and Taylor 2015), Canada (Adams and Walker 2023), China (Liu et al. 2007; Maharachchikumbura et al. 2012; Qian et al. 2014; Liu et al. 2017; Gu et al. 2022; Jiang et al. 2022; Peng et al. 2022; Qiu et al. 2022; Sun et al. 2023; Zheng et al. 2023b; Hsu et al. 2024; Razaghi et al. 2024), Italy (Maharachchikumbura et al. 2014b; Jayawardena et al. 2018; Vu et al. 2019), Japan (Nozawa et al. 2019), Portugal (Santos et al. 2022), Tunisia (Hlaiem et al. 2018).
Notes: Pestalotiopsis hainanensis was first introduced by Liu et al. (2007) from living stems of Podocarpus macrophyllus in China. Subsequently, Pe. intermedia and Pe. linearis were described by Maharachchikumbura et al. (2012), with Pe. intermedia identified as a saprobe or endophyte on unidentified trees in China, and Pe. linearis as an endophyte on living leaves of Trachelospermum sp. and Tsuga sp. in China. Later, Pe. chamaeropis was introduced by Maharachchikumbura et al. (2014b) from Chamaerops humilis in Italy. In 2022, Pe. appendiculata, Pe. daliensis, and Pe. rosarioides were isolated from healthy leaves of Rhododendron decorum in China (Gu et al. 2022), while Pe. tumida was introduced from living stems of Rosa chinensis in China (Peng et al. 2022).
Phylogenetic analyses reveal that these eight species—Pe. appendiculata, Pe. chamaeropis, Pe. daliensis, Pe. hainanensis, Pe. intermedia, Pe. linearis, Pe. rosarioides, and Pe. tumida—clustered together in both the combined three-gene phylogeny and single-gene trees (Fig. 9 and Fig. S2). The results of GCPSR, PTP, and mPTP analyses further support their classification as a single species. Moreover, the sequence similarity within this clade ranges from 99.07% to 100% for ITS, 98.86% to 100% for tef1, and 97.67% to 100% for tub2. Additionally, genome-wide similarity among five isolates within this clade exceeds 97.73%, with core genes comprising 91.12% of the pan-genome. While the type strains of these species exhibit some morphological differences in conidial characteristics (Fig. 12), minor morphological variations alone are not sufficient for species delimitation in this study, which primarily relies on molecular evidence (see discussion). Therefore, molecular data provide a more reliable basis for species classification. Based on these findings, we synonymize Pe. appendiculata, Pe. chamaeropis, Pe. daliensis, Pe. intermedia, Pe. linearis, Pe. rosarioides, and Pe. tumida under Pe. hainanensis.
Pestalotiopsis unicolor Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 122 (2012)
= Pestalotiopsis jiangxiensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 7 (2017)
= Pestalotiopsis taxicola Y.F. Wang & C.J. You, MycoKeys 102: 209 (2024)
See Maharachchikumbura et al. (2012) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Hunan Province, Yizhang County, Mangshan, on living leaves of Rhododendron sp., 12 April 2002, W.P. Wu (HMAS046974, holotype; MFLU 12-0417, isotype); ex-type NN0469740 = MFLUCC 12-0276.
Host range: Camellia sp., Eurya sp. (Liu et al. 2017), Pandanus sp. (Tibpromma et al. 2018a), Rhododendron sp. (Maharachchikumbura et al. 2012), Taxus chinensis (Wang et al. 2024).
Known distribution: China (Maharachchikumbura et al. 2012; Liu et al. 2017; Wang et al. 2024), Thailand (Tibpromma et al. 2018a).
Notes: Pestalotiopsis unicolor, Pe. jiangxiensis, and Pe. taxicola were all introduced from China, isolated from Rhododendron spp. (Maharachchikumbura et al. 2012), Camellia spp. (Liu et al. 2017), and diseased needles of Taxus chinensis (Wang et al. 2024), respectively. Phylogenetic analyses show that these three species form a well-supported clade (BS/PP = 95%/0.99, Fig. 9). GCPSR and PTP analyses further confirm that they represent a single species. Sequence similarity of type strain within this clade ranges from 99.80% to 100% for ITS, 97% for tef1, and 99.54% to 100% for tub2. Morphologically, Pe. jiangxiensis differs from Pe. unicolor and Pe. taxicola by producing longer and wider conidia, while Pe. taxicola is distinguished by having shorter and versicolorous three median cells compared to Pe. jiangxiensis and Pe. unicolor (Maharachchikumbura et al. 2012; Liu et al. 2017; Wang et al. 2024; Fig. 12, Table S12). However, given that morphological characteristics alone are insufficient for species delimitation within Pestalotiopsis, molecular data provide a more reliable means of classification (see discussion). Based on these findings, we synonymize Pe. jiangxiensis and Pe. taxicola under Pe. unicolor.
The Pestalotiopsis rosea clade
The phylogenetic network of Pestalotiopsis revealed a complex structure within the Pe. rosea clade (Fig. 8), which includes multiple taxa and exhibits a considerable number of parallel edges, indicating extensive gene conflict. The three-locus phylogenetic tree of Pestalotiopsis indicated that 58 species clustered into the Pe. rosea clade with high support values (71% BS/ 0.99 PP; Fig. 9). However, most evolutionary branches within the Pe. rosea clade were short and poorly supported. Based on these observations, a more focused three-locus combined phylogenetic tree and single-gene trees of the Pe. rosea clade were reconstructed (Figs. 24, S8a–c). The alignment for the three loci comprised 1,795 characters, including 533 from ITS, 494 from tef1, and 768 from tub2, with alignment gaps included. The analyses incorporated 139 ingroup taxa and two neighboring species, Pe. digitalis and Pe. jinchanghensis. Details of the alignment of ML and BI analyses for the Pe. rosea clade are provided in Table S11. For the entire Pe. rosea clade, the topologies derived from ML and BI analyses were consistent. However, discrepancies were observed in the branching patterns of certain subclades within the Pe. rosea clade. The ML tree is presented here, displaying bootstrap values and posterior probabilities for strongly supported clades.
Given the inconsistencies in the topological structures of phylogenetic trees and the low support values across individual loci, significant gene conflicts and the presence of numerous paraphyletic groups are apparent within the Pe. rosea clade (Figs. 24, S8a–c). These findings from multi-locus phylogenetic analyses highlight poorly resolved species boundaries, rendering it challenging to confidently classify certain isolates included in this study. Notably, long branches were observed in the tef1 and tub2 genes for certain strains, such as Pe. camelliae-japonicae in the tef1 dataset. Despite this, there is inadequate evidence to support the hypothesis that all taxa within the Pe. rosea clade belong to a single species. As a result, we propose recognizing the Pe. rosea clade as a Pe. rosea species complex.
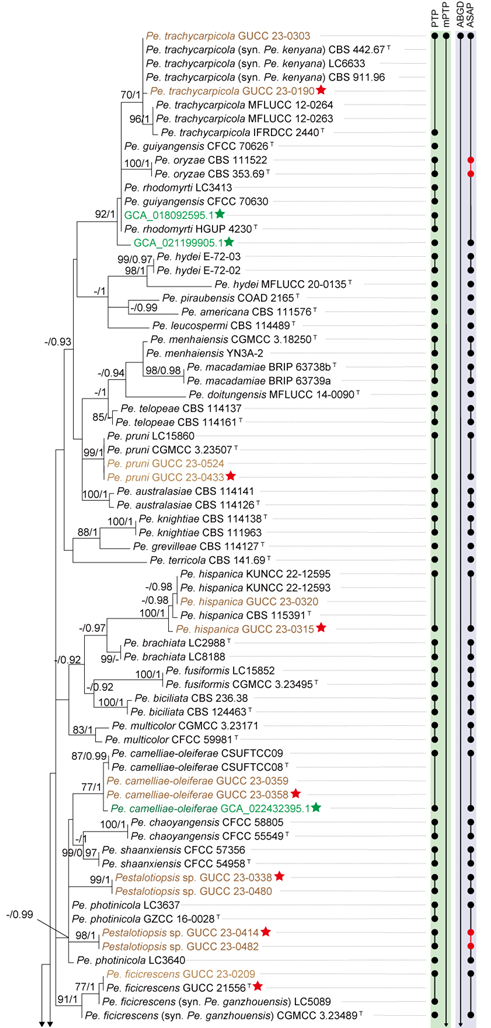
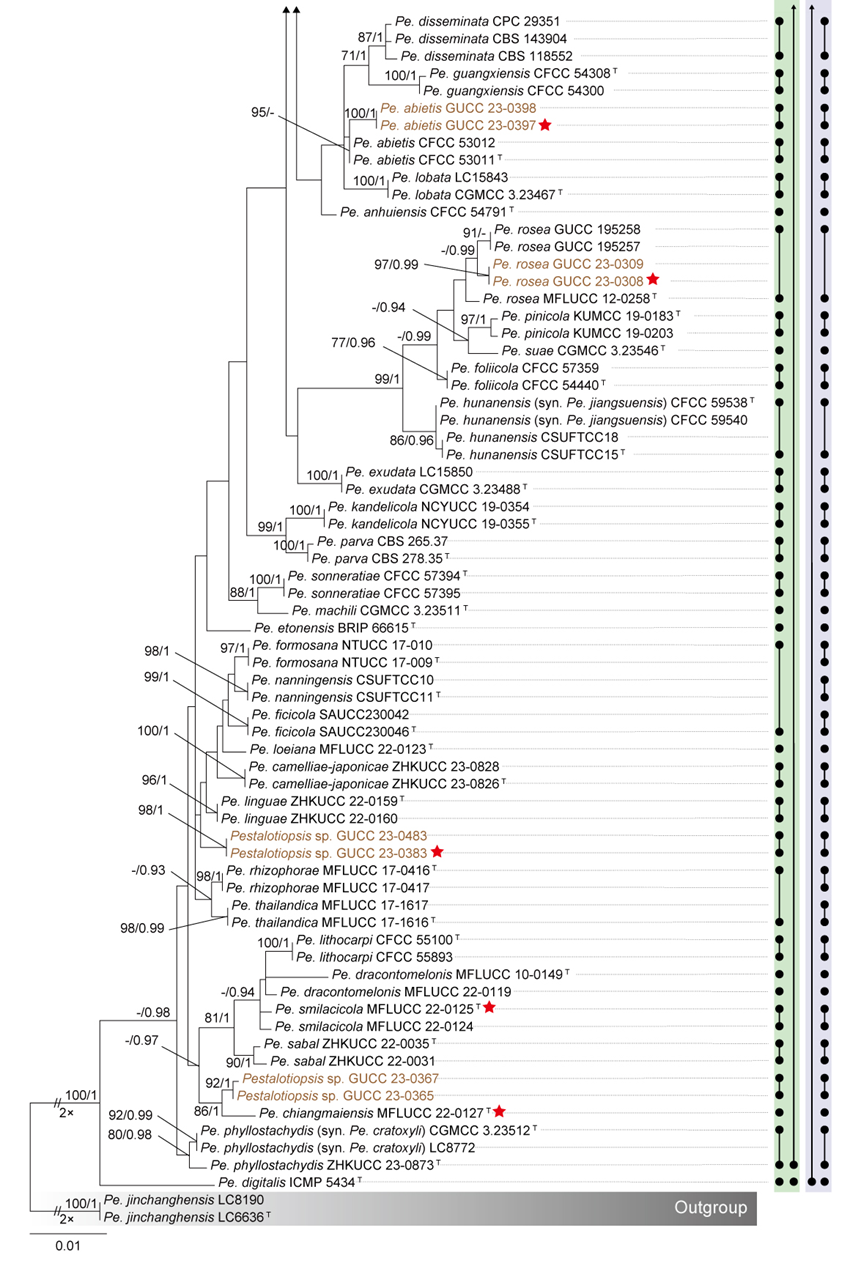
The phylogenetic analyses revealed incongruent topologies among individual gene trees, with generally low support values. In the multi-locus phylogeny, species boundaries within the Pe. rosea clade were indistinct, and several strains could not be reliably classified. Consequently, the GCPSR principle could not be applied for species delimitation. The PTP analysis identified 66 MOTUs (Fig. 24); however, it divided different strains of Pe. rhodomyrti and Pe. photinicola into separate MOTUs, likely due to the method’s tree-based algorithm, which can misclassify paraphyletic groups. The mPTP method recognized the entire Pe. rosea clade as a single MOTU, whereas the distance-based ABGD method merged this clade with the closely related Pe. digitalis into one MOTU. In contrast, ASAP delineated the Pe. rosea clade into 63 MOTUs.
The phylogenetic network of the Pe. rosea clade revealed several conflicting evolutionary signals, indicating the likelihood of recombination, which was statistically significant (P = 1.26 × 10⁻8; Fig. 25a). These conflicts are evident in the network's structure, characterized by boxlike polygons and irregular branch lengths. Given the extensive gene conflicts, frequent recombination, and inconclusive species boundaries observed in the phylogenetic network, we propose recognizing the Pe. rosea clade as a species complex. This designation reflects the high degree of genetic and evolutionary complexity within the clade and acknowledges the limitations of current delimitation methods in resolving its species structure. Within the Pe. rosea species complex the PHI test was conducted to assess species delimitations proposed by the PTP-based strategy. However, the test could not be effectively applied to Pe. ficicrescens and Pe. ganzhouensis (Fig. 25b) or Pe. cratoxyli and Pe. phyllostachydis (Fig. 25c) due to an insufficient number of informative sites, as these taxa contained none. In the Pe. formosana subclade (GUCC 23-0383, GUCC 23-0483, Pe. camelliae-japonicae, Pe. ficicola, Pe. formosana, Pe. linguae, Pe. loeiana, Pe. nanningensis, Pe. rhizophorae, and Pe. thailandica), the PHI test did not detect statistically significant evidence of recombination (p = 0.17; Fig. 25c). Similarly, no significant recombination was identified within the Pe. trachycarpicola subclade (Pe. guiyangensis, Pe. kenyana, Pe. oryzae, Pe. rhodomyrti, and Pe. trachycarpicola) (p = 1.0; Fig. 25e). However, it is noteworthy that this subclade was assessed using only 14 informative sites, suggesting a high degree of sequence similarity among these taxa. In contrast, within the Pe. rosea subclade (Pe. foliicola, Pe. hunanensis, Pe. jiangsuensis, Pe. pinicola, Pe. rosea, and Pe. suae), the PHI test detected statistically significant evidence of recombination (p = 0.02; Fig. 25d).
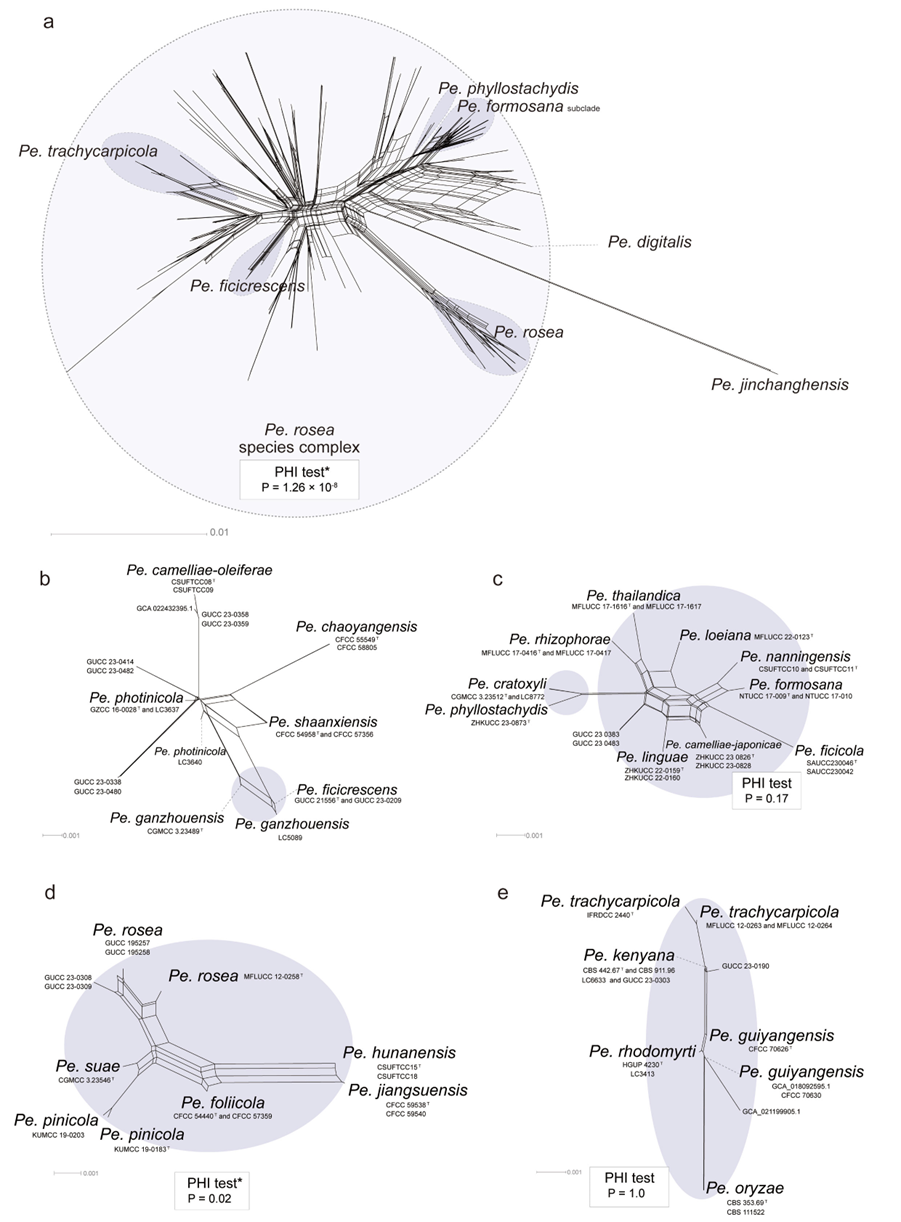
Species residing in the Pestalotiopsis rosea species complex
Pestalotiopsis abietis C.M. Tian & M. Gu, Phytotaxa 509 (1): 96 (2021)
Pestalotiopsis americana P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 240 (2024)
Pestalotiopsis anhuiensis Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 6 (2022)
Pestalotiopsis australasiae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 153 (2014)
Pestalotiopsis biciliata Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 156 (2014)
Pestalotiopsis brachiata F. Liu & L. Cai, Scientific Reports 7 (no. 866): 5 (2017)
Pestalotiopsis camelliae-japonicae Y.X. Shu & W. Dong, Mycosphere 14 (1): 2235 (2023)
Pestalotiopsis camelliae-oleiferae Qin Yang & He Li, Journal of Fungi 7 (12, no. 1080): 19 (2021)
Pestalotiopsis chaoyangensis Lu Lin & X.L. Fan, Journal of Fungi 9 (2, no. 271): 20 (2023)
Pestalotiopsis chiangmaiensis Y.R. Sun & Yong Wang bis, Microbiology Spectrum 11 (1, e03987-22): 12 (2023)
Pestalotiopsis disseminata (Thüm.) Steyaert, Bulletin du Jardin Botanique de l'État à Bruxelles 19: 174 (1948)
Pestalotiopsis doitungensis X.Y. Ma, K.D. Hyde & J.C. Kang, Phytotaxa 419 (3): 274 (2019)
Pestalotiopsis dracontomelonis Maharachch. & K.D. Hyde, Fungal Diversity 72: 15 (2015)
Pestalotiopsis etonensis C. Lock, Vitelli, Holdom, Y.P. Tan & R.G. Shivas, Persoonia 44: 437 (2020)
Pestalotiopsis exudata P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 242 (2024)
Pestalotiopsis ficicola Z.X. Zhang, J.W. Xia and X.G. Zhang, Microorganisms 11(7, no. 1627): 6 (2023)
Pestalotiopsis ficicrescens Qi Yang & Yong Wang bis, Mycosphere 14 (1): 717 (2023)
Pestalotiopsis foliicola Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 13 (2022)
Pestalotiopsis formosana Ariyaw. & K.D. Hyde, Mycosphere 9 (5): 1002 (2018)
Pestalotiopsis fusiformis P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 245 (2024)
Pestalotiopsis grevilleae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 162 (2014)
Pestalotiopsis guangxiensis Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 14 (2022)
Pestalotiopsis guiyangensis W.S. Zhang & X.L. Fan, Journal of Fungi 10 (7, no. 475): 10 (2024)
Pestalotiopsis hispanica F. Liu, L. Cai & Crous, Studies in Mycology 92: 362 (2019)
Pestalotiopsis hunanensis Qin Yang & He Li, Journal of Fungi 7 (12, no. 1080): 20 (2021)
Pestalotiopsis hydei Huanraluek & Jayaward., Phytotaxa 479 (1): 35 (2021)
Pestalotiopsis kandelicola Norph., C.H. Kuo & K.D. Hyde, Fungal Diversity 103: 233 (2020)
Pestalotiopsis knightiae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 168 (2014)
Pestalotiopsis leucospermi P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 246 (2024)
Pestalotiopsis linguae Hua Li, Manawas. & Y.X. Zhang, Phytotaxa 587 (3): 241 (2023)
Pestalotiopsis lithocarpi Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 17 (2022)
Pestalotiopsis lobata P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 247 (2024)
Pestalotiopsis loeiana Y.R. Sun & Yong Wang bis, Microbiology Spectrum 11 (1, e03987-22): 14 (2023)
Pestalotiopsis macadamiae R.G. Shivas & Akinsanmi, Plant Disease 101 (1): 49 (2017)
Pestalotiopsis machili P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 248 (2024)
Pestalotiopsis menhaiensis Y.C. Wang, X.C. Wang & Y.J. Yang, Plant Disease 103 (10): 2553 (2019)
Pestalotiopsis multicolor Y.F. Wang & C.J. You, MycoKeys 102: 210 (2024)
Pestalotiopsis nanningensis Qin Yang & He Li, Journal of Fungi 7 (12, no. 1080): 22 (2021)
Pestalotiopsis oryzae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 172 (2014)
Pestalotiopsis parva Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 175 (2014)
Pestalotiopsis photinicola Y.Y. Chen, K.D. Hyde, J.K. Liu & Maharachch., Mycosphere 8 (1): 107 (2017)
Pestalotiopsis phyllostachydis H.J. Zhao & W. Dong, Phytotaxa 633 (1): 76 (2024)
Pestalotiopsis pinicola Tibpromma, Karunaratha & Mortimer, Pathogens 8 (4, no. 285): 12 (2019)
Pestalotiopsis piraubensis V.P. Abreu & O.L. Pereira, Fungal Diversity 117: 149 (2022)
Pestalotiopsis pruni P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 252 (2024)
Pestalotiopsis rhizophorae Norph., T.C. Wen & K.D. Hyde, Mycosphere 10 (1): 552 (2019)
Pestalotiopsis rhodomyrti Yu Song, K. Geng, K.D. Hyde & Yong Wang bis, Phytotaxa 126 (1): 27 (2013)
Pestalotiopsis rosea Maharachch. & K.D. Hyde, Fungal Diversity 56 (1): 118 (2012)
Pestalotiopsis sabal Y.R. Xiong & Manawas., Current Research in Environmental & Applied Mycology 12 (1): 312 (2022)
Pestalotiopsis shaanxiensis Ning Jiang, Microbiology Spectrum 10 (6, e03272-22): 20 (2022)
Pestalotiopsis smilacicola Y.R. Sun & Yong Wang bis, Microbiology Spectrum 11 (1, e03987-22): 15 (2023)
Pestalotiopsis sonneratiae Ning Jiang, Mycotaxon 137 (4): 731 (2023)
Pestalotiopsis suae H.W. Shen, R. Gu & Z.L. Luo, Frontiers in Microbiology 13 (no. 1016782): 14 (2022)
Pestalotiopsis telopeae Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 178 (2014)
Pestalotiopsis terricola F. Liu, L. Cai & Crous, Studies in Mycology 92: 371 (2019)
Pestalotiopsis thailandica Norph., Doilom & K.D. Hyde, Mycosphere 10 (1): 555 (2019)
Pestalotiopsis trachycarpicola Y.M. Zhang & K.D. Hyde, Cryptogamie Mycologie 33 (3): 315 (2012)
Notes: The three-locus concatenated phylogeny (Fig. 9) showed that the Pestalotiopsis rosea clade formed a well-supported monophyletic lineage (BS/PP = 71%/0.99). However, some internal branches were short, and most nodes lacked strong support. Further phylogenetic analyses revealed incongruent topologies among individual gene trees with generally low support values. The multi-locus phylogeny also exhibited poorly defined species boundaries, with several strains difficult to classify precisely, thus preventing the application of the GCPSR principle for species delimitation. The results of different species delimitation methods were inconsistent: both PTP and ASAP divided the Pe. rosea clade into multiple MOTUs, whereas mPTP recognized the entire clade as a single MOTU. The phylogenetic network and PHI test further indicated extensive recombination events within this lineage. Genomic analyses showed high similarity among genomes within the Pe. rosea clade, with ANI values ranging from 94.36% to 99.22%. Taken together, the Pe. rosea clade exhibits evidence of gene tree conflict, high recombination rates, and ambiguous species boundaries, yet high genomic similarity among members. Therefore, we tentatively treat it as the Pestalotiopsis rosea species complex, pending the inclusion of additional type strain genomes to clarify its taxonomic status in the future.
Synonymies in the Pestalotiopsis rosea species complex
Pestalotiopsis ficicrescens Qi Yang & Yong Wang bis, Mycosphere 14 (1): 717 (2023)
= Pestalotiopsis ganzhouensis P. Razaghi, F. Liu & L. Cai, Studies in Mycology 109: 246 (2024)
See Hyde et al. (2023) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Guizhou Province, Guiyang City, leaf spots of Ficus tikoua, 17 July 2019, J. Yuan, HGUP 861 (HGUP 861, holotype); ex-type GUCC 21-556.
Host range: Ficus tikoua (Hyde et al. 2023), Cinnamomum camphora (Razaghi et al. 2024), Cleyera japonica (Razaghi et al. 2024), Oleaceae (Razaghi et al. 2024).
Known distribution: China (Hyde et al. 2023; Razaghi et al. 2024).
Notes: Pestalotiopsis ficicrescens was introduced by Hyde et al. (2023) from leaf spots on Ficus tikoua in China, while Pe. ganzhouensis was described by Razaghi et al. (2024) from Cinnamomum camphora in China. Upon examination of the genetic data for Pe. ficicrescens, we identified an error in its tef1 sequence. In this study, we provide a corrected tef1 sequence for Pe. ficicrescens. With this updated sequence, Pe. ficicrescens and the type strain of Pe. ganzhouensis form a well-supported sister clade (93% BS / 1 PP) in phylogenetic analyses (Fig. 9). Species delimitation analyses using the GCPSR and PTP frameworks recognize Pe. ficicrescens and Pe. ganzhouensis as a single species. The ex-type culture of Pe. ficicrescens (GUCC 21-556) exhibits high nucleotide similarity to the ex-type culture of Pe. ganzhouensis (CGMCC 3.23489), with sequence identities of 99.24% for ITS (524/528, including one gap), 99.38% for tef1 (480/483, including three gaps), and 99.87% for tub2 (757/758, no gaps). Morphologically, both species produce similar conidia, although Pe. ficicrescens has slightly narrower conidia (3–5.5 μm) (Hyde et al. 2023) compared to Pe. ganzhouensis (6–7.5 μm) (Razaghi et al. 2024; Fig. 12, Table S12). Given their phylogenetic placement and high genetic and morphological similarity, we synonymize Pe. ganzhouensis under Pe. ficicrescens.
Pestalotiopsis hunanensis Qin Yang & He Li, Journal of Fungi 7 (12, no. 1080): 20 (2021)
= Pestalotiopsis jiangsuensis Li Hua Zhu, Hui Li & D.W. Li, Journal of Fungi 10 (3, no. 230): 11 (2024)
See Li et al. (2021b) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Hunan Province, Xiangtan City, from leaf spots of Camellia oleifera, 7 November 2020, H. Li (CSUFT015, holotype); ex-type CSUFTCC15.
Host range: Camellia oleifera (Li et al. 2021b), Pinus massoniana (Li et al. 2024a).
Known distribution: China (Li et al. 2021b; Li et al. 2024a).
Notes: Phylogenetic analyses showed that Pestalotiopsis hunanensis and Pe. jiangsuensis clustered together with strong statistical support (BS/PP = 90%/0.99; Fig. 9). Both PTP and ASAP analyses recognized them as a single MOTU (Fig. 24). The ex-type strains of the two species exhibited extremely high sequence similarity across all loci examined: 100% for ITS (501/501), 100% for tef1 (481/481), and 99.77% for tub2 (439/440). Morphologically, they are nearly identical, differing only slightly in conidial width—Pe. hunanensis (7–10.5 μm) being slightly broader than Pe. jiangsuensis (6.2–8.7 μm) (Li et al. 2021b; Li et al. 2024a). However, such minor variation in conidial width is not considered sufficient for species delimitation within this genus (see discussion). Based on these molecular and morphological data, Pe. jiangsuensis is herein proposed as a synonym of Pe. hunanensis.
Pestalotiopsis phyllostachydis H.J. Zhao & W. Dong, Phytotaxa 633 (1): 76 (2024)
= Pestalotiopsis cratoxyli P. Razaghi, F. Liu, M. Raza & L. Cai, Studies in Mycology. 109: 241 (2024)
See Zhao et al. (2024) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Guangdong Province, Guanghzou City, South China National Botanical Garden, isolated from leaves of Phyllostachys sulphurea, 17 June 2021, H.J. Zhao, HNZW177 (MHZU 23-0119, holotype); ex-type culture ZHKUCC 23-0873.
Host range: Cratoxylum cochinchinense, Excoecaria cochinchinensis (Razaghi et al. 2024), Phyllostachys sulphurea (Zhao et al. 2024).
Known distribution: China (Razaghi et al. 2024; Zhao et al. 2024).
Notes: Pestalotiopsis phyllostachydis was introduced by Zhao et al. (2024) from Phyllostachys sulphurea in China. Later the same year, Razaghi et al. (2024) described Pe. cratoxyli from Cratoxylum cochinchinense and Excoecaria cochinchinensis in China. Phylogenetic analyses revealed that these two species clusters together with strong support (BS/PP = 92%/1, Fig. 9). Species delimitation using the GCPSR framework and the PTP method recognizes Pe. phyllostachydis and Pe. cratoxyli as a single species. The ex-type culture of Pe. phyllostachydis (ZHKUCC 23-0873) exhibits high nucleotide similarity to the ex-type culture of Pe. cratoxyli (CGMCC 3.23512), with sequence identities of 99.62% for ITS (526/528, no gaps), 99.37% for tef1 (475/478, no gaps), and 99.87% for tub2 (760/761, no gaps). Moreover, the morphological characteristics of Pe. phyllostachydis and Pe. cratoxyli are largely overlapping (Razaghi et al. 2024; Zhao et al. 2024; Fig. 12, Table S12). Hence, we synonymize Pe. cratoxyli under Pe. phyllostachydis.
Pestalotiopsis trachycarpicola Y.M. Zhang & K.D. Hyde, Cryptogamie Mycologie 33 (3): 315 (2012)
= Pestalotiopsis kenyana Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 166 (2014)
See Zhang et al. (2012a) for illustrations and descriptions of asexual morph and sexual morph.
Typus: China, Yunnan Province, Kunming, Kunming Botanical Gardens, on leaf spots on living leaves of Trachycarpus fortunei, March 2011, K.D. Hyde OP068 (IFRD 9026, holotype), ex-type IFRDCC 2440.
Host range: Camellia oleifera (Li et al. 2022), Camellia sinensis (Liu et al. 2017; Manawasinghe et al. 2021; Tsai et al. 2021; Xia et al. 2022), Camellia sp. (Liu et al. 2017), Castanea henryi (Jiang et al. 2021), Castanea mollissima (Jiang et al. 2021), Castanopsis fissa (Jiang et al. 2022), Castanopsis hystrix (Jiang et al. 2022), Celtis formosana (Tennakoon et al. 2021), Chrysophyllum sp. (Maharachchikumbura et al. 2012), Coffea sp. (Maharachchikumbura et al. 2014b), Cyclobalanopsis fleuryi (Jiang et al. 2022), Cyclobalanopsis glauca (Jiang et al. 2022), Cyclobalanopsis neglecta (Jiang et al. 2022), Dendrobium loddigesii (Ma et al. 2019), Historic documents (Escudero-Leyva et al. 2023), Magnolia garrettii (de Silva et al. 2021), Mangifera indica (Shu et al. 2020), Marine animals (Godinho et al. 2019), Ophiocordyceps sp. (Hsu et al. 2024), Paeonia suffruticosa (Li et al. 2022), Panax notoginseng (Lan et al. 2023), Photinia × fraseri (Xu et al. 2022), Pinus bungeana (Qi et al. 2021), Podocarous macrophyllus (Maharachchikumbura et al. 2012; Zhang et al. 2013b), Quercus aliena (Jiang et al. 2022), Raw material from agar-agar (Maharachchikumbura et al. 2014b), Rhododendron agastum (Li et al. 2024b), Rhododendron cyanocarpum (Gu et al. 2022), Rosa roxburghii (Zhang et al. 2021b), Schima sp. (Maharachchikumbura et al. 2012), Sorghum bicolor (Fan et al. 2021), Stereocaulon japonicum (Oh et al. 2020), Symplocos sp. (Maharachchikumbura et al. 2012), Taxus chinensis (Wang et al. 2024), Trachycarpus fortunei (Maharachchikumbura et al. 2012; Zhang et al. 2012a), Vaccinium corymbosum (Zheng et al. 2023b), Vaccinium sp. (Araujo et al. 2023), Vitis vinifera (Jayawardena et al. 2015; Ghuffar et al. 2018).
Known distribution: Antarctica (Godinho et al. 2019), Brazil (Araujo et al. 2023), China (Liu et al. 2017; Jiang et al. 2021; Manawasinghe et al. 2021; Tennakoon et al. 2021; Tsai et al. 2021; Jiang et al. 2022; Li et al. 2022; Xia et al. 2022; Hsu et al. 2024; Li et al. 2024b), Costa Rica (Escudero-Leyva et al. 2023), Kenya (Maharachchikumbura et al. 2014b), Korea (Oh et al. 2020), Pakistan (Ghuffar et al. 2018), Thailand (de Silva et al. 2021), Unknown country (Maharachchikumbura et al. 2014b).
Notes: Pestalotiopsis trachycarpicola clusters together with four other species—Pestalotiopsis guiyangensis, Pestalotiopsis kenyana, Pestalotiopsis oryzae, and Pestalotiopsis rhodomyrti—with high support in both the ITS and tub2 phylogenetic trees (BS ≥ 70%, PP ≥ 0.95, Fig. S8a, c). Furthermore, strong support is observed in the concatenated three-gene phylogenetic tree (88% BS / 1 PP, Fig. 9). However, we observed that Pestalotiopsis oryzae and GCA_021199905.1 form an anomalous branch, and in the tef1 phylogenetic tree, they are positioned relatively distant from Pe. trachycarpicola. Therefore, this study focuses only on the taxonomic status of Pestalotiopsis kenyana, which forms a short branch with Pe. trachycarpicola (BS 70% / PP 0.99, Fig. 9). Nucleotide similarities among the type strains of Pe. trachycarpicola and Pe. kenyana are 99.81% (539/540, no gaps) for ITS, 100% (488/488) for tef1, and 99.78% (446/447, no gaps) for tub2. Morphologically, Pe. trachycarpicola has smaller conidia (19–24.9 × 5.3–6.3 μm) than Pe. kenyana (22–29 × 7–9 μm) and shorter basal appendages (2.7–5.5 μm vs. 3–20 μm) (Fig. 12, Table S12). Since molecular data primarily guide species delineation in this study (see discussion), we synonymize Pe. kenyana under Pe. trachycarpicola.
Pseudopestalotiopsis
Phylogenetic analyses and phylogenetic species recognition of Pseudopestalotiopsis
To evaluate the genus Pseudopestalotiopsis, phylogenetic trees based on single loci (ITS, tef1, and tub2) and a three-locus concatenated tree were constructed (Fig. S9). The combined tree (Fig. 26) incorporated alignments of the ITS (530 bp), tef1 (496 bp), and tub2 (745 bp) regions, including alignment gaps. This analysis included 59 isolates, with Neopestalotiopsis acrostichi MFLUCC 17-1754 and N. rhizophorae MFLUCC 17-1550 as outgroup taxa. Among these isolates, 13 were newly sequenced in this study, while three sequences were retrieved from GenBank (originally identified as Pestalotiopsis fici GCA_000516985.1, Pseudopestalotiopsis chinensis GCA_028028555.1, and Pseudopestalotiopsis theae GCA_015881745.1). However, based on our re-evaluation, these sequences should be re-identified as Pseudopestalotiopsis chinensis GCA_000516985.1, Pseudopestalotiopsis chinensis GCA_028028555.1, and Pseudopestalotiopsis camelliae-sinensis GCA_015881745.1 (Figs. 26, 28). The ML analysis, conducted using RAxML-NG, resulted in a best-scoring tree (Fig. 26) with a final likelihood value of -5899.027326. Notably, the ML and BI trees shared consistent topologies. For clarity, only the ML tree is presented (Fig. 26), with bootstrap values and posterior probabilities provided for well-supported clades. Table S13 summarizes the alignment statistics and the parameters used for ML and BI analyses.
According to the phylogenetic analyses, Pseudopestalotiopsis is divided into multiple subclades. In this study, three isolates (GUCC 23-0429, GUCC 23-0443, and GUCC 23-0444), together with two genome assemblies from public databases (GCA_000516985.1, GCA_028028555.1), formed a clade with four strains identified as Ps. chinensis. Two additional isolates (GUCC 23-0387 and GUCC 23-0471) formed an independent lineage that is sister to Ps. ignota with maximum support values (100% BS / 1 PP). Additionally, two isolates (GUCC 23-0424 and GUCC 23-0430) clustered in a distinct clade, here referred to as the Ps. ampullacea clade, which included six strains representing three different Pseudopestalotiopsis species. Notably, branches within the Ps. ampullacea clade are extremely short, suggesting minimal genetic divergence among the included taxa. Furthermore, two other isolates (GUCC 23-0428 and GUCC 23-0442), along with one genome assembly (GCA_015881745.1), clustered with four strains identified as Ps. camelliae-sinensis. Lastly, isolates MFLUCC 22-0023a and MFLUCC 22-0023b formed a well-supported monophyletic lineage (97% BS / 1.00 PP).
To detect concordant branches and define evolutionary lineages within Pseudopestalotiopsis following the GCPSR principle, single-gene and multi-locus phylogenetic analyses were conducted. Conflicts among individual loci were evident (Fig. S9). The ITS phylogram (Fig. S9a) provided limited discriminatory power for most taxa, except for Ps. ignota, GUCC 23-0387, and GUCC 23-0471, which formed distinct long branches. In contrast, the tef1 phylogram (Fig. S9b) showed higher congruence with the concatenated tree and offered more robust support for species delineation. The multi-locus phylogenetic analyses significantly improved overall resolution and provided more reliable species boundaries for Pseudopestalotiopsis compared to single-locus trees (Fig. 26). The GCPSR framework was further applied to evaluate lineage concordance and refine species delimitations, leading to several proposed taxonomic adjustments. Within the Ps. cocos clade (Ps. ampullacea, Ps. avicenniae, Ps. camelliae-sinensis, Ps. chinensis, Ps. cocos, Ps. elaeidis, Ps. ignota, Ps. indica, and Ps. vietnamensis), the taxa clustered together with moderate to strong support in the three-locus concatenated phylogeny (BS/PP = 75%/0.93; Fig. 26). Similarly, in the tub2 phylogeny (Fig. S9c), they formed a short-branched clade with high support (BS/PP = 92%/1), and in the tef1 phylogeny (Fig. S9b), their branches were not clearly separated, making species delimitation difficult. These results conform to the GCPSR criterion, supporting their treatment as a IEL. However, within the three-locus tree, Ps. ignota, Ps. vietnamensis, GUCC 23-0387, and GUCC 23-0471 exhibited long branches, preventing a definitive decision on synonymization. As a preliminary revision, Ps. avicenniae and Ps. elaeidis are proposed here as potential synonyms of Ps. ampullacea. Pseudopestalotiopsis thailandica is considered a potential synonym of Ps. rhizophorae. In addition, two strains, MFLUCC 22-0023a and MFLUCC 22-0023b, formed a strongly supported monophyletic lineage (100% BS / 1 PP; Fig. 26), which was sister to Ps. ixorae, Ps. solicola, and Ps. theae within a well-supported clade (97% BS / 1 PP; Fig. 26). Separate analyses of tef1 and tub2 genes further confirmed that MFLUCC 22-0023a and MFLUCC 22-0023b represent a distinct evolutionary lineage. Based on combined molecular evidence, these two strains are recognized as representing a new species, which is formally described in the Taxonomy section below.
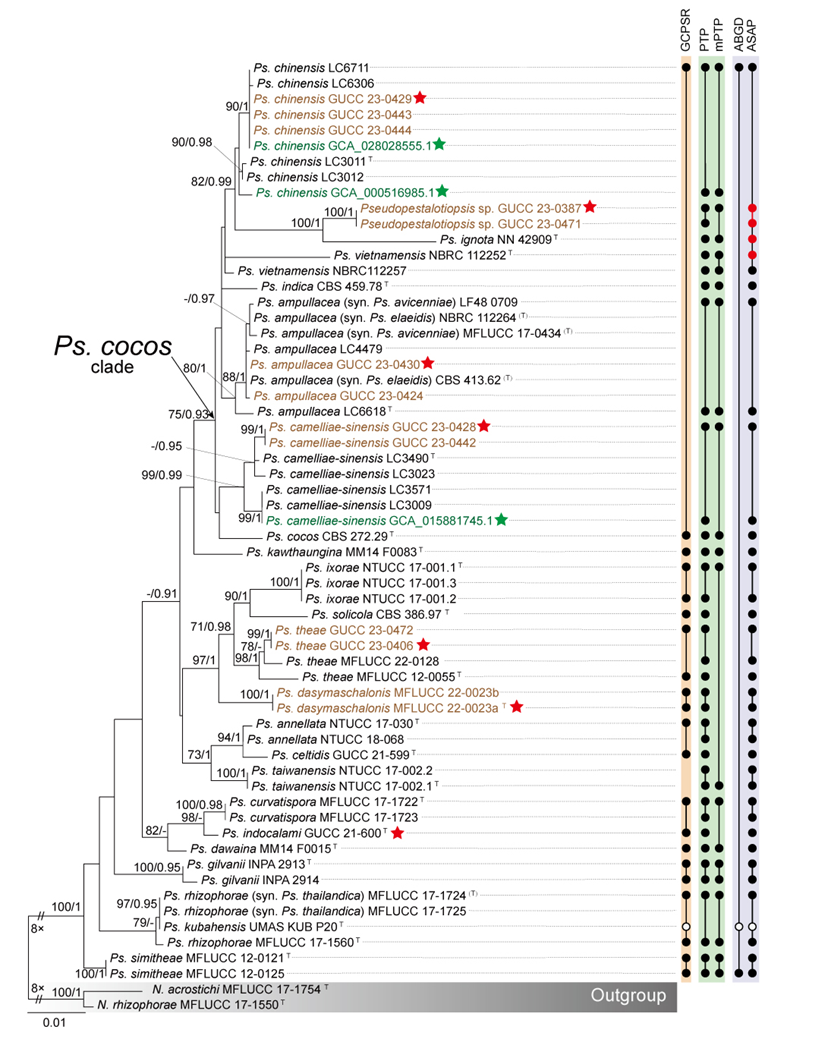
During the species delimitation analyses of Pseudopestalotiopsis using PTP, mPTP, ABGD, and ASAP, outgroup taxa were excluded to ensure more accurate delineation of species boundaries. The PTP analysis did not support the merger of several closely related taxa. For example, PTP recognized different strains of Ps. theae as separate species and clearly distinguished Ps. annellata from Ps. celtidis. In addition, PTP did not support the consolidation of the Ps. cocos clade or the merging of Ps. curvatispora and Ps. indocalami (Fig. 26). In contrast, the mPTP analysis yielded results that differed from those of PTP. Specifically, mPTP grouped the two strains of Ps. vietnamensis (NBRC 112252 and NBRC 112257) into a single MOTU, and likewise clustered Ps. ignota, GUCC 23-0387, and GUCC 23-0471 together in one MOTU. It also combined Ps. ixorae, Ps. solicola, Ps. theae, Ps. annellata, Ps. celtidis, and Ps. taiwanensis into a single MOTU, and merged Ps. curvatispora, Ps. indocalami, and Ps. dawaina into another. Unlike the tree-based methods, the distance-based ABGD approach recognized all Pseudopestalotiopsis taxa as a single species, indicating an overly conservative delimitation. The ASAP results were generally consistent with those of PTP, except for minor differences at the upper part of the phylogeny, where ASAP grouped Ps. chinensis and Ps. vietnamensis (NBRC 112257) into a single MOTU.
The discrepancies among these delimitation methods suggest the presence of potential recombination events within Pseudopestalotiopsis. The phylogenetic network analysis revealed numerous conflicting signals, which were statistically significant (P = 1.59 × 10⁻5; Fig. 27). These conflicting signals were mainly concentrated near the basal portion of the network, whereas individual species extended as nearly parallel branches. This pattern indicates historical or ongoing genetic exchange among lineages and highlights the need for caution when proposing new taxa. Nevertheless, the distinct taxonomic status of isolates MFLUCC 22-0023a and MFLUCC 22-0023b is strongly supported by both GCPSR and PTP analyses. Furthermore, their markedly long and isolated branch lengths within the phylogenetic network provide additional evidence supporting their recognition as a separate species.
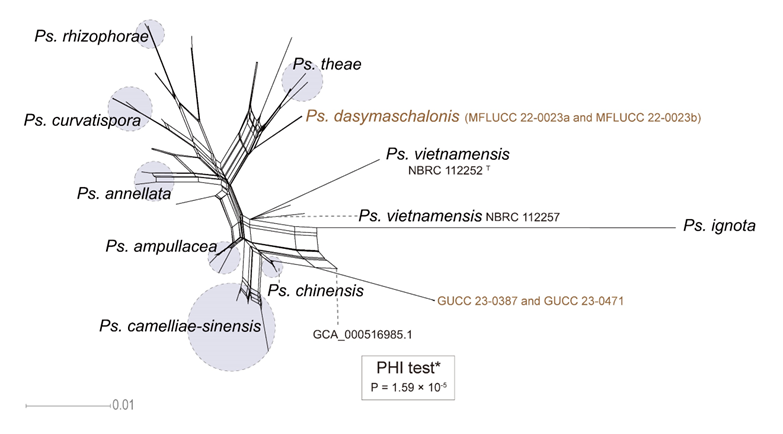
Whole-genome data and phylogenomic assessment of Pseudopestalotiopsis
Due to sampling limitations, this study includes only ten Pseudopestalotiopsis genomes, three of which were obtained from the NCBI genome database, while the remaining seven were newly sequenced in this study. These genomes represent five known species, one novel species, and one unidentified species. Using OrthoFinder, we identified 10,244 single-copy orthologous genes across all sequenced samples for phylogenetic analysis, incorporating ten Pseudopestalotiopsis strains along with the outgroup taxa Neopestalotiopsis acrostichi MFLUCC 17-1754 and N. rhizophorae MFLUCC 17-1550.
The resulting phylogenomic tree revealed that the two Ps. camelliae-sinensis strains (GCA_015881745.1 and GUCC 23-0428) and the three Ps. chinensis strains (GCA_028028555.1, GCA_000516985.1, and GUCC 23-0429) formed a closely related cluster, making them difficult to distinguish (Fig. 28). Moreover, Ps. camelliae-sinensis and Ps. chinensis exhibited short evolutionary branch lengths with GUCC 23-0387 and GUCC 23-0430 (the Ps. cocos clade, Fig. 28). To further delineate species boundaries, we assessed the ANI values among Pseudopestalotiopsis species. The genomic similarity between Ps. camelliae-sinensis and Ps. chinensis reached 98.83%, while their similarity to GUCC 23-0387 and GUCC 23-0430 exceeded 97.63% (the Ps. cocos clade, Fig. 28). A subsequent OrthoFinder analysis revealed that these seven strains share a core genome comprising 12,299 (90.92%) genes out of a total of 13,528, indicating a high degree of genetic conservation within this group (Fig. 29). Comparative genomic analysis revealed that strain MFLUCC 22-0023a exhibited less than 94.79% similarity to the other nine Pseudopestalotiopsis genomes, forming a distinct phylogenetic branch in the three-gene concatenated phylogeny and phylogenomic.
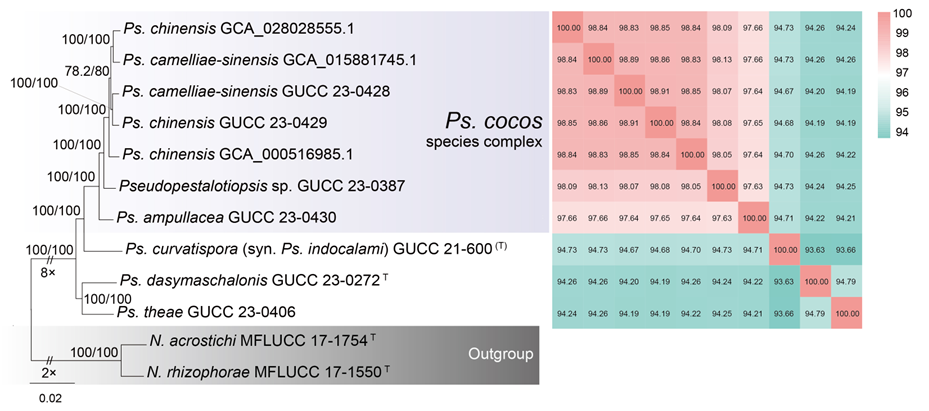
Morphology of Pseudopestalotiopsis
Figure 30 provides a detailed overview of conidial micromorphology for each Pseudopestalotiopsis species, aligned with their positions in the three-gene concatenated phylogenetic tree. Consistent with patterns observed in Neopestalotiopsis and Pestalotiopsis, the analysis reveals substantial discrepancies between molecular phylogeny and morphological characteristics. For instance, Ps. indocalami and Ps. simitheae exhibit nearly identical conidial morphology, yet they are phylogenetically distinct in the three-gene tree. Across Pseudopestalotiopsis, while the majority of species possess concolourous median cells, five species—Ps. annellata, Ps. avicenniae, Ps. curvatispora, Ps. rhizophorae, and Ps. thailandica—exhibit versicolourous median cells. Conidial lengths ranging from 18 to 38.5 μm and widths from 4.5 to 9.5 μm. Additionally, apical appendage morphology is highly variable, with lengths spanning 5.8 to 49.5 μm (Table S14). Importantly, this morphological variability shows no systematic relationship with phylogenetic affinities.
The MDS analysis yielded a stress value of 0.15, indicating that the dimensionality reduction effectively preserves the structural integrity of the dataset. Additionally, a high r-value of 0.92 signifies a strong positive correlation between the Euclidean distances in the original dataset and those in thereduced-dimensional space, confirming the reliability of the MDS configuration. While a few species exhibit morphological overlap, the majority are distinctly separated in the MDS plot (Fig. 31).
Taxonomy of Pseudopestalotiopsis
Pseudopestalotiopsis Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 180 (2014)
Notes: Pseudopestalotiopsis was segregated from Pestalotiopsis by Maharachchikumbura et al. (2014b), with Ps. theae designated as the type species. Members of the genus are characterized by conidia possessing concolourous, brown to dark brown or olivaceous median pigmented cells (Maharachchikumbura et al. 2014b). Pseudopestalotiopsis species occur mainly as plant pathogens (Darapanit et al. 2021; Ma et al. 2023; Pandey et al. 2024), particularly on leaves (Hyde et al. 2020c), but are occasionally reported as endophytes (Yu et al. 2020) or saprobes (Liu et al. 2019). In the present study, the Pseudopestalotiopsis cocos species complex is recognized, representing a group of closely related taxa with limited morphological and phylogenetic differentiation. Furthermore, three species are revised through synonymization based on molecular and phylogenetic evidence: Ps. avicenniae and Ps. elaeidis are synonymized under Ps. ampullacea, and Ps. thailandica is synonymized under Ps. rhizophorae. In addition, a new species, Pseudopestalotiopsis dasymaschalonis, is introduced.
The Pseudopestalotiopsis cocos species complex
Pseudopestalotiopsis ampullacea F. Liu & L. Cai, Scientific Reports 7 (no. 866): 12 (2017)
Pseudopestalotiopsis camelliae-sinensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 12 (2017)
Pseudopestalotiopsis chinensis F. Liu & L. Cai, Scientific Reports 7 (no. 866): 13 (2017)
Pseudopestalotiopsis cocos Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 181 (2014)
Pseudopestalotiopsis ignota Maharachch., L.D. Guo & K.D. Hyde, Mycological Progress 15 (no. 22): 5 (2016)
Pseudopestalotiopsis indica Maharachch., K.D. Hyde & Crous, Studies in Mycology 79: 182 (2014)
Pseudopestalotiopsis vietnamensis Nozawa & Kyoko Watan., Mycoscience 58 (5): 332 (2017)
Notes: The three-gene phylogenetic analyses revealed that seven species within the Ps. cocos clade (Ps. ampullacea, Ps. camelliae-sinensis, Ps. chinensis, Ps. cocos, Ps. ignota, Ps. indica, and Ps. vietnamensis) formed a well-supported monophyletic lineage (BS = 75%, PP = 0.93; Fig. 26). According to the GCPSR principle, these taxa can be regarded as representing a single MOTU. Similarly, the seven available genomes within the Ps. cocos clade (GUCC 23-0430, GUCC 23-0387, GUCC 23-0429, GCA 000516985.1, GCA 028028555.1, GUCC 23-0428, and GCA 015881745.1) clustered together in the whole-genome phylogenetic tree (Fig. 28), forming a short-branched lineage that indicates a high level of genomic concordance. All genomes exhibited ANI values exceeding 97.63%, and the proportion of core genes accounted for 90.92% of the pangenome, supporting their close genetic relatedness. However, given that the three-locus sequences of Ps. ignota, Ps. vietnamensis, and Ps. indica show relatively large genetic divergence from the remaining members of the clade, and that genomic data from their type strains are still lacking, taxonomic merging of the Ps. cocos clade is not proposed at this stage. Based on the combined evidence from multi-locus phylogenetic and genomic analyses, this lineage is herein recognized as the Pseudopestalotiopsis cocos species complex.
Pseudopestalotiopsis ampullacea F. Liu & L. Cai, Scientific Reports 7 (no. 866): 12 (2017)
= Pseudopestalotiopsis avicenniae Norph., T.C. Wen & K.D. Hyde, Mycosphere 10 (1): 558 (2019)
= Pseudopestalotiopsis elaeidis (C. Booth & J.S. Robertson) F. Liu, L. Cai & Crous, Studies in Mycology 92: 374 (2019)
See Liu et al. (2017) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: China, Yunnan Province, Xishuangbanna, Jing Mai, on Camellia sinensis, 17 April 2015, F. Liu (HMAS 247056, holotype), ex-type CGMCC 3.18157 = LC6618.
Host range: Acacia crassipes (Liu et al. 2019), Averrhoa carambola (Liu et al. 2019), Avicennia marina (Norphanphoun et al. 2019), Camellia chrysantha (Zhao et al. 2020), Camellia sinensis (Liu et al. 2017), Elaeis guineensis (Liu et al. 2019), Lauraceae (Liu et al. 2017; Liu et al. 2019), Magnolia candolli (de Silva et al. 2021), Mangifera indica (Shu et al. 2020), Oil palm hybrid (Betancourt-Ortiz et al. 2024).
Known distribution: China (Liu et al. 2017; Liu et al. 2019; Shu et al. 2020; Zhao et al. 2020; de Silva et al. 2021), Colombia (Betancourt-Ortiz et al. 2024), Indonesia (Liu et al. 2019), Myanmar (Liu et al. 2019), Nigeria (Liu et al. 2019), Thailand (Norphanphoun et al. 2019).
Notes: The phylogenetic results showed that Ps. elaeidis and Ps. avicenniae clustered close to Ps. ampullacea forming a distinct lineage (Fig. 26) with maximum support (BS =80%, PP = 1). Additionally, based on the relative distance and structure in the phylogenetic network (Fig. 27) and the results of PTP and mPTP (Fig. 26), which support that Ps. ampullacea, Ps. avicenniae, and Ps. elaeidis should be regarded as a single species. Furthermore, genetic similarity among the type strains of these three species is high, with ITS, tef1, and tub2 sequences exhibiting identity ranges of 100%, 97.9%–99.55%, and 99.86%–100%, respectively. Morphologically, the primary difference between these species in the size of their conidia. The conidia of Ps. elaeidis (31–38.5 × 6.5–9 μm) are longer than Ps. ampullacea (21–31.5 × 6.5–9 μm), and both are larger than the conidia of Ps. avicenniae (22–27 × 5–6.5 μm) (Liu et al. 2017; Nozawa et al. 2017; Norphanphoun et al. 2019). Although there are morphological differences in conidial size, the genetic similarity and species delimitation results suggest that these differences may not be sufficient to distinguish them as separate species. Based on phylogenetic evidence, we synonymize Ps. avicenniae and Ps. elaeidis under Ps. ampullacea.
Pseudopestalotiopsis dasymaschalonis Q. Zhang, C. S. Bhunjun & Yong Wang bis, sp. nov.
Index Fungorum number: IF904598; Fig. 32
Etymology: named after the host genus, Dasymaschalon.
Holotype: MFLU 23-0123
Sexual morph: Undetermined. Asexual morph: Conidiophores often reduced to conidiogenous cell. Conidiogenous cell discrete or integrated, ampulliform, hyaline, smooth or minutely verruculose, 4.5–19 × 1–4 μm. Conidia fusoid, ellipsoid, straight to slightly curved, 4-septate, 21–27.5 × 5–6.5 μm (av. ± SD = 24.77 ± 1.40 × 5.88 ± 0.36 μm); basal cell conic, hemispherical or obconic with a truncate base, hyaline, rugose and thin–walled, 3.5–6 μm long; three median cells doliiform, 13–18 μm (av. ± SD = 16.03 ± 0.94 μm) long, wall minutely verruculose, concoloured, septa darker than the rest of cell and conidium constructed at septum (second cell from base, 4.5–7.5 μm long; third cell 4.5–6 μm long; fourth cell 4.5–6.5 μm long); apical cell 4–6 μm long, hyaline, subcylindrical or obconic with a truncate base, thin-walled, slightly rugose; with 1–3 tubular apical appendages (occasionally 1), arising from the apical crest, unbranched, filiform, 17–41 μm long; basal appendage single, tubular, centric, 1.5–7 μm long.
Culture characteristics: Colonies on MEA aerial mycelium on surface flat or raised, with filiform margin (curled margin), fluffy,colony from above white, from below pale yellow, aerial mycelia flocculent, reaching 72–90 mm diam after 7 d at 25°C; on PDA flat with concave edge, low undulate margin, smooth, dense, colony from above white, from below pale yellow to ivory, aerial mycelia flocculent, reaching 65–73 mm diam after 7 d at 25°C; on SNA flat with entire edge, white, reaching 60–64 mm diam after 7 d at 25°C.
Specimen examined: Thailand, Chiang Rai Province, Doi Tung, 13 February 2022, on Dasymaschalon obtusipetalum, C. S. Bhunjun (MFLU 23-0123, holotype), ex-type MFLUCC 22-0023a; ibid., MFLUCC 22-0023b.
Notes: Based on the three-locus phylogenetic tree (Fig. 26), Ps. dasymaschalonis is clearly distinguished from other species within the genus. BLASTn searches against GenBank revealed that the closest matches for the ITS, tef1, and tub2 sequences of Ps. dasymaschalonis were Ps. theae (KM111476.1; 98.6% identity), Pseudopestalotiopsis sp. (LC114071.1; 96.75% identity), and Ps. theae (OP752136.1; 95.98% identity), respectively. Additionally, whole-genome similarity analysis indicated that Ps. dasymaschalonis (MFLUCC 22-0023a) shares 94.79% genome-wide similarity with Ps. theae (GUCC 23-0406). Based on both phylogenetic and genomic evidence, Ps. dasymaschalonis is introduced here as a new species.
Pseudopestalotiopsis rhizophorae Norph., T.C. Wen & K.D. Hyde, Mycosphere 10 (1): 560 (2019)
= Pseudopestalotiopsis thailandica Norph. & K.D. Hyde, Mycosphere 10 (1): 562 (2019)
See Norphanphoun et al. (2019) for illustrations and descriptions of asexual morph. Sexual morph not reported.
Typus: Thailand, Ngao, Ranong Province, Ngao Mangrove Forest Research Centre, leaf spots of Rhizophora apiculata, 6 December 2016, Norphanphoun Chada NG38a (MFLU 19-0793, holotype); ex-type MFLUCC 17-1560.
Host range: Camellia sinensis (Pandey et al. 2023), Rhizophora apiculate (Norphanphoun et al. 2019), Rhizophora mucronate (Norphanphoun et al. 2019).
Known distribution: India (Pandey et al. 2023), Thailand (Norphanphoun et al. 2019).
Notes: Pseudopestalotiopsis rhizophorae and Ps. thailandica were isolated from diseased leaves of Rhizophora apiculata and Rhizophora mucronata, respectively, in Thailand, as reported in the same paper by Norphanphoun et al. (2019). Pseudopestalotiopsis kubahensis was first described from the green leaves of Macaranga sp. in Malaysia (Lateef et al. 2015). These species form a closely related cluster in the phylogenetic tree (Fig. 26; BS/PP = 79%/-) and network (Fig. 27), and they are recognized as a distinct evolutionary unit, supported by GCPSR, PTP, mPTP, ASAP analyses. However, since Ps. kubahensis is only represented by ITS sequence data, with no available tef1 and tub2 sequences, we refrain from synonymizing Ps. kubahensis at this time. The ex-type culture of Ps. rhizophorae exhibited the following nucleotide similarities with the ex-type culture of Ps. thailandica: 99.62% (518/520, including two gaps) in the ITS region, 100% (463/463) in the tef1 region, and 99.23% (770/776, including three gaps) in the tub2 region. They exhibit morphological differences only in their apical appendages (Norphanphoun et al. 2019). Given the high genetic similarity and minimal morphological variation, Ps. thailandica is considered a synonym of Ps. rhizophorae.
| Species | Synonyms | Host | Known distribution |
| N. acrostichi | Acrostichum aureum (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| N. ageratinae | Ageratina adenophora (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. alpapicalis | Musa sp. and Phoenix roebelenii (Senanayake et al. 2020), Rhizophora apiculata and Rhizophora mucronate (Kumar et al. 2019) | China (Senanayake et al. 2020), Thailand (Kumar et al. 2019) | |
| N. zingiberis | N. amomi | Amomum villosum (Sun et al. 2023) | China (Sun et al. 2023) |
| N. anacardii | Mangifera indica (Kamil et al. 2012) | India (Kamil et al. 2012) | |
| N. aotearoa | Canvas (Maharachchikumbura et al. 2014b), Hevea brasiliensis (Li et al. 2021a) | New Zealand (Maharachchikumbura et al. 2014b), China (Li et al. 2021a) | |
| N. arecacearum | Caryota mitis, Dypsis lutescens, Dypsis madagascariensis, and Ptychosperma elegans (Guterres et al. 2023) | Brazil (Guterres et al. 2023) | |
| N. asiatica | Castanea mollissima (Jiang et al. 2021), Celtis formosana (Tennakoon et al. 2021), Mangifera indica (Shu et al. 2020), Unidentified tree (Maharachchikumbura et al. 2012), Vitis vinifera (Maharachchikumbura et al. 2016b) | China (Maharachchikumbura et al. 2012; Shu et al. 2020; Jiang et al. 2021), France (Maharachchikumbura et al. 2016b), and Taiwan, China (Tennakoon et al. 2021) | |
| N. australis | Byrsonima crassifolia (Moreno-Velázquez et al. 2022), Cordia dichotoma (Reddy et al. 2016), Eucalyptus sp. (Santos et al. 2020), Nopalea cochenillifera (Conforto et al. 2019), Soil (Park et al. 2016), Telopea sp. (Maharachchikumbura et al. 2014b) | Mexico (Moreno-Velázquez et al. 2022), India (Reddy et al. 2016), Brazil (Conforto et al. 2019; Santos et al. 2020), Korea (Park et al. 2016), Australia (Maharachchikumbura et al. 2014b) | |
| N. baotingensis | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) | |
| N. brachiata | Rhizophora apiculate (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| N. brasiliensis | Castanea mollissima (Jiang et al. 2021), Psidium guajava (Bezerra et al. 2018), Rosa roxburghii (Zhang et al. 2021b) | Brazil (Bezerra et al. 2018), China (Jiang et al. 2021; Zhang et al. 2021b) | |
| N. camelliae | Camellia japonica (This study) | China (This study) | |
| N. camelliae-oleiferae | Bamboo (Kwon et al. 2024), Camellia oleifera (Li et al. 2021b), Luma apiculata (Aiello et al. 2024), Tolypocladium sp. (Hsu et al. 2024) | Korea (Kwon et al. 2024), China (Li et al. 2021b), Italy (Aiello et al. 2024), Taiwan, China (Hsu et al. 2024) | |
| N. castanopsidis | Castanopsis boisii (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. caulicola | Rosa roxburghii (Zhang et al. 2024b) | China (Zhang et al. 2024b) | |
| N. caulicola | N. acericola | Acer palmatum (Zhang et al. 2024d) | China (Zhang et al. 2024d) |
| N. cavernicola | Cave (Liu et al. 2021) | China (Liu et al. 2021) | |
| N. celtidis | Celtis sinensis (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. cercidicola | Cercis chinensis (Zhang et al. 2024d) | China (Zhang et al. 2024d) | |
| N. chiangmaiensis | Ananas comosus (Tian et al. 2024), Magnolia candolli (de Silva et al. 2021), Pandanus sp. (Tibpromma et al. 2018b) | China (de Silva et al. 2021), Thailand (Tibpromma et al. 2018b; Tian et al. 2024) | |
| N. chrysea | Liquidambar formosana (Fan et al. 2022), Unidentified tree (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012; Fan et al. 2022) | |
| N. chrysea | N. umbrinospora | Unidentified host (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012) |
| N. clavispora | Acacia mearnsii (Duin et al. 2017), Actinidia arguta (Li et al. 2023b), Aegiceras corniculatum (Lin 2023), Anacardium occidentale (Rajashekara et al. 2023), Anthurium andraeanum (Daengsuwan et al. 2021), Argania spinosa (Bakry et al. 2009), Camellia chrysantha (Zhao et al. 2020), Camellia japonica (Zhang et al. 2013b), Camellia sinensis (Chen et al. 2018c), Carya illinoensis (Lazarotto et al. 2012), Carya illinoinensis (Lazarotto et al. 2014), Daphniphyllum macropodum (Xu et al. 2024), Decaying wood (Maharachchikumbura et al. 2014b), Egg mass of the sailfin sandfish (Park et al. 2018), Elaeagnus pungens (Qi et al. 2023a), Elettaria cardamomum (Biju et al. 2018), Eriobotrya japonica (Abbas et al. 2022), Euonymus japonicus (Song et al. 2022), Fragaria × ananassa (Obregón et al. 2018; Shi et al. 2022), Garcinia mangostana (Zhu et al. 2023a), Hevea brasiliensis (Solpot et al. 2024), Historic documents (Escudero-Leyva et al. 2023), Jatropha heynei (Ashoka et al. 2023), Kadsura coccinea (Xie et al. 2018), Ligustrum lucidum (Chen et al. 2019), Macadamia integrifolia (Santos et al. 2019), Macadamia sp. (Prasannath et al. 2020), Magnolia sp. (Maharachchikumbura et al. 2012), Malus domestica (Ebrahimi et al. 2022), Malus pumila (Shi et al. 2024), Mangifera indica (Shu et al. 2020), Mimusops elengi (Lokesh et al. 2017), Musa acuminata (Qi et al. 2023b), Myrica rubra (Lu et al. 2015), Orchids (Adit et al. 2022), Pandanus amaryllifolius (Gou et al. 2023), Parkinsonia aculeata (Steinrucken et al. 2017), Persea americana (Valencia et al. 2011), Phedimus aizoon (Yang et al. 2017), Phoebe bournei (Xu et al. 2023b), Phoenix dactylifera (Zhang et al. 2024c), Photinia bodinieri (Xu et al. 2023a), Photinia serratifolia (Yang et al. 2018), Platanus orientalis (Kurbetli et al. 2020), Psidium guajava (Lin et al. 2024), Quercus rubra (Maharachchikumbura et al. 2012), Rhizophora apiculata (Shara et al. 2023), Rhizophora mucronata (Hamzah et al. 2018), Rhizophoraceae mangrove plant (Xing and Guo 2011), Seaweeds (Goshima 2022), Selaginella kraussiana (McClymont et al. 2013), Spontaneously fermenting Chambourcin (Feng et al. 2021), Stryphnodendron adstringens (Carvalho et al. 2012), Styrax sp. (Elfiati et al. 2022), Suaeda australis (Khalmuratova et al. 2015), Suaeda glauca (Khalmuratova et al. 2015), Suaeda maritima (Khalmuratova et al. 2015), Syzygium cumini (Banerjee and Rana 2020), Taxus chinensis (Wang et al. 2019a), Terminalia arjuna (Tejesvi et al. 2008), Vaccinium corymbosum (Jevremović et al. 2022), Vaccinium sp. (Borrero et al. 2018), Vitis vinifera (Jayawardena et al. 2018) | Argentina (Obregón et al. 2018), Australia (Steinrucken et al. 2017), Brazil (Lazarotto et al. 2012), Canada (Bakry et al. 2009), Chile (Valencia et al. 2011), China (Li et al. 2023b), Costa Rica (Escudero-Leyva et al. 2023), India (Rajashekara et al. 2023), Indonesia (Shara et al. 2023), Iran (Basavand et al. 2020), Italy (Gilardi et al. 2019), Japan (Win et al. 2021), Korea (Lee et al. 2019), Malaysia (Hamzah et al. 2018), Morocco (Kenfaoui et al. 2024), New Zealand (McClymont et al. 2013), Pakistan (Abbas et al. 2022), Peru (Vega Gutierrez et al. 2020), Philippines (Solpot et al. 2024), Saudi Arabia (Hassan et al. 2018), Serbia (Jevremović et al. 2022), Spain (Chamorro et al. 2016), Sri Lanka (Maharachchikumbura et al. 2014b), Thailand (Daengsuwan et al. 2021), Turkey (Kurbetli et al. 2020), Uruguay (Machín et al. 2019), USA (Keith et al. 2006) | |
| N. cocoes | Cocos nucifera (Hyde et al. 2016), Liquidambar formosana (Fan et al. 2022) | China (Fan et al. 2022), Thailand (Hyde et al. 2016) | |
| N. coffeae-arabicae | Coffea arabica (Song et al. 2013) | China (Song et al. 2013) | |
| N. collariata | Diospyros kaki, Rhododendron sp. (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. concentrica | Rhapis excelsa (Zhang et al. 2024d), Rosa chinensis and Rosa rugosa (Peng et al. 2022) | China (Peng et al. 2022; Zhang et al. 2024d) | |
| N. cubana | Camellia oleifera (Li et al. 2021b), Careya arborea (Reddy et al. 2016), Cordia dichotoma (Reddy et al. 2016), Hevea brasiliensis (Pornsuriya et al. 2020), Leaf litter (Maharachchikumbura et al. 2014b), Psidium guajava (Huang et al. 2024), Stereospermum tetragonum (Reddy et al. 2016), | China (Li et al. 2021b), Cuba (Maharachchikumbura et al. 2014b), India (Reddy et al. 2016), Thailand (Pornsuriya et al. 2020) | |
| N. dendrobii | Dendrobium cariniferum (Ma et al. 2019), Leptolobium dsycarpum and Qualea parviflora (dos Reis et al. 2023) | Brazil (dos Reis et al. 2023), Thailand (Ma et al. 2019) | |
| N. dimorphospora | Eurya chinensis, Patrinia villosa (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. dolichoconidiophora | Aucuba japonica, Cycas revoluta (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. drenthii | Macadamia integrifolia (Prasannath et al. 2021) | Australia (Prasannath et al. 2021) | |
| N. egyptiaca | Magnolia candolli (de Silva et al. 2021), Mangifera indica (Crous et al. 2015), Psidium guajava (Bezerra et al. 2018), Rhizophora mucronate (Hamzah et al. 2018) | Brazil (Bezerra et al. 2018), China (de Silva et al. 2021), Egypt (Crous et al. 2015), Malaysia (Hamzah et al. 2018) | |
| N. elaeagni | Elaeagnus pungens (He et al. 2022) | China (He et al. 2022) | |
| N. elaeidis | Elaeis guineensis and Unidentified palm (Konta et al. 2023) | Thailand (Konta et al. 2023) | |
| N. ellipsospora | Acanthopanax divaricatus (Yun et al. 2015), Ardisia crenata (Maharachchikumbura et al. 2014b), Camellia sinensis (Wang et al. 2019b), Diospyros kaki (Qin et al. 2022), Diospyros montana (Reddy et al. 2016), Ipomoea batatas (Maharachchikumbura et al. 2016a), Unidentified tree (Maharachchikumbura et al. 2012) | China (Wang et al. 2019b), India (Reddy et al. 2016), Korea (Yun et al. 2015), Thailand (Maharachchikumbura et al. 2012) | |
| N. eucalypticola | Eucalyptus globulus (Maharachchikumbura et al. 2014b) | Unknown (Maharachchikumbura et al. 2014b) | |
| N. eucalyptorum | Eucalyptus globulus (Diogo et al. 2021) | Portugal (Diogo et al. 2021) | |
| N. fijiensis | Arachis hypogaea (Razaghi et al. 2024) | Fiji (Razaghi et al. 2024) | |
| N. fimbriata | Camellia sp., Cinnamomum camphora (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. foedans | Calliandra haematocephala (Maharachchikumbura et al. 2012), Cocos nucifera (Barbosa et al. 2023), Diospyros kaki (Watanabe et al. 2012), Egg mass of the sailfin sandfish (Park et al. 2018), Elaeis guineensis (Mohamed-Azni et al. 2022), Mangrove (Maharachchikumbura et al. 2012), Nectandra lineatifolia (Nelson et al. 2020), Neodypsis decaryi (Maharachchikumbura et al. 2012), Persea americana (Shetty et al. 2016), Podocarpus macrophyllus (Wei et al. 2007), Rhizophora mucronata (Hamzah et al. 2018), Thuja occidentalis (Maharachchikumbura et al. 2012), Vitis vinifera (Jayawardena et al. 2015) | Brazil (Barbosa et al. 2023), China (Maharachchikumbura et al. 2012), Ecuador (Nelson et al. 2020), Japan (Watanabe et al. 2012), Korea (Park et al. 2018), Malaysia (Mohamed-Azni et al. 2022), USA (Shetty et al. 2016) | |
| N. formicidarum | Amentotaxus yunnanensis (Vu et al. 2024), Ananas comosus (Tian et al. 2024), Calamus castaneus (Azuddin et al. 2021), Dead ant (Maharachchikumbura et al. 2014b), Dypsis leptocheilos (Xiong et al. 2022), Hevea brasiliensis (Pornsuriya et al. 2020), Jabuticaba (Lin et al. 2022), Kadsura coccinea (Zhonghou et al. 2023), Paullinia cupana (Gualberto et al. 2021), Photinia serratifolia (Sun et al. 2023), Plant debris (Maharachchikumbura et al. 2014b), Styrax sp. (Elfiati et al. 2022), Vitis vinifera (Kenfaoui et al. 2024) | Brazil (Gualberto et al. 2021), China (Xiong et al. 2022), Cuba (Maharachchikumbura et al. 2014b), Ghana (Maharachchikumbura et al. 2014b), Indonesia (Elfiati et al. 2022), Malaysia (Azuddin et al. 2021), Morocco (Kenfaoui et al. 2024), Thailand (Tian et al. 2024), Vietnam (Vu et al. 2024) | |
| N. chrysea | N. fragariae | Fragaria × ananassa (Prematunga et al. 2022) | China (Prematunga et al. 2022) |
| N. fructicola | Rosa roxburghii (Zhang et al. 2024b) | China (Zhang et al. 2024b) | |
| N. fuzhouensis | Acer buergerianum (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. guajavae | Psidium guajava (Haq et al. 2021) | Pakistan (Haq et al. 2021) | |
| N. guajavicola | Ananas comosus (Tian et al. 2024), Cocos nucifera (Tian et al. 2024), Nephrolepis sp. (Seifollahi et al. 2023), Psidium guajava (Haq et al. 2021) | Pakistan (Haq et al. 2021), Thailand (Tian et al. 2024) | |
| N. guangxiensis | Poaceae (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. guizhouensis | Air, unnamed karst cave (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. haikouensis | Ilex chinensis (Zhang et al. 2022), Ophiocordyceps sp. and Tolypocladium sp. (Hsu et al. 2024) | China (Zhang et al. 2022), Taiwan, China (Hsu et al. 2024) | |
| N. hispanica | Eucalyptus globulus (Diogo et al. 2021), Luma apiculata and Myrtus communis (Aiello et al. 2024) | Italy (Aiello et al. 2024), Portugal and Spain (Diogo et al. 2021) | |
| N. hispanica | N. vaccinii | Vaccinium corymbosum (Santos et al. 2022; Blagojević et al. 2024) | Portugal (Santos et al. 2022), Serbia (Blagojević et al. 2024) |
| N. honoluluana | Telopea sp. (Maharachchikumbura et al. 2014b) | America (Maharachchikumbura et al. 2014b) | |
| N. honoluluana | N. hadrolaeliae | Hadrolaelia jongheana (Freitas et al. 2019) | Brazil (Freitas et al. 2019) |
| N. honoluluana | N. zimbabwana | Leucospermum cunciforme (Maharachchikumbura et al. 2014b), Xylaria sp. (Hermawan et al. 2021) | Zimbabwe (Maharachchikumbura et al. 2014b) |
| N. hydeana | Alpinia malaccensis and Artocarpus heterophyllus (Huanaluek et al. 2021), Cyclosorus sp. (Seifollahi et al. 2023) | Thailand (Huanaluek et al. 2021) | |
| N. hyperici | Hypericum monogynum (Sun et al. 2023), Musa basjoo (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. iberica | Bamboo (Kwon et al. 2024), Camellia oleifera (Li et al. 2021b), Eucalyptus globulus (Li et al. 2021b), Myrtus communis (Aiello et al. 2024) | China (Li et al. 2021b), Italy (Aiello et al. 2024), Korea (Kwon et al. 2024), Portugal and Spain (Li et al. 2021b) | |
| N. iranensis | Fragaria × ananassa (Ayoubi and Soleimani 2016) | Iran (Ayoubi and Soleimani 2016) | |
| N. javaensis | Cocos nucifera (Maharachchikumbura et al. 2014b), stone fruit trees (Singha et al. 2024), Vitis vinifera (Maharachchikumbura et al. 2016b) | France (Maharachchikumbura et al. 2016b), Indonesia (Maharachchikumbura et al. 2014b), Korea (Ahn et al. 2016) and South Africa (Singha et al. 2024) | |
| N. jiangxiensis | Rhododendron latoucheae (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. keteleeriae | Keteleeria pubescens (Song et al. 2014a), Magnolia candolli (de Silva et al. 2021) | China (Song et al. 2014a; de Silva et al. 2021) | |
| N. keteleeriae | N. rhapidis | Podocarpus macrophyllus (Sun et al. 2023), Rhapis excelsa (Yang et al. 2021) | China (Yang et al. 2021; Sun et al. 2023) |
| N. liquidambaris | Liquidambar formosana (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. longiappendiculata | Bamboo (Kwon et al. 2024), Camellia oleifera, Camellia sasanqua (Wang et al. 2022), Eucalyptus globulus, Eucalyptus nitens (Diogo et al. 2021) | China (Wang et al. 2022), Korea (Kwon et al. 2024) and Portugal (Diogo et al. 2021) | |
| N. lusitanica | Eucalyptus globulus (Diogo et al. 2021) | Portugal (Diogo et al. 2021) | |
| N. macadamiae | Macadamia integrifolia (Akinsanmi et al. 2017), Macadamia sp. (Prasannath et al. 2023) | Australia (Akinsanmi et al. 2017; Prasannath et al. 2023) | |
| N. machili | Machilus yunnanensis (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. maddoxii | Macadamia integrifolia (Prasannath et al. 2021) | Australia (Prasannath et al. 2021) | |
| N. magna | Pteridium sp. (Maharachchikumbura et al. 2014a) | France (Maharachchikumbura et al. 2014a) | |
| N. megabetaspora | Brachiaria sp., Poaceae (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. mesopotamica | Achras sapota (Maharachchikumbura et al. 2014b), Bridelia retusa, Cordia dichotoma (Reddy et al. 2016), Eucalyptus sp. (Maharachchikumbura et al. 2014b), Fragaria × ananassa (Ayoubi and Soleimani 2016; Hidrobo-Chavez et al. 2022), Pinus brutia (Maharachchikumbura et al. 2014b) | Ecuador (Hidrobo-Chavez et al. 2022), India (Maharachchikumbura et al. 2014b; Reddy et al. 2016), Iran (Ayoubi and Soleimani 2016), Iraq (Maharachchikumbura et al. 2014b) and Turkey (Maharachchikumbura et al. 2014b) | |
| N. mianyangensis | Paeonia suffruticosa (Li et al. 2022) | China (Li et al. 2022) | |
| N. moniliformis | Phyllostachys sp. (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. musae | Asplenium nidus (Seifollahi et al. 2023), Musa sp. (Hyde et al. 2016) | Thailand (Hyde et al. 2016; Seifollahi et al. 2023) | |
| N. nanningensis | Ixora chinensis (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. natalensis | Acacia mollissima (Maharachchikumbura et al. 2014b), Camellia sinensis (Pandey et al. 2023) | South Africa (Maharachchikumbura et al. 2014b), India (Pandey et al. 2023) | |
| N. nebuloides | Sporobolus elongatus (Crous et al. 2020), Sporobolus indicus (Steinrucken et al. 2022) | Australia (Crous et al. 2020; Steinrucken et al. 2022) | |
| N. neomaricae | Neomarica longifolia (Hyde et al. 2024) | Thailand (Hyde et al. 2024) | |
| N. oblatespora | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) | |
| N. zingiberis | N. olivaceous | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) |
| N. olumideae | Macadamia integrifolia (Prasannath et al. 2021) | Australia (Prasannath et al. 2021) | |
| N. oxyphylla | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) | |
| N. paeoniae | Paeoniae (Liu et al. 2019) | Unknown (Liu et al. 2019) | |
| N. paeoniae-suffruticosae | Paeonia suffruticosa (Li et al. 2022) | China (Li et al. 2022) | |
| N. pandanicola | Cyclosorus sp. (Seifollahi et al. 2023), Pandanus sp. (Tibpromma et al. 2018b) | Thailand (Seifollahi et al. 2023), China (Tibpromma et al. 2018b) | |
| N. pernambucana | Vismia guianensis (Silvério et al. 2016) | Brazil (Silvério et al. 2016) | |
| N. perukae | Psidium guajava (Haq et al. 2021) | Pakistan (Haq et al. 2021) | |
| N. petila | Rhizophora mucronate (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| N. phangngaensis | Cyclosorus sp. (Seifollahi et al. 2023), Ficus septica (Tennakoon et al. 2021), Pandanus sp. (Tibpromma et al. 2018b) | Taiwan, China (Tennakoon et al. 2021), Thailand (Tibpromma et al. 2018b; Seifollahi et al. 2023) | |
| N. phoenicis | Phoenix canariensis (Zhang et al. 2024d) | China (Zhang et al. 2024d) | |
| N. photiniae | Dendrobium sp. (Xing et al. 2011), Photinia serratifolia, Photinia sp. (Sun et al. 2023) | China (Sun et al. 2023), Vietnam (Xing et al. 2011) | |
| N. phyllostachydis | Castanopsis sp., Phyllostachys sp., Rhododendron arboreum (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. piceana | Ananus comosus (Bhattacharya et al. 2019), Cinnamomum malabatrum (Reddy et al. 2016), Cocos nucifera (Maharachchikumbura et al. 2014b), Homo sapiens (Borgohain et al. 2020), Mangifera indica, Picea sp. (Maharachchikumbura et al. 2014b) | England (Maharachchikumbura et al. 2014b), India (Bhattacharya et al. 2019; Borgohain et al. 2020), Indonesia (Maharachchikumbura et al. 2014b), Unknown country (Maharachchikumbura et al. 2014b) | |
| N. poae | Poaceae (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. protearum | Bulbophyllum thouars (Ran et al. 2017), Camellia oleifera (Tang et al. 2021), Eucalyptus (Belisário et al. 2020), Leucospermum cuneiforme (Crous et al. 2011), Nopalea cochenillifera (Conforto et al. 2019), Qualea parviflora (dos Reis et al. 2023), Rhizophora mucronate (Hamzah et al. 2018) | Brazil (Conforto et al. 2019; Belisário et al. 2020; dos Reis et al. 2023), China (Ran et al. 2017; Tang et al. 2021), Malaysia (Hamzah et al. 2018), Zimbabwe (Crous et al. 2011) | |
| N. psidii | Nephrolepis cordifolia (Seifollahi et al. 2023), Psidium guajava (Haq et al. 2021) | Pakistan (Haq et al. 2021), Thailand (Seifollahi et al. 2023) | |
| N. rhizophorae | Mangifera indica (Adikaram et al. 2023), Rhizophora mucronate (Norphanphoun et al. 2019) | Sri Lanka (Adikaram et al. 2023), Thailand (Norphanphoun et al. 2019) | |
| N. rhododendri | Dracaena fragrans (Sun et al. 2023), Rhododendron simsii (Yang et al. 2021) | China (Yang et al. 2021), Thailand (Sun et al. 2023) | |
| N. rhododendricola | Rhododendron sp. (Chaiwan et al. 2022) | China (Chaiwan et al. 2022) | |
| N. rosae | Callistemon viminalis (Aiello et al. 2024), Citrus reticulata (Ma et al. 2023b), Egg mass of the sailfin sandfish (Park et al. 2018), Eucalyptus sp. (Belisário et al. 2020; Santos et al. 2020), Fragaria × ananassa (Sun et al. 2021; Lawrence et al. 2023), Fraxinus excelsior (Langer et al. 2024), Luma apiculata, Myrtus communis (Aiello et al. 2024), Paeonia suffruticosa (Maharachchikumbura et al. 2014b), Parmotrema praesorediosum (Oh et al. 2020), Persea americana (Fiorenza et al. 2022), Punica granatum (Xavier et al. 2021), Rosa sp. (Maharachchikumbura et al. 2014b), Vaccinium corymbosum (Rodríguez-Gálvez et al. 2020; Santos et al. 2022; Lamderos Gálvez et al. 2024) | America (Maharachchikumbura et al. 2014b; Lawrence et al. 2023), Brazil (Belisário et al. 2020; Santos et al. 2020), China (Sun et al. 2021; Hsu et al. 2022; Ma et al. 2023b), Egypt (Essa et al. 2018), Germany (Langer et al. 2024), Italy (Fiorenza et al. 2022; Aiello et al. 2024), Korea (Park et al. 2018; Oh et al. 2020), Mexico (Rebollar-Alviter et al. 2020; Lamderos Gálvez et al. 2024), New Zealand (Maharachchikumbura et al. 2014b), Paraguay(Fernández-Ozuna et al. 2023), Peru (Rodríguez-Gálvez et al. 2020), Portugal (Santos et al. 2022), Russia(Tsvetkova and Kuznetsova 2022), Serbia (Santos et al. 2022), Turkey (Erdurmuş et al. 2023) | |
| N. rosicola | Myrtus communis (Aiello et al. 2024), Rosa chinensis (Jiang et al. 2018) | China (Jiang et al. 2018), Italy (Aiello et al. 2024) | |
| N. samarangensis | Syzygium samarangense (Maharachchikumbura et al. 2013b), Unidentified tree (Maharachchikumbura et al. 2014b) | China (Maharachchikumbura et al. 2014b), Thailand (Maharachchikumbura et al. 2013b) | |
| N. saprophytica | Calamus castaneus (Azuddin et al. 2021), Camellia sinensis (Koodalugodaarachchi et al. 2024), Cocos nucifera (Tian et al. 2024), Cordia dichotoma (Reddy et al. 2016), Cyclosorus sp. (Seifollahi et al. 2023), Diospyros kaki (Qin et al. 2023), Elaeis guineensis (Ismail et al. 2017), Erythropalum scandens (Yang et al. 2021), Litsea rotundifolia (Maharachchikumbura et al. 2012; Maharachchikumbura et al. 2014a), Magnolia sp. (Maharachchikumbura et al. 2012), Mangifera indica (Shu et al. 2020), Paphiopedilum micranthum (Qin et al. 2020), Roystonea regia (Ismail et al. 2022), Vitis vinifera (Maharachchikumbura et al. 2016b) | China (Yang et al. 2021; Qin et al. 2023), France (Maharachchikumbura et al. 2016b), India (Reddy et al. 2016), Malaysia (Ismail et al. 2017; Azuddin et al. 2021), Thailand (Seifollahi et al. 2023; Koodalugodaarachchi et al. 2024; Tian et al. 2024) | |
| N. scalabiensis | Vaccinium corymbosum (Santos et al. 2022) | Portugal (Santos et al. 2022) | |
| N. sichuanensis | Castanea mollissima (Jiang et al. 2021) | China (Jiang et al. 2021) | |
| N. siciliana | Persea americana (Fiorenza et al. 2022) | Italy (Fiorenza et al. 2022) | |
| N. smilacis | Smilax lanceifolia (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| N. sonneratiae | Sonneronata alba (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| N. steyaertii | Eucalyptus viminalis (Maharachchikumbura et al. 2014a) | Australia (Maharachchikumbura et al. 2014a) | |
| N. subepidermalis | Rosa chinensis, Rosa rugosa (Peng et al. 2022) | China (Peng et al. 2022) | |
| N. suphanburiensis | Unidentified host (Sun et al. 2023) | Thailand (Sun et al. 2023) | |
| N. surinamensis | Egg mass of the sailfin sandfish (Park et al. 2018), Oil palm hybrid (Betancourt-Ortiz et al. 2024), Protea eximia, Soil under Elaeis guineensis (Maharachchikumbura et al. 2014b), Vellozia gigantea (Ferreira et al. 2017) | Brazil (Ferreira et al. 2017), Colombia (Betancourt-Ortiz et al. 2024), Korea (Park et al. 2018), Suriname, Zimbabwe (Maharachchikumbura et al. 2014b) | |
| N. termitarii | Termite nest in the tea agro-eco-system (Kamil et al. 2012) | India (Kamil et al. 2012) | |
| N. terricola | Cocos nucifera (Tian et al. 2024), Olea europaea, Paeonia suffruticosa (Li et al. 2022) | China (Li et al. 2022), Thailand (Tian et al. 2024) | |
| N. thailandica | Mangrove (Sahibu et al. 2024), Rhizophora mucronate (Norphanphoun et al. 2019) | Malaysia (Sahibu et al. 2024), Thailand (Norphanphoun et al. 2019) | |
| N. vacciniicola | Vaccinium corymbosum (Santos et al. 2022) | Portugal (Santos et al. 2022) | |
| N. vheenae | Macadamia integrifolia (Prasannath et al. 2021) | Australia (Prasannath et al. 2021) | |
| N. vitis | Manilkara zapota (Nuthan et al. 2021), Vitis vinifera (Jayawardena et al. 2016; Kenfaoui et al. 2024) | China (Jayawardena et al. 2016; Jayawardena et al. 2018), India (Nuthan et al. 2021), Morocco (Kenfaoui et al. 2024) | |
| N. wuzhishanensis | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) | |
| N. xishuangbannaensis | Kerivoula hardwickii (Liu et al. 2023) | China (Liu et al. 2023) | |
| N. yongxunensis | Alpinia oxyphylla (Cui et al. 2024) | China (Cui et al. 2024) | |
| N. zakeelii | Callistemon viminali (Aiello et al. 2024), Macadamia integrifolia (Prasannath et al. 2021), Myrtus communis (Aiello et al. 2024) | Australia (Prasannath et al. 2021), Italy (Aiello et al. 2024) | |
| N. zingiberis | Zingiber officinale (He et al. 2022) | China (He et al. 2022) | |
| Pe. abietis | Abies fargesii (Gu et al. 2021) | China (Gu et al. 2021) | |
| Pe. adusta | Clerodendrum canescens (Xu et al. 2016), Cocos nucifera (Rosado et al. 2015; Tian et al. 2024), Dictyosperma album (Zhu et al. 2015), Ericaceae (Kohout and Tedersoo 2017), Hevea brasiliensis (de Oliveira Amaral et al. 2022), Immature coconut (Rosado et al. 2015), Mangifera indica (Shu et al. 2020; Adikaram et al. 2023), Nectandra lineatifolia (Nelson et al. 2020), Podocarpus macrophyllus (Wei et al. 2007), Prunus cerasus (Maharachchikumbura et al. 2012), Refrigerator door PVC gasket (Maharachchikumbura et al. 2012), Rhizophora mucronate (Apurillo et al. 2019), Rubus idaeus (Yan et al. 2019), Sinopodophyllum hexandrum (Xiao et al. 2017), Smilax nipponica (Watanabe et al. 2010), Syagrus oleracea (Cardoso et al. 2017), Syzygium sp. (Maharachchikumbura et al. 2012), Vaccinium corymbosum (Zheng et al. 2023b) | America (Maharachchikumbura et al. 2012), Brazil (Rosado et al. 2015; Cardoso et al. 2017; de Oliveira Amaral et al. 2022), China (Zhu et al. 2015; Xu et al. 2016; Shu et al. 2020), Ecuador (Nelson et al. 2020), Fiji (Maharachchikumbura et al. 2012), Japan (Watanabe et al. 2010), Philippines (Apurillo et al. 2019), South Africa (Kohout and Tedersoo 2017), Sri Lanka (Adikaram et al. 2023), Thailand (Maharachchikumbura et al. 2012; Tian et al. 2024) | |
| Pe. adusta | Pe. papuana | Celtis formosana (Tennakoon et al. 2021), Cocos nucifera (Maharachchikumbura et al. 2014b), Oryza sp. (Pak et al. 2017), Palm (Zhang et al. 2024c), Soil along the coast (Maharachchikumbura et al. 2014b), Unknown grass species (Pak et al. 2017) | Australia (Pak et al. 2017), China (Tennakoon et al. 2021), Papua New Guinea (Maharachchikumbura et al. 2014b), Thailand (Zhang et al. 2024c) |
| Pe. aggestorum | Camellia sinensis (Liu et al. 2017) | China (Liu et al. 2017) | |
| Pe. alloschemones | Alloschemone occidentalis (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. americana | Leucospermum cunei × conocarpodendron (Razaghi et al. 2024) | America (Razaghi et al. 2024) | |
| Pe. anacardiacearum | Mangifera indica (Maharachchikumbura et al. 2013c, Shu et al. 2020) | China (Maharachchikumbura et al. 2013c, Shu et al. 2020) | |
| Pe. anhuiensis | Cyclobalanopsis glauca (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. aporosae-dioicae | Aporosa dioica (Yin et al. 2024) | China (Yin et al. 2024) | |
| Pe. arceuthobii | Arceuthobium campylopodum (Maharachchikumbura et al. 2014b) | America (Maharachchikumbura et al. 2014b) | |
| Pe. arengae | Arenga undulatifolia (Maharachchikumbura et al. 2014b), Oil palm hybrid (Betancourt-Ortiz et al. 2024) | Colombia (Betancourt-Ortiz et al. 2024), Singapore (Maharachchikumbura et al. 2014b) | |
| Pe. australasiae | Knightia sp., Protea neriifolia × susannae cv. Pink Ice (Maharachchikumbura et al. 2014b), Sabal maritima (Zhang et al. 2024c), Vellozia gigantea (Ferreira et al. 2017) | Australia (Maharachchikumbura et al. 2014b), Brazil (Ferreira et al. 2017), China (Zhang et al. 2024c), New Zealand (Maharachchikumbura et al. 2014b) | |
| Pe. australis | Grevillea sp. (Maharachchikumbura et al. 2014b), Pinus pinea (Silva et al. 2020), Protea neriifolia × susannae cv. Pink Ice (Maharachchikumbura et al. 2014b), Vaccinium corymbosum (Santos et al. 2022) | Australia (Maharachchikumbura et al. 2014b), Portugal (Silva et al. 2020; Santos et al. 2022), South Africa (Maharachchikumbura et al. 2014b) | |
| Pe. biappendiculata | Rhododendron sp. (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. biciliata | Egg mass of the sailfin sandfish (Park et al. 2018), Eucalyptus sp. (Morales‐Rodríguez et al. 2019), Myrtus communis (Aiello et al. 2024), Paeonia sp. (Maharachchikumbura et al. 2014b), Parkinsonia aculeata (Steinrucken et al. 2017), Pinus pinea (Silva et al. 2020), Pistacia lentiscus (Hlaiem et al. 2022), Platanus × hispanica (Maharachchikumbura et al. 2014b), Quercus coccifera (Hlaiem et al. 2022), Shorea macrophylla (Lateef et al. 2018), Taxus baccata (Maharachchikumbura et al. 2014b), Vaccinium corymbosum (Santos et al. 2022), Vitis vinifera (Maharachchikumbura et al. 2016b; Lorenzini and Zapparoli 2018) | Australia (Steinrucken et al. 2017), France (Maharachchikumbura et al. 2016b), Italy (Maharachchikumbura et al. 2014b; Lorenzini and Zapparoli 2018; Morales‐Rodríguez et al. 2019; Aiello et al. 2024), Korea (Park et al. 2018), Malaysia (Lateef et al. 2018), Netherlands (Maharachchikumbura et al. 2014b), Portugal (Silva et al. 2020; Santos et al. 2022), Slovakia (Maharachchikumbura et al. 2014b), Tunisia (Hlaiem et al. 2022) | |
| Pe. brachiata | Camellia sp. (Liu et al. 2017) | China (Liu et al. 2017) | |
| Pe. brassicae | Brassica napus (Maharachchikumbura et al. 2014b) | New Zealand (Maharachchikumbura et al. 2014b) | |
| Pe. brassicae | Pe. hollandica | Cupressus sempervirens (Crous et al. 2018), Pinus pinea (Silva et al. 2020), Sciadopitys verticillate (Maharachchikumbura et al. 2014b) | Netherlands (Maharachchikumbura et al. 2014b), Portugal (Silva et al. 2020), Spain (Crous et al. 2018) |
| Pe. camelliae | Camellia japonica (Zhang et al. 2012b), Camellia sinensis (Chen et al. 2018c; Win et al. 2018; Tsai et al. 2021; Win et al. 2021) | China (Zhang et al. 2012b; Chen et al. 2018c; Tsai et al. 2021), Japan (Win et al. 2018; Win et al. 2021), Turkey (Vu et al. 2019) | |
| Pe. camelliae | Pe. yanglingensis | Camellia sinensis (Liu et al. 2017; Tsai et al. 2021) | China (Liu et al. 2017; Tsai et al. 2021) |
| Pe. camelliae-japonicae | Camellia japonica (Dong et al. 2023) | China (Dong et al. 2023) | |
| Pe. camelliae-oleiferae | Camellia oleifera (Li et al. 2021b) | China (Li et al. 2021b) | |
| Pe. cangshanensis | Rhododendron delavayi (Gu et al. 2022) | China (Gu et al. 2022) | |
| Pe. castanopsidis | Castanopsis hystrix, Castanopsis lamontii (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. changjiangensis | Castanopsis hainanensis, Castanopsis tonkinensis, Cyclobalanopsis austrocochinchinensis (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. chaoyangensis | Euonymus japonicus (Lin et al. 2023) | China (Lin et al. 2023) | |
| Pe. chiangmaiensis | Phyllostachys edulis (Sun et al. 2023) | Thailand (Sun et al. 2023) | |
| Pe. chiaroscuro | Sporobolus indicus (Steinrucken et al. 2022), Sporobolus natalensis (Crous et al. 2022) | Australia (Crous et al. 2022; Steinrucken et al. 2022) | |
| Pe. chinensis | Camellia sinensis (Liu et al. 2017), Taxus sp. (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012; Liu et al. 2017) | |
| Pe. clavata | Buxus sp., Euonymus sp. (Maharachchikumbura et al. 2012), Rhododendron delavayi (Zhang et al. 2013b), Soil (Lim et al. 2022) | China (Maharachchikumbura et al. 2012; Zhang et al. 2013b), Korea (Lim et al. 2022) | |
| Pe. lushanensis | Camellia sinensis (Chen et al. 2018b; Manawasinghe et al. 2021), Camellia sp. (Liu et al. 2017), Dendrobium sp. (Ma et al. 2019), Podocarpus macrophyllus (Zheng et al. 2022), Quercus serrata (Jiang et al. 2022), Sarcandra glabra (Zhang et al. 2021a) | China (Chen et al. 2018b; Manawasinghe et al. 2021; Zhang et al. 2021a; Jiang et al. 2022; Zheng et al. 2022), Thailand (Ma et al. 2019) | |
| Pe. pini | Pinus oocarpa (Ortiz et al. 2022), Pinus pinaster, Pinus pinea (Silva et al. 2020) | Honduras (Ortiz et al. 2022), Portugal (Silva et al. 2020) | |
| Pe. lushanensis | Pe. iberica | Pinus radiata, Pinus sylvestris (Monteiro et al. 2022) | Spain (Monteiro et al. 2022) |
| Pe. rhododendri | Egg mass of the sailfin sandfish (Park et al. 2018), Rhododendron sinogrande (Zhang et al. 2013b) | China (Zhang et al. 2013b), Korea (Park et al. 2018) | |
| Pe. colombiensis | Eucalyptus eurograndis (Maharachchikumbura et al. 2014b), Oryza australiensis (Pak et al. 2017) | Australia (Pak et al. 2017), Colombia (Maharachchikumbura et al. 2014b) | |
| Pe. cyclobalanopsidis | Cyclobalanopsis glauca (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. dianellae | Dianella sp. (Crous et al. 2017) | Australia (Crous et al. 2017) | |
| Pe. digitalis | Digitalis purpurea (Liu et al. 2015) | New Zealand (Liu et al. 2015) | |
| Pe. dilucida | Camellia sinensis (Liu et al. 2017) | China (Liu et al. 2017) | |
| Pe. diploclisiae | Algae (Bunyapaiboonsri et al. 2023), Areca triandra (Xiong et al. 2022), Butia sp. (Senanayake et al. 2020), Diploclisia glaucescens, Psychotria tutcheri (Maharachchikumbura et al. 2014b), Sabal mexicana (Xiong et al. 2022), Syzygium sp. (Lateef et al. 2018) | China (Maharachchikumbura et al. 2014b; Senanayake et al. 2020; Xiong et al. 2022), Malaysia (Lateef et al. 2018), Thailand (Bunyapaiboonsri et al. 2023) | |
| Pe. humicola | Egg mass of the sailfin sandfish (Park et al. 2018), Ilex cinerea (Maharachchikumbura et al. 2014b), Mangrove (Li et al. 2016), Seaweed (Park et al. 2021), Soil in tropical rain forest (Maharachchikumbura et al. 2014b) | China (Maharachchikumbura et al. 2014b; Li et al. 2016), Korea (Park et al. 2018), Micronesia (Park et al. 2021), Papua New Guinea (Maharachchikumbura et al. 2014b) | |
| Pe. spatholobi | Pe. pyrrosiae-linguae | Pyrrosia lingua (Dong et al. 2023) | China (Dong et al. 2023) |
| Pe. spatholobi | Spatholobus suberectus (Zhang et al. 2023) | China (Zhang et al. 2023) | |
| Pe. disseminata | Dead hardwood branch (Deyrup et al. 2006), Espeletia sp. (Miles et al. 2012), Eucalyptus botryoides (Crous et al. 2006; Liu et al. 2019), Eucalyptus sp. (Liu et al. 2019), Ficus macrocarpa (Ding et al. 2015), Pieris japonica (Watanabe et al. 2012), Pinus armandii (Hu et al. 2007), Pinus pentaphylla (Watanabe et al. 2010), Pinus pinea (Silva et al. 2020), Podocarpus macrophyllus (Wei et al. 2007), Psidium guajava (Keith et al. 2006), Shorea macrophylla (Lateef et al. 2018) | America (Deyrup et al. 2006; Keith et al. 2006), Brazil (Paulino et al. 2019), China (Hu et al. 2007; Wei et al. 2007), Colombia (Miles et al. 2012), Hong Kong (Ding et al. 2015), Japan (Watanabe et al. 2010; Watanabe et al. 2012), Malaysia (Lateef et al. 2018), New Zealand (Crous et al. 2006; Liu et al. 2019), North Island (Liu et al. 2019), Portugal (Silva et al. 2020) | |
| Pe. diversiseta | Rhododendron sp. (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012) | |
| Pe. doitungensis | Dendrobium sp. (Ma et al. 2019) | Thailand (Ma et al. 2019) | |
| Pe. dracaenae | Celtis formosana (Tennakoon et al. 2021), Dracaena fragrans (Ariyawansa et al. 2015) | China (Ariyawansa et al. 2015; Tennakoon et al. 2021) | |
| Pe. dracaenicola | Dracaena sp. (Chaiwan et al. 2020) | Thailand (Chaiwan et al. 2020) | |
| Pe. dracontomelonis | Dipterocarpus alatus (Samaradiwakara et al. 2023), Dracontomelon dao (Liu et al. 2015), Nephrolepis cordifolia (Seifollahi et al. 2023), Podocarpus sp. (Sun et al. 2023) | Thailand (Liu et al. 2015; Samaradiwakara et al. 2023; Seifollahi et al. 2023; Sun et al. 2023) | |
| Pe. eleutherococci | Eleutherococcus brachypus (Tian et al. 2022) | China (Tian et al. 2022) | |
| Pe. endophytica | Magnolia garrettii (de Silva et al. 2021) | Thailand (de Silva et al. 2021) | |
| Pe. ericacearum | Rhododendron delavayi (Zhang et al. 2013b) | China (Zhang et al. 2013b) | |
| Pe. etonensis | Sporobolus indicus (Steinrucken et al. 2022), Sporobolus jacquemontii (Crous et al. 2020) | Australia (Crous et al. 2020; Steinrucken et al. 2022) | |
| Pe. exudata | Aucuba japonica (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. ficicola | Ficus macrocarpa (Zhang et al. 2023) | China (Zhang et al. 2023) | |
| Pe. ficicrescens | Cleyera japonica (Razaghi et al. 2024), Ficus tikoua (Hyde et al. 2023), Oleaceae (Razaghi et al. 2024) | China (Hyde et al. 2023; Razaghi et al. 2024) | |
| Pe. ficicrescens | Pe. ganzhouensis | Cinnamomum camphora (Razaghi et al. 2024) | China (Razaghi et al. 2024) |
| Pe. foliicola | Castanopsis faberi (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. formosana | Dead grass (Ariyawansa and Hyde 2018), Grass (Tennakoon et al. 2021), Ophiocordyceps sp. (Hsu et al. 2024) | China (Ariyawansa and Hyde 2018; Tennakoon et al. 2021; Hsu et al. 2024) | |
| Pe. furcata | Camellia sinensis (Maharachchikumbura et al. 2013a; Liu et al. 2017) | China (Liu et al. 2017), Thailand (Maharachchikumbura et al. 2013a) | |
| Pe. fusiformis | Rhododendron sp. (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. fusoidea | Rhododendron delavayi (Gu et al. 2022) | China (Gu et al. 2022) | |
| Pe. gibbosa | Gaultheria forrestii (Zhang et al. 2013b), Gaultheria shallon (Watanabe et al. 2018) | Canada (Watanabe et al. 2018), China (Zhang et al. 2013b) | |
| Pe. grevilleae | Grevillea sp. (Maharachchikumbura et al. 2014b) | Australia (Maharachchikumbura et al. 2014b) | |
| Pe. guangdongensis | Arenga pinnata (Xiong et al. 2022) | China (Xiong et al. 2022) | |
| Pe. guangxiensis | Quercus griffithii (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. guiyangensis | Eriobotrya japonica (Zhang et al. 2024d) | China (Zhang et al. 2024d) | |
| Pe. guizhouensis | Cyclobalanopsis glauca (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. hainanensis | Artemisia argyi (Qian et al. 2014), Podocarpus macrophyllus (Liu et al. 2007) | China (Liu et al. 2007; Qian et al. 2014) | |
| Pe. hainanensis | Pe. appendiculata | Rhododendron decorum (Gu et al. 2022) | China (Gu et al. 2022) |
| Pe. hainanensis | Pe. chamaeropis | Camellia sp. (Liu et al. 2017), Castanopsis fissa (Jiang et al. 2022), Chamaerops humilis (Maharachchikumbura et al. 2014b), Decaying woody (Adams and Walker 2023), Erica arborea (Hlaiem et al. 2018), Eurya nitida (Qiu et al. 2022), Ophiocordyceps sp. (Hsu et al. 2024), Peristrophe japonica (Sun et al. 2023), Pieris japonica sub sp. Japonica (Nozawa et al. 2019), Prostanthera rotundifolia (Moslemi and Taylor 2015), Quercus acutissima, Quercus aliena, Quercus variabilis (Jiang et al. 2022), Rosa chinensis (Peng et al. 2022), Vaccinium corymbosum (Santos et al. 2022; Zheng et al. 2023b), Vandopsis gigantea (Ran et al. 2017), Vitis vinifera (Jayawardena et al. 2018) | Australia (Moslemi and Taylor 2015), Canada (Adams and Walker 2023), China (Liu et al. 2017; Jiang et al. 2022; Peng et al. 2022; Qiu et al. 2022; Sun et al. 2023; Zheng et al. 2023b; Hsu et al. 2024), Italy (Maharachchikumbura et al. 2014b; Jayawardena et al. 2018; Vu et al. 2019), Japan (Nozawa et al. 2019), Portugal (Santos et al. 2022), Tunisia (Hlaiem et al. 2018) |
| Pe. hainanensis | Pe. daliensis | Rhododendron decorum (Gu et al. 2022) | China (Gu et al. 2022) |
| Pe. hainanensis | Pe. intermedia | Unidentified host (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012) |
| Pe. hainanensis | Pe. linearis | Trachelospermum sp. (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012) |
| Pe. hainanensis | Pe. rosarioides | Rhododendron decorum (Gu et al. 2022) | China (Gu et al. 2022) |
| Pe. hainanensis | Pe. tumida | Rosa chinensis (Peng et al. 2022) | China (Peng et al. 2022; Razaghi et al. 2024) |
| Pe. hawaiiensis | Leucospermum cv. Coral (Maharachchikumbura et al. 2014b) | America (Maharachchikumbura et al. 2014b) | |
| Pe. hispanica | Cryptomeria japonica (Fan et al. 2023), Ophiocordyceps sp. (Hsu et al. 2024), Peristrophe japonica (Sun et al. 2023), Protea ‘Susara’ (Liu et al. 2019) | China (Fan et al. 2023; Sun et al. 2023; Hsu et al. 2024), Spain (Liu et al. 2019) | |
| Pe. hunanensis | Camellia oleifera (Li et al. 2021b) | China (Li et al. 2021b) | |
| Pe. hunanensis | Pe. jiangsuensis | Pinus massoniana (Li et al. 2024a) | China (Li et al. 2024a) |
| Pe. hydei | Cyclosorus sp. (Seifollahi et al. 2023), Litsea elliptica, Litsea petiolata (Huanaluek et al. 2021), Unidentified host (Sun et al. 2023) | China (Sun et al. 2023), Thailand (Huanaluek et al. 2021; Seifollahi et al. 2023) | |
| Pe. inflexa | Kashiwadia orientalis (Oh et al. 2020), Unidentified host (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012), Korea (Oh et al. 2020) | |
| Pe. italiana | Cupressus glabra (Liu et al. 2015) | Italy (Liu et al. 2015) | |
| Pe. jesteri | Acer saccharum (Das et al. 2008), Fragraea bodenii (Maharachchikumbura et al. 2014b), Pinus strobus (Broders et al. 2015) | America (Das et al. 2008; Broders et al. 2015), Papua New Guinea (Maharachchikumbura et al. 2014b) | |
| Pe. jinchanghensis | Camellia sinensis (Liu et al. 2017), Vaccinium dunalianum (Fan et al. 2020) | China (Liu et al. 2017; Fan et al. 2020) | |
| Pe. kandelicola | Kandelia candel (Hyde et al. 2020b), Sabal mexicana (Xiong et al. 2022) | China (Hyde et al. 2020b; Xiong et al. 2022) | |
| Pe. knightiae | Banksia ericifolia, Banksia serrata (Mertin et al. 2022), Knightia sp. (Maharachchikumbura et al. 2014b), Sea sand (Park et al. 2021), Stereocaulon japonicum (Oh et al. 2020), Wood (Park et al. 2021) | Australia (Mertin et al. 2022), Korea (Oh et al. 2020), Micronesia (Park et al. 2021), New Zealand (Maharachchikumbura et al. 2014b) | |
| Pe. krabiensis | Pandanus sp. (Tibpromma et al. 2018b) | Thailand (Tibpromma et al. 2018b) | |
| Pe. leucadendri | Leucadendron sp. (Liu et al. 2019) | South Africa (Liu et al. 2019) | |
| Pe. leucospermi | Leucospermum cv. Pink Ice (Razaghi et al. 2024) | America (Razaghi et al. 2024) | |
| Pe. licualacola | Licuala grandis (Geng et al. 2013) | China (Geng et al. 2013) | |
| Pe. lijiangensis | Castanopsis carlesii var. spinulosa (Zhou et al. 2018) | China (Zhou et al. 2018) | |
| Pe. linguae | Pyrrosia lingua (Li et al. 2023a) | China (Li et al. 2023a) | |
| Pe. lithocarpi | Lithocarpus chiungchungensis (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. lobata | Lithocarpus glaber (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. loeiana | Unidentified host (Sun et al. 2023) | Thailand (Sun et al. 2023) | |
| Pe. macadamiae | Macadamia integrifolia (Akinsanmi et al. 2017), Macadamia sp. (Prasannath et al. 2023) | Australia (Akinsanmi et al. 2017; Prasannath et al. 2023) | |
| Pe. machili | Machilus sp. (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. malayana | Macaranga triloba colonised by ants (Maharachchikumbura et al. 2014b), Traded seed (Franić et al. 2019) | Asia (Franić et al. 2019), Malaysia (Maharachchikumbura et al. 2014b) | |
| Pe. manyueyuanensis | Stroma of Ophiocordyceps sp. parasitic on an insect (Hsu et al. 2024) | China (Hsu et al. 2024) | |
| Pe. matildae | Thamnochortus spicigerus (Lee et al. 2006) | South Africa (Lee et al. 2006) | |
| Pe. menhaiensis | Camellia sinensis (Wang et al. 2019c) | China (Wang et al. 2019c) | |
| Pe. menhaiensis | Pe. kaki | Diospyros kaki (Das et al. 2021) | Korea (Das et al. 2021) |
| Pe. monochaeta | Quercus robur, Taxus baccata (Maharachchikumbura et al. 2014b) | Baarn, Netherlands (Maharachchikumbura et al. 2014b) | |
| Pe. montellica | Dead plant material (Ariyawansa et al. 2015) | China (Ariyawansa et al. 2015) | |
| Pe. multiappendiculata | – | China (Razaghi et al. 2024) | |
| Pe. multicolor | Camellia sp. (Razaghi et al. 2024), Taxus chinensis (Wang et al. 2024) | China (Razaghi et al. 2024; Wang et al. 2024) | |
| Pe. nanningensis | Camellia oleifera (Li et al. 2021b), Pyrrosia lingua (Li et al. 2023a) | China (Li et al. 2021b; Li et al. 2023a) | |
| Pe. nannuoensis | Rotted leaves (Yin et al. 2024) | China (Yin et al. 2024) | |
| Pe. neolitseae | Liquidambar formosana (Fan et al. 2022), Lithocarpus amygdalifolius (Jiang et al. 2022), Magnolia garrettii (de Silva et al. 2021), Morus australis (Tennakoon et al. 2021), Neolitsea villosa (Ariyawansa and Hyde 2018), Photinia × fraseri (Xu et al. 2023c) | China (Ariyawansa and Hyde 2018; Tennakoon et al. 2021; Fan et al. 2022; Jiang et al. 2022), Thailand (de Silva et al. 2021) | |
| Pe. novae-hollandiae | Banksia grandis (Maharachchikumbura et al. 2014b) | Australia (Maharachchikumbura et al. 2014b) | |
| Pe. oryzae | Dysosma versipellis (Tan et al. 2018), Oryza sativa, Telopea sp., Unknown substrate (Maharachchikumbura et al. 2014b) | China (Tan et al. 2018), Denmark, Italy, America (Maharachchikumbura et al. 2014b) | |
| Pe. pallidotheae | Camptotheca cuminata (Su et al. 2014), Citrus limon (Douanla-Meli et al. 2013), Pieris japonica (Watanabe et al. 2010) | Cameroon (Douanla-Meli et al. 2013), China (Su et al. 2014), Japan (Watanabe et al. 2010)S | |
| Pe. pandanicola | Pandanus sp. (Tibpromma et al. 2018b) | Thailand (Tibpromma et al. 2018b) | |
| Pe. parva | Airborne, aquatic, and marshland soil (Choi et al. 2024), Delonix regia, Leucothoe fontanesiana (Maharachchikumbura et al. 2014b), Macaranga peltate (Reddy et al. 2016), Macaranga sp. (Lateef et al. 2018), Metrosideros polymorpha (Chock et al. 2021), Morus australis (Tennakoon et al. 2021), Phoenix sp. (Senanayake et al. 2020), Syzygium jambos (Chock et al. 2021) | America (Chock et al. 2021), China (Senanayake et al. 2020; Tennakoon et al. 2021), India (Reddy et al. 2016), Korea (Choi et al. 2024), Malaysia (Lateef et al. 2018), Unknown country (Maharachchikumbura et al. 2014b) | |
| Pe. phoebes | Phoebe zhennan (Zhang et al. 2023) | China (Zhang et al. 2023) | |
| Pe. photinicola | Mangifera indica (Shu et al. 2020), Photinia serrulate (Chen et al. 2017) | China (Chen et al. 2017; Shu et al. 2020) | |
| Pe. phyllostachydis | Phyllostachys sulphurea (Zhao et al. 2024) | China (Zhao et al. 2024) | |
| Pe. phyllostachydis | Pe. cratoxyli | Cratoxylum cochinchinense, Excoecaria cochinchinensis (Razaghi et al. 2024) | China (Razaghi et al. 2024) |
| Pe. pinicola | Pinus armandii (Tibpromma et al. 2019) | China (Tibpromma et al. 2019) | |
| Pe. piraubensis | Psidium guajava (Jayawardena et al. 2022) | Brazil (Jayawardena et al. 2022) | |
| Pe. portugallica | Camellia chekiangoleosa, Camellia japonica, Camellia sp. (Liu et al. 2017), Ficus septica (Tennakoon et al. 2021), Seaweeds (Goshima 2022), Unidentified host (Maharachchikumbura et al. 2014b) | China (Liu et al. 2017; Tennakoon et al. 2021), Japan (Goshima 2022), Portugal (Maharachchikumbura et al. 2014b) | |
| Pe. pruni | Prunus cerasoides (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. rhaphiolepidis | Rhaphiolepis indica (Yin et al. 2024) | China (Yin et al. 2024) | |
| Pe. rhizophorae | Rhizophora apiculate (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| Pe. rhodomyrti | Camellia sinensis (Liu et al. 2017; Manawasinghe et al. 2021), Cyclobalanopsis augustinii, Quercus aliena (Jiang et al. 2022), Rhodomyrtus tomentosa (Song et al. 2013), Rosa chinensis, Rosa multiflora, Rosa rugosa (Peng et al. 2022) | China (Song et al. 2013; Liu et al. 2017; Manawasinghe et al. 2021; Jiang et al. 2022; Peng et al. 2022) | |
| Pe. rosea | Ericaceae (Kohout and Tedersoo 2017), Pinus sp. (Maharachchikumbura et al. 2012), Vicia faba (Wei et al. 2023) | China (Maharachchikumbura et al. 2012; Wei et al. 2023), South Africa (Kohout and Tedersoo 2017) | |
| Pe. rubrae | Plagiogyria glauca, Quercus rubra (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. sabal | Sabal mexicana (Xiong et al. 2022) | China (Xiong et al. 2022) | |
| Pe. scoparia | Chamaecyparis sp. (Maharachchikumbura et al. 2014b), Egg mass of the sailfin sandfish (Park et al. 2018), Rhododendron delavayi (Zhu et al. 2023b) | China (Zhu et al. 2023b), Korea (Park et al. 2018) | |
| Pe. sequoiae | Sequoia sempervirens (Hyde et al. 2016) | Italy (Hyde et al. 2016) | |
| Pe. shaanxiensis | Quercus variabilis (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. shorea | Shorea obtuse (Song et al. 2014b) | Thailand (Song et al. 2014b) | |
| Pe. sichuanensis | Camellia sinensis (Wang et al. 2019c) | China (Wang et al. 2019c) | |
| Pe. sichuanensis | Pe. nanjingensis | Camellia oleifera (Li et al. 2021b), Panax quinquefolius (Liyanapathiranage et al. 2023), Quercus aliena (Jiang et al. 2022) | China (Li et al. 2021b; Jiang et al. 2022), America (Liyanapathiranage et al. 2023) |
| Pe. silvicola | Cyclobalanopsis kerrii (Jiang et al. 2022) | China (Jiang et al. 2022) | |
| Pe. smilacicola | Smilax china, Smilax sp. (Sun et al. 2023) | Thailand (Sun et al. 2023) | |
| Pe. sonneratiae | Sonneratia apetala (Jiang et al. 2023) | China (Jiang et al. 2023) | |
| Pe. spathulata | Guevina avellana (Maharachchikumbura et al. 2014b) | Chile (Maharachchikumbura et al. 2014b) | |
| Pe. spathuliappendiculata | Phoenix canariensis (Liu et al. 2019) | Australia (Liu et al. 2019) | |
| Pe. suae | Rhododendron delavayi (Gu et al. 2022) | China (Gu et al. 2022) | |
| Pe. telopeae | Macadamia sp. (Prasannath et al. 2023), Protea neriifolia × susannae cv. Pink Ice, Telopea sp. (Maharachchikumbura et al. 2014b) | Australia (Maharachchikumbura et al. 2014b; Prasannath et al. 2023) | |
| Pe. terricola | Soil (Liu et al. 2019) | Pacific Islands (Liu et al. 2019) | |
| Pe. thailandica | Polyethylene terephthalate waste (Kim et al. 2022b), Rhizophora apiculate (Norphanphoun et al. 2019) | Korea (Kim et al. 2022b), Thailand (Norphanphoun et al. 2019) | |
| Pe. trachycarpicola | Camellia sinensis (Liu et al. 2017; Tsai et al. 2021; Xia et al. 2022), Celtis formosana (Tennakoon et al. 2021), Chrysophyllum sp. (Maharachchikumbura et al. 2012), Dendrobium loddigesii (Ma et al. 2019), Historic documents (Escudero-Leyva et al. 2023), Mangifera indica (Shu et al. 2020), Ophiocordyceps sp. (Hsu et al. 2024), Panax notoginseng (Lan et al. 2023), Photinia x fraseri (Xu et al. 2022), Pinus bungeana (Qi et al. 2021), Podocarous macrophyllus (Maharachchikumbura et al. 2012), Podocarpus macrophyllus (Zhang et al. 2013b), Rhododendron cyanocarpum (Gu et al. 2022), Rosa roxburghii (Zhang et al. 2021b), Schima sp. (Maharachchikumbura et al. 2012), Sorghum bicolor (Fan et al. 2021), Symplocos sp. (Maharachchikumbura et al. 2012), Taxus chinensis (Wang et al. 2024), Trachycarpus fortunei (Maharachchikumbura et al. 2012; Zhang et al. 2012a), Vaccinium sp. (Araujo et al. 2023), Vaccinium corymbosum (Zheng et al. 2023b), Vitis vinifera (Jayawardena et al. 2015; Ghuffar et al. 2018) | Brazil (Araujo et al. 2023), China (Liu et al. 2017; Tennakoon et al. 2021; Tsai et al. 2021; Xia et al. 2022; Hsu et al. 2024), Costa Rica (Escudero-Leyva et al. 2023), Pakistan (Ghuffar et al. 2018) | |
| Pe. trachycarpicola | Pe. kenyana | Camellia oleifera (Li et al. 2022), Camellia sinensis (Liu et al. 2017; Manawasinghe et al. 2021), Camellia sp. (Liu et al. 2017), Castanea henryi, Castanea mollissima (Jiang et al. 2021), Castanopsis fissa, Castanopsis hystrix (Jiang et al. 2022), Coffea sp. (Maharachchikumbura et al. 2014b), Cyclobalanopsis fleuryi, Cyclobalanopsis glauca, Cyclobalanopsis neglecta (Jiang et al. 2022), Historic documents (Escudero-Leyva et al. 2023), Magnolia garrettii (de Silva et al. 2021), Marine animals (Godinho et al. 2019), Paeonia suffruticosa (Li et al. 2022), Quercus aliena (Jiang et al. 2022), Raw material from agar-agar (Maharachchikumbura et al. 2014b), Rhododendron agastum (Li et al. 2024b), Stereocaulon japonicum (Oh et al. 2020) | Antarctica (Godinho et al. 2019), China (Liu et al. 2017; Jiang et al. 2021; Manawasinghe et al. 2021; Jiang et al. 2022; Li et al. 2022; Li et al. 2024b), Costa Rica (Escudero-Leyva et al. 2023), Kenya (Maharachchikumbura et al. 2014b), Korea (Oh et al. 2020), Thailand (de Silva et al. 2021), Unknown country (Maharachchikumbura et al. 2014b) |
| Pe. unicolor | Rhododendron sp. (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012) | |
| Pe. unicolor | Pe. jiangxiensis | Camellia sp., Eurya sp. (Liu et al. 2017), Pandanus sp. (Tibpromma et al. 2018a) | China (Liu et al. 2017), Thailand (Tibpromma et al. 2018a) |
| Pe. unicolor | Pe. taxicola | Taxus chinensis (Wang et al. 2024) | China (Wang et al. 2024) |
| Pe. verruculosa | Clematis vitalba (Phukhamsakda et al. 2020), Rhododendron sp. (Maharachchikumbura et al. 2012) | China (Maharachchikumbura et al. 2012), Italy (Phukhamsakda et al. 2020) | |
| Pe. wulichongensis | Poaceae (Razaghi et al. 2024) | China (Razaghi et al. 2024) | |
| Pe. yunnanensis | Podocarpus macrophyllus (Wei et al. 2013) | China (Wei et al. 2013) | |
| Pe. zhaoqingensis | Unidentified host (Dong et al. 2023) | China (Dong et al. 2023) | |
| Ps. ampullacea | Camellia chrysantha (Zhao et al. 2020), Camellia sinensis, Lauraceae (Liu et al. 2017), Magnolia candolli (de Silva et al. 2021), Mangifera indica (Shu et al. 2020), Oil palm hybrid (Betancourt-Ortiz et al. 2024) | China (Liu et al. 2017; Shu et al. 2020; Zhao et al. 2020; de Silva et al. 2021), Colombia (Betancourt-Ortiz et al. 2024) | |
| Ps. ampullacea | Ps. avicenniae | Avicennia marina (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) |
| Ps. ampullacea | Ps. elaeidis | Acacia crassipes, Averrhoa carambola, Elaeis guineensis, Lauraceae (Liu et al. 2019) | China, Indonesia, Myanmar, Nigeria (Liu et al. 2019) |
| Ps. annellata | Camellia sinensis (Tsai et al. 2021) | China (Tsai et al. 2021) | |
| Ps. celtidis | Celtis sinensis (Yang et al. 2022) | China (Yang et al. 2022) | |
| Ps. camelliae-sinensis | Camellia sinensis (Liu et al. 2017; Chen et al. 2018c; Wang et al. 2019b; Tsai et al. 2021; Pandey et al. 2023), Ficus ampelas (Tennakoon et al. 2021), Styrax sp. (Elfiati et al. 2022), Vitis vinifera (Jayawardena et al. 2018) | China (Liu et al. 2017; Chen et al. 2018c; Wang et al. 2019b; Tennakoon et al. 2021; Tsai et al. 2021), India (Pandey et al. 2023), Indonesia (Elfiati et al. 2022), Italy (Jayawardena et al. 2018) | |
| Ps. chinensis | Camellia sinensis (Liu et al. 2017; Wang et al. 2019b; Manawasinghe et al. 2021; Tsai et al. 2021; Undugoda et al. 2023; Koodalugodaarachchi et al. 2024), Pteridophytes (Singha et al. 2024) | China (Liu et al. 2017; Wang et al. 2019b; Manawasinghe et al. 2021; Tsai et al. 2021), India (Singha et al. 2024), Sri Lanka (Undugoda et al. 2023), Thailand (Koodalugodaarachchi et al. 2024) | |
| Ps. cocos | Cocos nucifera (Maharachchikumbura et al. 2014b), Oil palm hybrid (Betancourt-Ortiz et al. 2024) | Colombia (Betancourt-Ortiz et al. 2024), Indonesia (Maharachchikumbura et al. 2014b) | |
| Ps. curvatispora | Rhizophora apiculata, Rhizophora mucronate (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| Ps. dasymaschalonis | Dasymaschalon yunnanense (This study) | Thailand (This study) | |
| Ps. dawaina | Caryota mitis (Catarino et al. 2020), Unidentified host (Nozawa et al. 2018) | Brazil (Catarino et al. 2020), Myanmar (Nozawa et al. 2018) | |
| Ps. gilvanii | Paullinia cupana var. sorbilis (Gualberto et al. 2021) | Brazil (Gualberto et al. 2021) | |
| Ps. ignota | Camellia sinensis (Maharachchikumbura et al. 2016c) | China (Maharachchikumbura et al. 2016c) | |
| Ps. indica | Hibiscus rosa-sinensis (Maharachchikumbura et al. 2014b), Sea sand, Seaweed (Park et al. 2021) | India (Maharachchikumbura et al. 2014b), Micronesia (Park et al. 2021) | |
| Ps. indocalami | Indocalamus tessellatus (Yang et al. 2022) | China (Yang et al. 2022) | |
| Ps. ixorae | Ixora sp. (Tsai et al. 2018) | China (Tsai et al. 2018) | |
| Ps. kawthaungina | Unidentified host (Nozawa et al. 2018) | Myanmar (Nozawa et al. 2018) | |
| Ps. kubahensis | Macaranga sp. (Lateef et al. 2015) | Malaysia (Lateef et al. 2015) | |
| Ps. rhizophorae | Rhizophora apiculate (Norphanphoun et al. 2019) | Thailand (Norphanphoun et al. 2019) | |
| Ps. rhizophorae | Ps. thailandica | Camellia sinensis (Pandey et al. 2023), Rhizophora mucronate (Norphanphoun et al. 2019) | India (Pandey et al. 2023), Thailand (Norphanphoun et al. 2019) |
| Ps. simitheae | Magnolia candolli (de Silva et al. 2021), Pandanus odoratissimus (Song et al. 2014b; Maharachchikumbura et al. 2016c) | China (de Silva et al. 2021), Thailand (Song et al. 2014b; Maharachchikumbura et al. 2016c) | |
| Ps. solicola | Soil in tropical forest (Liu et al. 2019) | Papua New Guinea (Liu et al. 2019) | |
| Ps. taiwanensis | Hevea brasiliensis (de Oliveira Amaral et al. 2022), Ixora sp. (Tsai et al. 2018) | Brazil (de Oliveira Amaral et al. 2022), China (Tsai et al. 2018) | |
| Ps. theae | Airborne, aquatic, and marshland soil (Choi et al. 2024), Aloe vera (Ahmmed et al. 2022), Ananus comosus (Bhattacharya et al. 2019), Caloncoba welwitschia (Akone et al. 2022), Camellia macrophyllus, Camellia nitidissima, Camellia reticulata (Wei et al. 2007), Camellia sinensis (Wei et al. 2007; Maharachchikumbura et al. 2013a; Maharachchikumbura et al. 2014b; Wang et al. 2017; Tsai et al. 2021; Win et al. 2021; Pandey et al. 2023), Ceriops tagal (Sun et al. 2023), Citrus limon (Douanla-Meli et al. 2013), Clarias gariepinus (Zakaria et al. 2021), Coffea canephora (Sumaya et al. 2023), Cordia dichotoma (Reddy et al. 2016), Diospyros crassiflora (Douanla-Meli and Langer 2009), Elaeis guineensis (Suwannarach et al. 2013; Mohamed-Azni et al. 2022), Euonymus japonicus (Ma et al. 2023a), Gyrinops versteegii (Hidayat et al. 2019), Holarrhena antidysenterica (Tejesvi et al. 2008), Homo sapiens (Sane et al. 2019; Borgohain et al. 2020), Ixora chinensis (Mukhtar et al. 2019), Khaya anthotheca (Linnakoski et al. 2012), Library air (Okpalanozie et al. 2018), Macaranga sp. (Lateef et al. 2018), Mangifera indica (Sandoval-Sánchez et al. 2013), Musa sp. (Zakaria and Aziz 2018), Orchids (Adit et al. 2022), Palm (Hyde et al. 2020a), Pearl millet (Nandhini et al. 2018), Phoenix dactylifera (Tao et al. 2021), Rhaphiolepis umbellate (Watanabe et al. 2010), Rhizophora racemose (Yu et al. 2020), Seleniferous agricultural soils (Kaur et al. 2022), Soil (Watanabe et al. 2010), Terminalia arjuna (Tejesvi et al. 2008), Thea sinensis (Watanabe et al. 2010), Turraeanthus longipes (Akone et al. 2013) | Bangladesh (Ahmmed et al. 2022), Cameroon (Douanla-Meli and Langer 2009; Akone et al. 2013; Douanla-Meli et al. 2013; Akone et al. 2022), China (Wei et al. 2007; Maharachchikumbura et al. 2014b; Wang et al. 2017; Mukhtar et al. 2019; Tsai et al. 2021; Ma et al. 2023a), Ghana (Linnakoski et al. 2012), India (Tejesvi et al. 2008; Reddy et al. 2016; Bhattacharya et al. 2019; Sane et al. 2019; Borgohain et al. 2020; Adit et al. 2022; Kaur et al. 2022; Pandey et al. 2023), Indonesia (Hidayat et al. 2019), Japan (Watanabe et al. 2010; Win et al. 2021), Korea (Choi et al. 2024), Malaysia (Lateef et al. 2018; Zakaria and Aziz 2018; Zakaria et al. 2021; Mohamed-Azni et al. 2022), Mexico (Sandoval-Sánchez et al. 2013), Nigeria (Okpalanozie et al. 2018; Yu et al. 2020), Philippines (Sumaya et al. 2023), Thailand (Maharachchikumbura et al. 2013a; Suwannarach et al. 2013; Maharachchikumbura et al. 2014b; Sun et al. 2023) | |
| Ps. vietnamensis | Fragaria sp. (Nozawa et al. 2017) | Vietnam (Nozawa et al. 2017) |
Discussion
In this study, we employed an integrative taxonomic framework combining multi-locus phylogenetic analyses, genome-scale data, and multiple species delimitation approaches to reassess pestalotiopsis-like fungi. In Neopestalotiopsis, seven previously described taxa were synonymized, and one new species, N. camelliae, was introduced. In Pestalotiopsis, four species complexes were recognized and 18 taxa were synonymized based on phylogenetic topology, sequence similarity, and genomic evidence. In Pseudopestalotiopsis, one species complex was established, three taxa were synonymized, and one new species, Ps. dasymaschalonis, was described. The integration of GCPSR, PTP, mPTP, genomic ANI, and core-genome similarity analyses, together with comprehensive assessments of recombination signals and sequence polymorphisms, provided a robust and reliable framework for accurately delineating species boundaries within pestalotiopsis-like fungi. Collectively, this study refines the taxonomy of Pestalotiopsis-like fungi, reduces species-level redundancy, and establishes a phylogenomic foundation for future systematic, ecological, and evolutionary research within the family Sporocadaceae.
Limitations of Morphological Characteristics in the Taxonomy of Pestalotiopsis-like taxa
The morphological characteristics of conidia have historically been the most widely used features for intergeneric and interspecific classification within pestalotiopsis-like taxa (Maharachchikumbura et al. 2014b). Maharachchikumbura et al. (2014b) proposed colour of three median cells in conidial as a key diagnostic feature at the generic level within Neopestalotiopsis, Pestalotiopsis, and Pseudopestalotiopsis. However, with the growing number of newly described species, numerous exceptions to this classification criterion have arisen. For instance, Neopestalotiopsis, originally defined by its versicolored conidia, has been found to include species with concolorous conidia, such as N. ageratinae, N. amomi, N. castanopsidis, N. chiangmaiensis, N. dimorphospora, N. dolichoconidiophora, N. fijiensis, N. fimbriata, N. fructicola, N. fuzhouensis, N. hyperici, N. megabetaspora, N. moniliformis, N. olivaceous, N. pandanicola, N. phangngaensis, and N. rhododendricola (Tibpromma et al. 2018b; Chaiwan et al. 2022; Sun et al. 2023; Zhang et al. 2024b; Cui et al. 2024; Razaghi et al. 2024). Conversely, Pestalotiopsis and Pseudopestalotiopsis, which are generally characterized by concolorous conidia, have been found to include species with versicolored conidia, such as Pe. aggestorum (Liu et al. 2017), Pe. appendiculata (Gu et al. 2022), Pe. biappendiculata (Razaghi et al. 2024), Pe. dianellae (Crous et al. 2017), Pe. doitungensis (Ma et al. 2019), Pe. gibbosa (Watanabe et al. 2018), Pe. guiyangensis (Zhang et al. 2024d), Pe. kandelicola (Hyde et al. 2020b), Pe. matildae (Lee et al. 2006), Pe. multicolor (Wang et al. 2024), Pe. pinicola (Tibpromma et al. 2019), Pe. pyrrosiae-linguae (Dong et al. 2023), Pe. sabal (Xiong et al. 2022), Pe. suae (Gu et al. 2022), Pe. taxicola (Wang et al. 2024), Pe. thailandica, Ps. avicenniae, Ps. curvatispora, Ps. rhizophorae, and Ps. thailandica (Norphanphoun et al. 2019). In addition, in the description of new species, concolorous and versicolored conidia can be observed, as in the cases of N. brasiliensis (Bezerra et al. 2018) and Pe. zhaoqingensis (Dong et al. 2023). These inconsistencies highlight the limitations of using colour of three median cells in conidial as a reliable taxonomic criterion for pestalotiopsis-like taxa.
Conidial size has historically been considered an important criterion for species delimitation and has been widely used to differentiate pestalotiopsis-like taxa. However, findings from this study and previous literature suggest that species with similar conidial dimensions may belong to distinct evolutionary clades (Maharachchikumbura et al. 2014b). Li et al. (2022) observed that the conidial morphology, size, and colour of Neopestalotiopsis terricola, originally isolated from Paeonia suffruticosa, underwent significant changes after being inoculated onto olive leaves. Similarly, the conidia of N. paeonia-suffruticosa produced on PDA medium were shorter than those naturally formed on woody plants (Li et al. 2022). Such variations are also observed in Seiridium, a genus within the same family (Sporocadaceae) of pestalotiopsis-like fungi. For instance, different strains of S. cupressi (IMI 52254 and IMI 52255) exhibit considerable differences in conidial size when observed in herbarium material (Bonthond et al. 2018). Additionally, strains of S. neocupressi (CBS 142625, CBS 142627, and CBS 142626) displayed variations in conidial size even when cultured under identical conditions on SNA medium (Bonthond et al. 2018). These findings indicate that conidial size is not always a dependable taxonomic marker, as it can vary considerably due to genetic differences, environmental factors, and growth conditions.
Conidial appendages play a significant taxonomic role in many coelomycetous genera (Crous et al. 2012). Among coelomycetous fungi with appendages, pestalotiopsis-like taxa exhibit highly variable appendage morphology (Crous et al. 2012; Maharachchikumbura et al. 2014b). Appendage characteristics have been widely used in the classification of Neopestalotiopsis, Pestalotiopsis, and Pseudopestalotiopsis, aiding in species identification and the establishment of new taxonomic units (Liu et al. 2019; Norphanphoun et al. 2019; Gu et al. 2022). These characteristics include the length, number, shape, branching pattern, presence of swollen apices, and attachment position on the conidium. However, whether conidial appendages alone can serve as reliable taxonomic markers remains a question worth careful consideration. The key to determining whether a morphological trait is suitable for species delimitation is its stability and reproducibility. In pestalotiopsis-like fungi, even within a single strain, appendage number can vary considerably, and this is not an isolated phenomenon. For instance, Neopestalotiopsis caulicola produces conidia with 1–4 apical appendages within the same colony on PDA medium (Zhang et al. 2024b), while N. olivaceous produces conidia with 2–5 apical appendages under the same conditions (Cui et al. 2024). Liu et al. (2017) conducted a comprehensive study on Pestalotiopsis species, examining characteristics such as apical and basal appendages, conidiogenous cells, and conidial morphology. The study observed differences only in basal appendages, and while these morphological traits partially corresponded with phylogenetic analyses, they did not reveal clear evolutionary relationships (Liu et al. 2017). Strains of Seiridium cupressi (IMI 52254 and IMI 52255) exhibited large differences in basal appendage length when observed in herbarium material, while strains of S. unicorne (IMI 5816 and CBS 538.82) displayed significant variation in both apical and basal appendage lengths under different growth conditions (Bonthond et al. 2018). This raises the question of whether species primarily distinguished by appendage morphology might face the same taxonomic challenges as genera that were historically separated based on appendage characteristics. For example, Dinemasporium and Stauronema, as well as Seimatosporium and Vermisporium, were once classified as distinct genera based on appendage morphology but were later recognized as representing only two valid genera, Dinemasporium and Seimatosporium (Barber et al. 2011; Crous et al. 2012). Crous et al. (2012) cautioned against evaluating appendage function in isolation, emphasizing that their role is often linked to ecological functions such as spore dispersal, release, deposition, and colonization of new substrates or ecological niches (Crous et al. 2012; Maharachchikumbura et al. 2014b).
Therefore, although conidial morphological characteristics still hold some taxonomic significance in classifying fungi, they exhibit instability among pestalotiopsis-like species. Consequently, depending exclusively on morphology for species delimitation is questionable, especially when genetic differences between species are minimal, yet morphological variation is substantial. The current description of new species primarily relies on phylogenetic analyses using a three-gene dataset (ITS, tef1, tub2). Researchers first position isolates within phylogenetic trees and then compare their morphology with closely related species. Therefore, we recommend using molecular data as the primary criterion for species identification, with morphological characteristics serving only as supplementary references. Recent studies further support this perspective. Razaghi et al. (2024) revised several species based on three-gene (ITS, tef1, and tub2) sequence similarity and phylogenetic distance, demonstrating that species with significant morphological differences but minor molecular variations should be synonymized. For instance, Neopestalotiopsis rhapidis (conidial size: 22–25.5 × 4–6 μm, av. = 23 × 5.2 μm; three median cells versicolorous) was synonymized with N. keteleeriae (18.5–24 × 7–9.5 μm, av. = 22 × 8.5 μm; three median cells concolorous or sometimes versicolorous). Similarly, N. vaccinii (conidial size: 11.0–15.2 × 4.9–7.5 μm, av. = 13.6 × 6.4 μm; basal appendage length: 1.7–6.6 μm) was synonymized with N. hispanica (conidial size: 21.5–27.1 × 5.9–8.7 μm, av. = 24.9 × 7.5 μm; basal appendage length: 5.1–15.5 μm). Additionally, Pestalotiopsis kaki was synonymized with Pe. menhaiensis, and Pe. nanjingensis with Pe. sichuanensis.
Use of Molecular Barcodes
Maharachchikumbura et al. (2012) identified ITS, tef1, and tub2 as the most effective markers for distinguishing pestalotiopsis-like fungi based on their PCR amplification efficiency and capability to delineate species boundaries. This research established a standardized molecular marker system for the classification of pestalotiopsis-like fungi, enhancing data sharing and comparability among various studies. Following this, most classification efforts on pestalotiopsis-like fungi have predominantly utilized these three loci, creating a cohesive framework for global research on this group (He et al. 2022; Peng et al. 2022; Sun et al. 2023; Yin et al. 2024). This study aimed to further validate the effectiveness of ITS, tef1, and tub2 by comparing the performance of these markers in three-gene phylogenetic analyses, whole-genome phylogenetic analyses, and species delimitation based on GCPSR. The ITS region is considered the standard fungal DNA barcode and is commonly utilized in fungal systematics to define species and elucidate their evolutionary relationships (Schoch et al. 2012). Nonetheless, the effectiveness of ITS has been challenged because of various known issues, particularly concerning its resolution and the potential existence of non-homologous ITS copies in genomes (Lücking et al. 2020). Research indicates that ITS often lacks adequate resolution among many closely related species, especially within indoor and foodborne molds (e.g., Aspergillus, Penicillium) and economically and medically significant plant, human, and animal pathogens (e.g., Alternaria, Cladosporium, Colletotrichum, Fusarium). This study faced a similar challenge, as ITS was unable to identify numerous branches of pestalotiopsis-like fungi, which have short evolutionary branches. Although ITS struggles with defining species boundaries in pestalotiopsis-like fungi, it effectively places strains within their corresponding genera and clades. Additionally, ITS still offers a higher sequencing success rate compared to many other molecular markers (Maharachchikumbura et al. 2012). Thus, it remains an appropriate method for examining genetic diversity, population structure, and species relationships with their hosts and geographic distributions. A crucial strategy to address ITS limitations involves integrating it with other genetic markers.
Protein-coding genes typically show greater phylogenetic resolution compared to ribosomal gene regions (Hofstetter et al. 2007). The tef1 gene has been recognized as a significant marker for clarifying the evolutionary links among cryptic species of pestalotiopsis-like fungi (Maharachchikumbura et al. 2012). In this study, the tef1 gene exhibited the highest phylogenetic informativeness, with longer branches observed in most clades of Neopestalotiopsis and Pestalotiopsis compared to the ITS and tub2 phylogenies (Figs. S1, S2). This conclusion matches the topology of the multi-gene combined analysis, showing strong consistency with the phylogenetic tree of tef1, particularly in the consistency of terminal branches. However, genome-based phylogenetic trees and ANI analysis suggest that such a highly polymorphic locus may lead to significant taxonomic inflation in pestalotiopsis-like fungi. To further assess the impact of tef1 variability, we constructed single-gene trees for ITS, tef1, and tub2, as well as a three-gene combined tree (Figs. S10, S11), using the same set of strains as in the genome-based phylogenetic analysis. Within the Pe. adusta species complex, strains GUCC 23-0363 and GUCC 23-0366 were placed distantly in the tef1 phylogeny, and this divergence also influenced the structure of the three-gene tree, causing the formation of distinct branches. In contrast, these two strains clustered together with short branches in the genome-based phylogenetic tree. Moreover, their ANI similarity reached 99.31%, suggesting they belong to the same species. Recent studies on Diaporthe have also confirmed that highly polymorphic loci may lead to significant taxonomic inflation issues (Pereira & Phillips 2024). While the tub2 gene may not provide sufficient resolution for certain species, its results align more closely with the phylogenomic tree and ANI analysis. Consequently, future species classification should not concentrate only on genes with informative sites. Instead, it should seek to find more stable loci that accurately depict the overall evolutionary relationships among species. This approach enables clearer branching and greater support values in multi-gene phylogenetic analyses.
Further analysis of the combined effect of ITS, tef1, and tub2 on species delimitation reveals notable differences between Neopestalotiopsis and Pestalotiopsis. We compared the phylogenetic trees constructed from whole genomes and from the three genes (ITS, tef1, and tub2) using the same set of strains. Although not all strains used in this analysis are type strains, their sources are clearly documented and their sequence data are of high quality. Therefore, the use of non-type strains does not compromise the validity of the comparison. Our goal here is not to define species boundaries, but rather to compare the differences in phylogenetic resolution between genome-scale data and commonly used three-gene datasets when applied to the same strains. For Neopestalotiopsis, the three-gene (ITS, tef1, and tub2) phylogenetic tree (Fig. S10b), constructed with the same strains as the genome phylogenetic tree (Fig. S10a), exhibited shorter branches and lower or even no statistical support. Moreover, the topologies of the three-gene and genome-based trees were largely incongruent, indicating that the three commonly used markers lack sufficient phylogenetic signal to accurately resolve interspecific relationships. This suggests that species classification within Neopestalotiopsis based solely on ITS, tef1, and tub2 should be approached with caution. Further research is required to identify and evaluate more informative genetic markers for reliable species delimitation in this genus. In contrast, for Pestalotiopsis, the three-gene tree (ITS, tef1, and tub2) (Fig. S11b) displayed similar branch lengths and topologies compared to the genome-based tree (Fig. S11a). This indicates that the observed short branches in the three-gene phylogeny of Pestalotiopsis are likely due to species over-splitting rather than a lack of resolution in the markers themselves.
The Necessity of a Comprehensive Taxonomy for Pestalotiopsis-like Fungi
With advancements in technology, various methods for species delimitation have gradually emerged to tackle the problem of unclear species boundaries (Sklenář et al. 2022). Different data sources and analytical methods have been employed for the fundamental biological proposition of species delimitation, ranging from reliance on morphological characteristics and phylogenetic methods based on molecular fragments, to genetic cluster analysis informed by population genetics and the multi-species coalescent theory, as well as more in-depth and extensive population genomics techniques (Gutiérrez and Garbino 2018; Jorna et al. 2021; Oliveira‐Silva et al. 2023). Nonetheless, the outcomes of species delimitation using various classification evidence are frequently inconsistent. For example, this is similar to an earlier study using multi-locus sequence data (Spjut et al. 2020), robust, consistent species delimitations were difficult to infer, even with thousands of RADseq loci for lichen fungi (Jorna et al. 2021). This study yielded some minor discrepancies across different methods. For instance, species delimitation for certain clades within Pestalotiopsis and Pseudopestalotiopsis demonstrated that ABGD, ASAP, and mPTP merged more species compared to PTP. This also shows that species evolution often exhibits significant heterogeneity in molecular, and population characteristics. Therefore, recognizing that any species delimitation approach infers hypotheses of species boundaries, rather than providing the ultimate answer/solution and the extensive use of different data sources and different analytical approaches for species delimitation and the comprehensive analysis of the results, is imperative (Jorna et al. 2021; Sklenář et al. 2022).
Recent studies employing comprehensive species delimitation methods have often led to a decrease in the number of recognized species, even when phylogenomic approaches are used (Kim et al. 2022a; Obiol et al. 2023; Dissanayake et al. 2024; Pereira & Phillips 2024). These studies typically support employing a dataset with considerable variability and various analytical methods for species delimitation, as well as for an in-depth analysis of the results (Jorna et al. 2021; Sklenář et al. 2022). Regrettably, this criterion poses challenges in fungal taxonomic research, as it is hard to acquire numerous genetically similar isolates from diverse locations, particularly in microfungi. As a result, fungal species are often characterized using a small set of isolates that show limited variability (Hyde et al. 2023; Zhang et al. 2024a). As sampling increases, true intraspecific variability often becomes apparent only after sufficient diverse strains are gathered. By then, the species may have already been classified into multiple new species, though we cannot overlook the contributions made. This is primarily due to the efforts of earlier taxonomists and their contributions to increasing group classifications can guide future research and aid in the reassessment, leading to clearer identification of more robust species boundaries.
When described as new taxa, pestalotiopsis-like fungi are frequently based on a limited number of isolates or even a single isolate, resulting in low intraspecific diversity (Jiang et al. 2021; Prasannath et al. 2021; He et al. 2022). Currently, the number of pestalotiopsis-like taxa has reached a historical peak, with 123/132 records of Neopestalotiopsis, 448/489 records of Pestalotiopsis, and 31/34 records of Pseudopestalotiopsis in Index Fungorum (February 2025; http://www.indexfungorum.org/names/Names.asp) and MycoBank (Crous et al. 2004). Despite the ongoing addition of new species and the support of molecular methods, some isolates cannot be adequately identified to the species level, which may result in further confusion. This is particularly apparent for newly described taxa, where significant topological differences and short branches exist across different studies (Liu et al. 2017; Jiang et al. 2021). To determine whether they represent distinct species or populations, we initially identified species clusters using phylogenetic trees and networks. We then carried out species delimitation employing the GCPSR principle, two heuristic methods (namely, PTP and mPTP), and two genetic distance-based approaches (specifically, ABGD and ASAP). Furthermore, to enhance our assessment of species boundaries, we constructed a genomic phylogenetic tree, conducted ANI analysis, and calculated the core genome percentages.
The Application of Phylogenetics and GCPSR in Fungal Species Delimitation
Multi-locus sequence analysis is regarded as the gold standard for exploring the evolutionary relationships among pestalotiopsis-like fungi (Maharachchikumbura et al.2012; Tsai et al. 2021; Razaghi et al. 2024). Nevertheless, variations in evolutionary rates and histories across different genes (or sequence fragments) may create interactions within the same dataset, potentially resulting in conflicts (Li et al. 2021c). In this study, phylogenetic analyses based on concatenated datasets reveal several strongly supported lineages within pestalotiopsis-like fungi, which are often interpreted as distinct species. In contrast, our single-gene trees exhibit pronounced phylogenetic discordance among these lineages, including inconsistent node placements, conflicting topologies, and weak statistical support. Additionally, various strains of the same species frequently exhibit polyphyly or paraphyly in multiple single-gene phylogenies. This leads to decreased resolution in multi-locus phylogenetic trees and introduces uncertainty in the support for internal nodes across different phylogenetic trees. For instance, in Neopestalotiopsis and the Pe. rosea clade of Pestalotiopsis, while some units have diverged into well-supported terminal branches, the phylogenetic relationships between them remain unclear. This kind of phylogenetic discordance is common in pestalotiopsis-like fungi (Sun et al. 2023; Razaghi et al. 2024). This concern is frequently neglected during the introduction of new species. As a result, it has been consistently advised that delimiting species in fungi should depend on both multi-locus phylogenetic analyses and the strict application of genealogical concordance (GCPSR) (Taylor et al. 2000; Dissanayake et al. 2024; Pereira & Phillips 2024)
According to the GCPSR principle, the absence of lineage concordance between various gene trees suggests that the sampled diversity has not yet attained the species level. Conversely, phylogenetic congruence provides compelling evidence that different evolutionary lineages experience a lack of gene flow and thus possess independent evolutionary histories (Taylor et al. 2000). Using the GCPSR principle, we successfully identified most of the branches within Pestalotiopsis, with results corresponding to the same species, showing a genomic ANI greater than 97.30% and a core genome proportion higher than 85%. However, for the Pe. rosea clade within Pestalotiopsis and some taxa of Neopestalotiopsis, due to the absence of clear, independent branches in single-gene phylogenetic trees and low support in multi-gene phylogenetic analyses, we could not conclusively delimit species using GCPSR. Whether these taxa represent distinct evolutionary lineages or are simply in the early stages of species differentiation still requires further investigation for validation.
Species Delimitation Integrative Taxonomy in Pestalotiopsis-like Fungi
Overall, genome ANI, along with the proportion of core genes within the pangenome, offers a reliable approach for species delimitation. In this study, due to the unavailability of genome data for most pestalotiopsis-like fungi species, we only relied on genomic data to confirm the species delimitation derived from the three-gene phylogenetic analysis methods. In bacterial taxonomy, a 95% ANI threshold has been widely accepted as a standard criterion for species delineation, effectively replacing traditional DNA-DNA hybridization (DDH) (Jain et al. 2018). DDH is a classical molecular biology technique used to assess genomic similarity and phylogenetic relationships among bacterial populations, with a similarity threshold of ≥70% typically indicating conspecificity (Jain et al. 2018). However, no universally accepted ANI threshold has been established for fungal species identification, and a single cutoff may not be applicable across all fungal taxa. For example, Razaghi et al. (2024) recommended a 98% ANI threshold for species delimitation in Neopestalotiopsis. In our study, species boundaries within pestalotiopsis-like taxa, as determined based on the GCPSR framework, generally exhibited ANI values exceeding 97.30%, with core genes accounting for more than 85% of the pangenome. Similarly, McCarthy & Fitzpatrick (2019) analyzed 12 strains of Aspergillus fumigatus, 25 strains of Cryptococcus neoformans, 34 strains of Candida albicans, and 100 strains of Saccharomyces cerevisiae, demonstrating that the proportion of core genes within the pangenome for conspecific strains typically falls between 80% and 90%.
Heuristic methods and genetic distance-based approaches have been successfully applied to delineate cryptic species complexes across multiple fungal genera, including Aspergillus (Glässnerová et al. 2022; Sklenář et al. 2022) and Diaporthe (Dissanayake et al. 2024; Pereira & Phillips 2024). These methods are especially useful when morphological differentiation is subtle, as traditional polyphasic taxonomic approaches often depend significantly on multi-locus phylogenetic analysis in such instances. Furthermore, they assist in circumventing arbitrary taxonomic boundaries. Our findings indicate that species delimitations inferred using the PTP method closely align with those based on the GCPSR framework, suggesting that PTP outperforms mPTP and genetic distance-based methods for species recognition in pestalotiopsis-like taxa fungi. In contrast, over-lumping was observed in analyses employing mPTP, ABGD, and ASAP. Recent studies on Diaporthe have also underscored the propensity of these methods to over-merge species (Dissanayake et al. 2024; Pereira & Phillips 2024). ABGD and ASAP depend on discontinuities in DNA sequence variation, rendering them less effective for species with a strong population genetic structure or low interspecific divergence. When intraspecific variation is high and interspecific differentiation is low, the pairwise distance distributions produced by ABGD and ASAP can prove difficult to interpret. This is likely why species delimitations based on ABGD and ASAP demonstrate high variability and conflict when compared to those based on GCPSR results.
This study consistently demonstrated that species delimitation using GCPSR, heuristic methods, genetic distance-based approaches, and genome analyses revealed a significant number of species within pestalotiopsis-like fungi overestimated. Using a comprehensive approach, a total of seven species were placed in synonymy in Neopestalotiopsis, 18 species were synonymized in Pestalotiopsis, and three species were synonymized in Pseudopestalotiopsis, as outlined in Table 1. The merged species names follow the priority rules of the International Code of Nomenclature for algae, fungi, and plants (Turland et al. 2018). For example, N. zingiberis is the oldest published name in the N. zingiberis lineage. Due to missing sequences, long branch or inconsistent evolutionary placements in the ML analysis and Bayesian analysis for N. pandanicola KUMCC 17-0175, N. anacardii ITCC 6524, and N. termitarii ITCC 6233 in Neopestalotiopsis; Pe. chinensis MFLUCC 12-0273, Pe. matildae CBS 118155, Pe. yunnanensis HMAS 96359, and Pe. sequoiae MFLUCC 13-0399 in Pestalotiopsis; and Ps. kubahensis UMAS-KUB-P20 in Pseudopestalotiopsis, pending further investigation.
In the multi-gene phylogenetic tree, Ps. chinensis is distinctly separated from the Ps. camelliae-sinensis clade (Fig. 26). However, in the whole-genome phylogeny (Fig. 28), Ps. chinensis is nested within Ps. camelliae-sinensis, with an average genome-wide ANI exceeding 98.83% (Fig. 28) and core genes comprising more than 90% of the pangenome. Despite these results, the presence of Ps. vietnamensis NBRC 112252 and Ps. ignota NN 42909 as long-branch taxa in the multi-gene phylogenetic analysis further complicates the delineation of Ps. chinensis and Ps. camelliae-sinensis. This observed discrepancy requires further investigation. If sequencing errors can be ruled out, the inconsistency may stem from unequal evolutionary rates among different lineages. Unfortunately, due to the unavailability of type material, we were unable to generate additional sequences to confirm their taxonomic identities in this study.
Conclusions
Although the number of species in pestalotiopsis-like fungi has increased dramatically over the last decade, this surge is largely attributed to a lack of deep understanding of intra- and interspecific diversification mechanisms within the group. This study demonstrates that intraspecific variation has often been misinterpreted as evidence of new species in pestalotiopsis-like fungi, leading to taxonomic inflation. Our findings emphasize the need for a critical reassessment of the phylogenetic relationships within this group to prevent an overestimation of species diversity. Although this study does not completely clarify the taxonomy of Neopestalotiopsis, the genus Pestalotiopsis has been investigated in greater detail, providing valuable insights into the application of empirical species delimitation methods in a non-model context system. Going forward, a polyphasic approach should be adopted to stabilize the classification of pestalotiopsis-like fungi. Future research ought to build upon the hypotheses presented here, incorporating additional lines of evidence to refine the taxonomy of this group. Further investigations may assist in clarifying species boundaries within pestalotiopsis-like fungi, ensuring a more accurate and comprehensive classification system.
Suggestions for future research on pestalotiopsis-like species include the following:
1) To avoid species over-splitting and underestimation, it is essential to integrate multiple data types and analytical methods for a more comprehensive approach to taxonomy and species identification.
2) For Neopestalotiopsis, the Pestalotiopsis rosea species complex, and the Pseudopestalotiopsis cocos species complex, if a strain does not form a sufficiently long and distinct branch from other species in single-gene phylogenetic trees, and also fails to demonstrate a clearly differentiated branch in the three-gene concatenated tree, we recommend against the introduction of new species, as multiple lines of evidence suggest minimal genetic and protein variation differences.
Acknowledgements
Thanks for the computing support of the State Key Laboratory of Public Big Data, Guizhou University. This research is supported by the following projects: National Natural Science Foundation of China (No. 31972222), the Natural Science Special Research Fund of Guizhou University, Special Post (Gui Da Ling Jun He Zi [2024] 07), Program of Introducing Talents of Discipline to Universities of China (111 Program, D20023), Guizhou Science, Technology Department of International Cooperation Base project ([2018]5806). Kevin D. Hyde and Fatimah Al-Otibi extend their appreciation to the ongoing Research Funding Program (ORF-2026-114), King Saud University, Riyadh, Saudi Arabia. Chitrabhanu S. Bhunjun would like to thank Martin van de Bult, Narong Apichai, and the Doi Tung Development Project for sample collection (permission number 7700/17142 with the title “The diversity of saprobic fungi on selected hosts in forest northern Thailand”), Chiang Rai, Thailand. Chitrabhanu S. Bhunjun would like to thank the National Research Council of Thailand (NRCT) grant ‘Total fungal diversity in a given forest area with implications towards species numbers, chemical diversity and biotechnology (grant number N42A650547). Hong Long acknowledges support from the Guizhou Science and Technology Support Project (Qian [2024]082).
Author Contributions
All authors contributed to the study conception and design. Material preparation was performed by Ya-Ru Sun, Fatimah Al-Otibi, and Shi-Xian Zeng. Data collection and analysis were performed by Qian Zhang, Cheng Li, Chitrabhanu Sharma Bhunjun and Hui Long. The first draft of the manuscript was written by Qian Zhang. Kevin D. Hyde, Yong Wang, Sajeewa S. N. Maharachchikumbura, and Feng-Quan Liu revised the manuscript and guided the analysis of this study. All authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
ORCID
Qian Zhang: https://orcid.org/0009-0003-7937-6982
Yong Wang: https://orcid.org/0000-0003-3831-2117
Feng-Quan Liu: https://orcid.org/0000-0001-9325-1500
Sajeewa S. N. Maharachchikumbura: https://orcid.org/0000-0001-9127-0783
Conflict of Interest Statement
Yong Wang and Sajeewa S. N. Maharachchikumbura are editorial board members of Fungal Diversity. They were not involved in the journal’s review of, or decisions related to, this manuscript. The authors declare no other competing interests.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Supplemental Information
The online version contains supplemental information available at https://doi.org/10.65390/fdiv.2026.136003, Supplementary File to this study.
Rights and permissions
The Author(s) 2026. Published by BioAcademic Press on behalf of Kunming Institute of Botany,Chinese Academy of Sciences (CAS) and Mushroom Research Foundation. This is an open accessarticle under the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0), which permits use, distribution and reproduction in any medium, provided the originalwork is properly cited.
References
- Abbas MF, Batool S, Khan T, Rashid M (2022) First report of Neopestalotiopsis clavispora causing postharvest fruit rot of loquat in Pakistan. Journal of Plant Pathology 104:459. https://doi.org/10.1007/s42161-021-01007-9
- Adams SJ, Walker AK (2023) Diversity of fungi from marine inundated wood from the Bay of Fundy, Nova Scotia, Canada. Botanica Marina 66:319–329. https://doi.org/10.1515/bot-2023-0005
- Adikaram NKB, Maharachchikumbura SSN, Yakandawala DMD, Manawadu LN, Dissanayake DMS, Jayasinghe L (2023) Postharvest stem-end browning (SEB) disease in ripe mango (Mangifera indica L.) cultivar TomEJC. European Journal of Plant Pathology 165:447–464. https://doi.org/10.1007/s10658-022-02616-5
- Adit A, Koul M, Kapoor R, Tandon R (2022) Topological analysis of orchid-fungal endophyte interaction shows lack of phylogenetic preference. South African Journal of Botany 149:339–346. https://doi.org/10.1016/j.sajb.2022.06.025
- Ahmmed MS, Sikder MM, Sarker SR, Hossain MS, Shubhra RD, Alam N (2022) First report of Pseudopestalotiopsis theae causing Aloe vera leaf spot disease in Bangladesh. American Journal of Plant Sciences 13:193–204. https://doi.org/10.4236/ajps.2022.132012
- Ahn GR, Kim BY, Lee GS, Hyun MW, Lee CJ, Kim SH (2016) A report of five unrecorded fungal species of Korea. The Korean Journal of Mycology 44:240–246. https://doi.org/10.4489/KJM.2016.44.4.240
- Aiello D, Gusella G, Leonardi GR, Polizzi G, Voglmayr H (2024) Bottlebrush and myrtle twig canker caused by Neopestalotiopsis species: an emerging canker-causing group of fungi in Italy. Mycokeys 106:133–151. https://doi.org/10.3897/mycokeys.106.121520
- Akinsanmi OA, Nisa S, Jeff-Ego OS, Shivas RG, Drenth A (2017) Dry flower disease of Macadamia in Australia caused by Neopestalotiopsis macadamiae sp. nov. and Pestalotiopsis macadamiae sp. nov. Plant Disease 101:45–53. https://doi.org/10.1094/PDIS-05-16-0630-RE
- Akone SH, El Amrani M, Lin W, Lai D, Proksch P (2013) Cytosporins F–K, new epoxyquinols from the endophytic fungus Pestalotiopsis theae. Tetrahedron Letters 54:6751–6754. https://doi.org/10.1016/j.tetlet.2013.10.005
- Akone SH, Wang H, Mouelle ENM, Mándi A, Kurtán T, Koliye PR, Hartmann R, Bhatia S, Yang J, Müller WEG, Lai D, Liu Z, Kalscheuer R, Proksch P (2022) Prenylated cyclohexene-type meroterpenoids and sulfur-containing xanthones produced by Pseudopestalotiopsis theae. Phytochemistry 197:113124. https://doi.org/10.1016/j.phytochem.2022.113124
- Amezrou R, Ducasse A, Compain J, Lapalu N, Pitarch A, Dupont L, Confais J, Goyeau H, Kema GHJ, Croll D, Amselem J, Sanchez-Vallet A, Marcel TC (2024) Quantitative pathogenicity and host adaptation in a fungal plant pathogen revealed by whole-genome sequencing. Nature Communications 15:1933. https://doi.org/10.1038/s41467-024-46191-1
- Apurillo CCS, Cai L, dela Cruz TEE (2019) Diversity and bioactivities of mangrove fungal endophytes from Leyte and Samar, Philippines. Philippine Science Letters 12:33–48.
- Araujo L, Ferreira Pinto FAM, de Andrade CCL, Mituti T, Falkenbach BR, Gomes LB, Duarte V (2023) Pestalotiopsis trachycarpicola causes leaf spot disease on blueberry in Santa Catarina, Brazil. Australasian Plant Disease Notes 18:14. https://doi.org/10.1007/s13314-023-00500-7
- Ariyawansa HA, Hyde KD (2018) Additions to Pestalotiopsis in Taiwan. Mycosphere 9:999–1013. https://doi.org/10.5943/mycosphere/9/5/4
- Ariyawansa HA, Hyde KD, Jayasiri SC, Buyck B, Chethana KWT, Dai DQ, Dai YC, Daranagama DA, Jayawardena RS, Lücking R, Ghobad-Nejhad M, Niskanen T, Thambugala KM, Voigt K, Zhao RL et al. (2015) Fungal diversity notes 111–252—taxonomic and phylogenetic contributions to fungal taxa. Fungal Diversity 75:27–274. https://doi.org/10.1007/s13225-015-0346-5
- Ashoka GB, Nischitha R, Gagana SL, Shivanna MB (2023) Diversity of endophytic fungal community in Jatropha heynei N.P. Balakr. Journal of Mycopathological Research 61:489–501. https://doi.org/10.57023/JMycR.61.4.2023.489
- Ayoubi N, Soleimani MJ (2016) Strawberry Fruit Rot Caused by Neopestalotiopsis iranensis sp. nov., and N. mesopotamica. Current Microbiology 72:329–336. https://doi.org/10.1007/s00284-015-0955-y
- Ayukawa Y, Asai S, Gan P, Tsushima A, Ichihashi Y, Shibata A, Komatsu K, Houterman PM, Rep M, Shirasu K, Arie T (2021) A pair of effectors encoded on a conditionally dispensable chromosome of Fusarium oxysporum suppress host-specific immunity. Communications Biology 4:707. https://doi.org/10.1038/s42003-021-02245-4
- Azuddin NF, Mohd MH, Rosely NFN, Mansor A, Zakaria L (2021) Molecular phylogeny of endophytic fungi from rattan (Calamus castaneus Griff.) spines and their antagonistic activities against plant pathogenic fungi. Journal of Fungi 7:301. https://doi.org/10.3390/jof7040301
- Bakry M, Bussières G, Lamhamedi MS, Margolis HA, Stowe DC, Abourouh M, Blais M, Bérubé JA (2009) A first record of Pestalotiopsis clavispora in Argan mass cutting propagation: prevalence, prevention and consequences for plant production. Phytoprotection 90:117–120. https://doi.org/10.7202/045780ar
- Banerjee A, Rana D (2020) Leaf spot disease of Syzygium cumini caused by Neopestalotiopsis clavispora newly reported in India. Plant Disease 104:1255. https://doi.org/10.1094/PDIS-09-19-2018-PDN
- Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Pham S, Prjibelski AD (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology 19:455−477. https://doi.org/10.1089/cmb.2012.0021
- Bao W, Kojima KK, Kohany O (2015) Repbase update, a database of repetitive elements in eukaryotic genomes. Mobile DNA 6:11. https://doi.org/10.1186/s13100-015-0041-9
- Barber PA, Crous PW, Groenewald JZ, Pascoe IG, Keane P (2011) Reassessing Vermisporium (Amphisphaeriaceae), a genus of foliar pathogens of eucalypts. Persoonia 27:90–118. https://doi.org/10.3767/003158511X617381
- Barbosa TJA, Feijó MGL, Silva GC, Infante NB, Feijó FM, Assunção IP, Lima GSA (2023) First report of Neopestalotiopsis foedans causing pestalotia spot in leaf on coconut in Brazil. Plant Disease 107:2552. https://doi.org/10.1094/PDIS-12-22-2874-PDN
- Basavand E, Pakdin-Parizi A, Mirhosseini H-A, Dehghan-Niri M (2020) Occurrence of leaf spot disease on date palm caused by Neopestalotiopsis clavispora in Iran. Journal of Plant Pathology 102:959. https://doi.org/10.1007/s42161-020-00530-5
- Belisário R, Aucique‐Pérez CE, Abreu LM, Salcedo SS, de Oliveira WM, Furtado GQ (2020) Infection by Neopestalotiopsis spp. occurs on unwounded eucalyptus leaves and is favoured by long periods of leaf wetness. Plant Pathology 69:194–204. https://doi.org/10.1111/ppa.13132
- Betancourt-Ortiz WF, Medina-Cardenas HC, Padilla-Agudelo JL, Varon FH, Mestizo-Garzón YA, Morales-Rodríguez A, Sarria-Villa GA (2024) Foliar lesions induced by Pestalotiopsis arengae in oil palm (O × G) in the Colombian Southwest Palm Zone. Journal of Fungi 10:24. https://doi.org/10.3390/jof10010024
- Bezerra JDP, Machado AR, Firmino AL, Rosado AWC, de Souza CAF, de Souza-Motta CM, de Sousa Freire KTL, Paiva LM, Magalhães OMC, Pereira OL, Crous PW, de Oliveira TGL, de Abreu VP, Fan X (2018) Mycological diversity description I. Acta Botanica Brasilica 32:656–666. https://doi.org/10.1590/0102-33062018abb0154
- Bhattacharya S, Debnath S, Das P, Saha AK (2019) Diversity of fungal endophyte of Ananus comosu L. var. kew from Unokoti district, tripura with bioactive potential of Neopestalotiopsis piceana. Asian Journal of Pharmacy and Pharmacology 5:353–360. https://doi.org/10.31024/ajpp.2019.5.2.19
- Biju CN, Peeran MF, Gowri R (2018) Identification and characterization of Neopestalotiopsis clavispora associated with leaf blight of small cardamom (Elettaria cardamomum Maton). Journal of Phytopathology 166:532–546. https://doi.org/10.1111/jph.12715
- Blagojević J, Aleksić G, Vučurović I, Starović M, Ristić D (2024) Exploring the phylogenetic diversity of Botryosphaeriaceae and Diaporthe species causing dieback and shoot blight of blueberry in Serbia. Phytopathology 114:1333–1345. https://doi.org/10.1094/PHYTO-04-23-0133-R
- Blanco E, Parra G, Guigó R (2007) Using geneid to identify genes. Current Protocols in Bioinformatics:4.3.1–4.3.28. https://doi.org/10.1002/0471250953.bi0403s18
- Boeckmann B, Bairoch A, Apweiler R, Blatter MC, Estreicher A, Gasteiger E, Martin MJ, Michoud K, O'Donovan C, Phan I, Pilbout S, Schneider M (2003) The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Research 31:365−370. https://doi.org/10.1093/nar/gkg095
- Bonthond G, Sandoval-Denis M, Groenewald JZ, Crous PW (2018) Seiridium (Sporocadaceae): an important genus of plant pathogenic fungi. Persoonia 40:96–118. https://doi.org/10.3767/persoonia.2018.40.04
- Borgohain P, Barua P, Mahanta J, Saikia LR (2020) Pestalotioid fungi: a rare agent of onychomycosis among agriculture workers. Current Medical Mycology 6:23–29. https://doi.org/10.18502/CMM.6.2.2839
- Borrero C, Castaño R, Avilés M (2018) First report of Pestalotiopsis clavispora (Neopestalotiopsis clavispora) causing canker and twig dieback on blueberry bushes in Spain. Plant Disease 102:1178. https://doi.org/10.1094/PDIS-10-17-1529-PDN
- Broders K, Munck I, Wyka S, Iriarte G, Beaudoin E (2015) Characterization of fungal pathogens associated with white pine needle damage (WPND) in northeastern North America. Forests 6:4088–4104. https://doi.org/10.3390/f6114088
- Bunyapaiboonsri T, Yoiprommarat S, Nithithanasilp S, Choowong W, Preedanon S, Suetrong S (2023) Two new farnesyl hydroquinones from Pestalotiopsis diploclisia (BCC 35283), the fungus associated with algae. Natural Product Research 37:24–30. https://doi.org/10.1080/14786419.2021.1946536
- Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
- Cardoso JKB, Nanami DSY, Zanutto CA, Grabicoski EMG, Vida JB, Bock CH, Nunes WMC (2017) Description and identification of two new diseases of guariroba palm (Syagrus oleraceae) in Brazil. Journal of Phytopathology 165:610–619. https://doi.org/10.1111/jph.12599
- Carvalho CR, Gonçalves VN, Pereira CB, Johann S, Galliza IV, Alves TMA, Rabello A, Sobral MEG, Zani CL, Rosa CA, Rosa LH (2012) The diversity, antimicrobial and anticancer activity of endophytic fungi associated with the medicinal plant Stryphnodendron adstringens (Mart.) Coville (Fabaceae) from the Brazilian savannah. Symbiosis 57:95–107. https://doi.org/10.1007/s13199-012-0182-2
- Catarino AdM, Hanada RE, de Queiroz CA, Sousa TF, Lima ÍN, Gasparotto L, da Silva GF (2020) First report of Pseudopestalotiopsis dawaina causing spots in Caryota mitis in Brazil. Plant Disease 104:989. https://doi.org/10.1094/PDIS-08-19-1771-PDN
- Chaiwan N, Jeewon R, Pem D, Jayawardena RS, Nazurally N, Mapook A, Promputtha I, Hyde KD (2022) Fungal species from Rhododendron sp.: Discosia rhododendricola sp. nov, Neopestalotiopsis rhododendricola sp.nov and Diaporthe nobilis as a new host record. Journal of Fungi 8:907. https://doi.org/10.3390/jof8090907
- Chaiwan N, Wanasinghe DN, Mapook A, Jayawardena RS, Norphanphoun C, Hyde KD (2020) Novel species of Pestalotiopsis fungi on Dracaena from Thailand. Mycology 11:306–315. https://doi.org/10.1080/21501203.2020.1801873
- Chamorro M, Aguado A, De los Santos B (2016) First report of root and crown rot caused by Pestalotiopsis clavispora (Neopestalotiopsis clavispora) on strawberry in Spain. Plant Disease 100:1495. https://doi.org/10.1094/PDIS-11-15-1308-PDN
- Chen S, Zhou Y, Chen Y, Gu J (2018a) Fastp: an ultra-fast all-in-one FASTQ preprocessor. Bioinformatics 34:i884–i890. https://doi.org/10.1093/bioinformatics/bty560
- Chen TG, Wang X, Hu JC, Yang QQ, Liang WX (2019) First report of Neopestalotiopsis clavispora causing leaf spots on Ligustrum lucidum in China. Plant Disease 103:1034. https://doi.org/10.1094/PDIS-06-18-0988-PDN
- Chen Y, Zeng L, Shu N, Jiang M, Wang H, Huang Y, Tong H (2018c) Pestalotiopsis-like species causing gray blight disease on Camellia sinensis in China. Plant Disease 102:98–106. https://doi.org/10.1094/PDIS-05-17-0642-RE
- Chen YJ, Zeng L, Meng Q, Tong HR (2018b) Occurrence of Pestalotiopsis lushanensis causing grey blight disease on Camellia sinensis in China. Plant Disease 102:2654. https://doi.org/10.1094/PDIS-04-18-0640-PDN
- Chen YY, Maharachchikumbura SSN, Liu JK, Hyde KD, Nanayakkara RR, Zhu GS, Liu ZY (2017) Fungi from Asian Karst formations I. Pestalotiopsis photinicola sp. nov., causing leaf spots of Photinia serrulata. Mycosphere 8:103–110. https://doi.org/10.5943/mycosphere/8/1/9
- Chock MK, Hoyt BK, Amend AS (2021) Mycobiome transplant increases resistance to Austropuccinia psidii in an endangered Hawaiian plant. Phytobiomes Journal 5:326–334. https://doi.org/10.1094/PBIOMES-09-20-0065-R
- Choi YJ, Eom H, Park J, Park J, Cheon S, Ro HS (2024) Fungal diversity in Nam River and their biodegradative activities. Mycobiology 52:102–110. https://doi.org/10.1080/12298093.2024.2324575
- Conesa A, Götz S, García-Gómez JM, Terol J, Talón M, Robles M (2005) Blast2GO: a universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 21:3674–3676. https://doi.org/10.1093/bioinformatics/bti610
- Conforto C, Lima NB, Silva FJA, Câmara MPS, Maharachchikumbura S, Michereff SJ (2019) Characterization of fungal species associated with cladode brown spot on Nopalea cochenillifera in Brazil. European Journal of Plant Pathology 155:1179–1194. https://doi.org/10.1007/s10658-019-01847-3
- Crous PW, Boers J, Holdom D, Osieck ER, Steinrucken TV, Tan YP, Vitelli JS, Shivas RG, Barrett M, Boxshall AG, Broadbridge J, Larsson E, Lebel T, Pinruan U, Sommai S et al. (2022) Fungal Planet description sheets: 1383–1435. Persoonia 48:261–371. https://doi.org/10.3767/persoonia.2022.48.08
- Crous PW, Gams W, Stalpers JA, Robert V, Stegehuis G (2004) MycoBank: an online initiative to launch mycology into the 21st century. Studies in Mycology 50:19–22.
- Crous PW, Schumacher RK, Wingfield MJ, Akulov A, Denman S, Roux J, Braun U, Burgess TI, Carnegie AJ, Váczy KZ, Guatimosim E, Schwartsburd PB, Barreto RW, Hernández-Restrepo M, Lombard L, Groenewald JZ (2018) New and interesting fungi. 1. Fungal Systematics and Evolution 1:169–216. https://doi.org/10.3114/fuse.2018.01.08
- Crous PW, Summerell BA, Swart L, Denman S, Taylor JE, Bezuidenhout CM, Palm ME, Marincowitz S, Groenewald JZ (2011) Fungal pathogens of Proteaceae. Persoonia 27:20–45. https://doi.org/10.3767/003158511X606239
- Crous PW, Verkley GJM, Christensen M, Castañeda-Ruiz RF, Groenewald JZ (2012) How important are conidial appendages? Persoonia 28:126–137. https://doi.org/10.3767/003158512X652624
- Crous PW, Verkley GJM, Groenewald JZ (2006) Eucalyptus microfungi known from culture. 1. Cladoriella and Fulvoflamma genera nova, with notes on some other poorly known taxa. Studies in Mycology 55:53–63. https://doi.org/10.3114/sim.55.1.53
- Crous PW, Wingfield MJ, Burgess TI, Carnegie AJ, Hardy GESJ, Smith D, Summerell BA, Cano-Lira JF, Guarro J, Houbraken J, Lombard L, Martín MP, Sandoval-Denis M, Alexandrova AV, Barnes CW et al. (2017) Fungal Planet description sheets: 625–715. Persoonia 39:270–467. https://doi.org/10.3767/persoonia.2017.39.11
- Crous PW, Wingfield MJ, Chooi YH, Gilchrist CLM, Lacey E, Pitt JI, Roets F, Swart WJ, Cano-Lira JF, Valenzuela-Lopez N, Hubka V, Shivas RG, Stchigel AM, Holdom DG, Jurjević Ž et al. (2020) Fungal Planet description sheets: 1042–1111. Persoonia 44:301–459. https://doi.org/10.3767/persoonia.2020.44.11
- Crous PW, Wingfield MJ, Le Roux JJ, Richardson DM, Strasberg D, Shivas RG, Alvarado P, Edwards J, Moreno G, Sharma R, Sonawane MS, Tan YP, Altés A, Barasubiye T, Barnes CW et al. (2015) Fungal Planet description sheets: 371–399. Persoonia 35:264–327. https://doi.org/10.3767/003158515x690269
- Cui X, Hao Z, Chen M, Song S, Zhang J, Li Y, Li J, Liu Y, Luo L (2024) Identification and pathogenicity of pestalotioid species on Alpinia oxyphylla in Hainan province, China. Journal of Fungi 10:371. https://doi.org/10.3390/jof10060371
- Daengsuwan W, Wonglom P, Arikit S, Sunpapao A (2021) Morphological and molecular identification of Neopestalotiopsis clavispora causing flower blight on Anthurium andraeanum in Thailand. Horticultural Plant Journal 7:573–578. https://doi.org/10.1016/j.hpj.2020.10.004
- Darapanit A, Boonyuen N, Leesutthiphonchai W, Nuankaew S, Piasai O (2021) Identification, pathogenicity and effects of plant extracts on Neopestalotiopsis and Pseudopestalotiopsis causing fruit diseases. Scientific Reports 11:22606. https://doi.org/10.1038/s41598-021-02113-5
- Das K, Lee SY, Jung HY (2021) Pestalotiopsis kaki sp. nov., a novel species isolated from persimmon tree (Diospyros kaki) bark in Korea. Mycobiology 49:54–60. https://doi.org/10.1080/12298093.2020.1852703
- Das M, Royer TV, Leff LG (2008) Fungal communities on decaying leaves in streams: a comparison of two leaf species. Mycological Progress 7:267–275. https://doi.org/10.1007/s11557-008-0569-x
- de Oliveira Amaral A, e Ferreira AFTAF, da Silva Bentes JL (2022) Fungal endophytic community associated with Hevea spp.: diversity, enzymatic activity, and biocontrol potential. Brazilian Journal of Microbiology 53:857–872. https://doi.org/10.1007/s42770-022-00709-1
- de Silva NI, Maharachchikumbura SSN, Thambugala KM, Bhat DJ, Karunarathna SC, Tennakoon DS, Phookamsak R, Jayawardena RS, Lumyong S, Hyde KD (2021) Morpho-molecular taxonomic studies reveal a high number of endophytic fungi from Magnolia candolli and M. garrettii in China and Thailand. Mycosphere 12:163–237. https://doi.org/10.5943/mycosphere/12/1/3
- Deyrup ST, Swenson DC, Gloer JB, Wicklow DT (2006) Caryophyllene sesquiterpenoids from a fungicolous isolate of Pestalotiopsis disseminata. Journal of Natural Products 69:608–611. https://doi.org/10.1021/np050460b
- Ding S, Hu H, Gu JD (2015) Fungi colonizing wood sticks of Chinese fir incubated in subtropical urban soil growing with Ficus microcarpa trees. International Journal of Environmental Science and Technology 12:3781–3790. https://doi.org/10.1007/s13762-015-0802-5
- Diogo E, Gonçalves CI, Silva AC, Valente C, Bragança H, Phillips AJL (2021) Five new species of Neopestalotiopsis associated with diseased Eucalyptus spp. in Portugal. Mycological Progress 20:1441–1456. https://doi.org/10.1007/s11557-021-01741-5
- Dissanayake AJ, Zhu JT, Chen YY, Maharachchikumbura SSN, Hyde KD, Liu JK (2024) A re-evaluation of Diaporthe: refining the boundaries of species and species complexes. Fungal Diversity 126:1–125. https://doi.org/10.1007/s13225-024-00538-7
- Dong W, Hyde KD, Jeewon R, Liao CF, Zhao HJ, Kularathnage ND, Li H, Yang YH, Pem D, Shu YX, Gafforov Y, Manawasinghe IS, Doilom M (2023) Mycosphere notes 449–468: Saprobic and endophytic fungi in China, Thailand, and Uzbekistan. Mycosphere 14:2208–2262. https://doi.org/10.5943/mycosphere/14/1/26
- dos Reis JBA, Pappas Junior GJ, Lorenzi AS, Pinho DB, Costa AM, da Cunha Bustamante MM, do Vale HMM (2023) How deep can the endophytic mycobiome go? A case study on six woody species from the Brazilian Cerrado. Journal of Fungi 9:508. https://doi.org/10.3390/jof9050508
- Douanla-Meli C, Langer E (2009) Pestalotiopsis theae (Ascomycota, Amphisphaeriaceae) on seeds of Diospyros crassiflora (Ebenaceae). Mycotaxon 107:441–448.
- Douanla-Meli C, Langer E, Mouafo FT (2013) Fungal endophyte diversity and community patterns in healthy and yellowing leaves of Citrus limon. Fungal Ecology 6:212–222. https://doi.org/10.1016/j.funeco.2013.01.004
- Duin IM, Higa AR, dos Santos ÁF, Coelho TAdV, Rezende EH, Auer CG (2017) Etiology and possible inoculum sources for the occurrence of rot in black wattle minicuttings. Summa Phytopathologica 43:297–302. https://doi.org/10.1590/0100-5405/174647
- Ebrahimi L, Hatami Rad S, Etebarian HR (2022) Apple endophytic fungi and their antagonism against apple scab disease. Frontiers in Microbiology 13:1024001. https://doi.org/10.3389/fmicb.2022.1024001
- Elfiati D, Faulina SA, Rahayu LM, Aryanto A, Dewi RT, Rachmat HH, Turjaman M, Royyani MF, Susilowati A, Hidayat A (2022) Culturable endophytic fungal assemblages from Styrax sumatrana and Stryax benzoin and their potential as antifungal, antioxidant, and alpha-glucosidase inhibitory resources. Frontiers in Microbiology 13:974526. https://doi.org/10.3389/fmicb.2022.974526
- Emms DM, Kelly S (2015) OrthoFinder: solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biology 16:157. https://doi.org/10.1186/s13059-015-0721-2
- Erdurmuş D, Palacıoğlu G, Erdurmuş G, Bayraktar H (2023) First report of Neopestalotiopsis rosae causing leaf spot and crown rot of strawberry in Turkey. Journal of Plant Pathology 105:315. https://doi.org/10.1007/s42161-022-01218-8
- Escudero-Leyva E, Vieto S, Avendaño R, Rojas-Gätjens D, Agüero P, Pacheco C, Montero ML, Chaverri P, Chavarría M (2023) Fungi with history: unveiling the mycobiota of historic documents of Costa Rica. PLoS ONE 18:e0279914. https://doi.org/10.1371/journal.pone.0279914
- Essa TA, Kamel SM, Ismail AM, El-Ganainy SM (2018) Characterization and chemical control of Neopestalotiopsis rosae the causal agent of strawberry root and crown rot in Egypt. Egyptian Journal of Phytopathology 46:1–19. https://doi.org/10.21608/ejp.2018.87411
- Fan J, Qiu HB, Long HJ, Zhao W, Xiang XL, Hu AL (2021) First report of leaf spot on sorghum caused by Pestalotiopsis trachycarpicola in China. Journal of Plant Pathology 103:1043-1044. https://doi.org/10.1007/s42161-021-00853-x
- Fan M, Chen X, Luo X, Zhang H, Liu Y, Zhang Y, Wu J, Zhao C, Zhao P (2020) Diversity of endophytic fungi from the leaves of Vaccinium dunalianum. Letters in Applied Microbiology 71:479–489. https://doi.org/10.1111/lam.13345
- Fan S, Su J, Fang T, Pan A, He X, Fan G, Wang Z, Hu H (2022) Pathogen of Liquidambar formosana leaf spot. Mycosystema 41:420–434. https://doi.org/10.13346/j.mycosystema.210375
- Fan SB, Liu XG, Su JY, Hu YP, Jia Y, Liang XY, Wang ZH, Hu HL (2023) Pestalotiopsis hispanica, a new pathogenic fungus responsible for Cryptomeria japonica red blight. Acta Phytopathologica Sinica 53:321–325. https://doi.org/10.13926/j.cnki.apps.000815
- Feng CT, Du X, Wee J (2021) Microbial and chemical analysis of non-Saccharomyces yeasts from Chambourcin hybrid grapes for potential use in winemaking. Fermentation 7:15. https://doi.org/10.3390/fermentation7010015
- Fernández-Ozuna YA, Gini Álvarez AR, Lopez-Nicora HD, Arrúa Alvarenga AA, Colmán AA (2023) First report of Neopestalotiopsis rosae causing leaf spot and crown rot on strawberry (Fragaria× ananassa) in Paraguay. New Disease Reports 48:e12239. https://doi.org/10.1002/ndr2.12239
- Ferreira MC, Cantrell CL, Wedge DE, Gonçalves VN, Jacob MR, Khan S, Rosa CA, Rosa LH (2017) Diversity of the endophytic fungi associated with the ancient and narrowly endemic neotropical plant Vellozia gigantea from the endangered Brazilian rupestrian grasslands. Biochemical Systematics and Ecology 71:163–169. https://doi.org/10.1016/j.bse.2017.02.006
- Finn RD, Clements J, Eddy SR (2011) HMMER web server: interactive sequence similarity searching. Nucleic Acids Research 39:W29–W37. https://doi.org/10.1093/nar/gkr367
- Fiorenza A, Gusella G, Aiello D, Polizzi G, Voglmayr H (2022) Neopestalotiopsis siciliana sp. nov. and N. rosae causing stem lesion and dieback on avocado plants in Italy. Journal of Fungi 8:562. https://doi.org/10.3390/jof8060562
- Flynn JM, Hubley R, Goubert C, Rosen J, Clark AG, Feschotte C, Smit AF (2020) RepeatModeler2 for automated genomic discovery of transposable element families. Proceedings of the National Academy of Sciences 117: 9451−9457. https://doi.org/10.1073/pnas.1921046117
- Franić I, Prospero S, Hartmann M, Allan E, Auger‐Rozenberg MA, Grünwald NJ, Kenis M, Roques A, Schneider S, Sniezko R, Williams W, Eschen R (2019) Are traded forest tree seeds a potential source of nonnative pests? Ecological Applications 29:e01971. https://doi.org/10.1002/eap.1971
- Freitas EFS, Da Silva M, Barros MVP, Kasuya MCM (2019) Neopestalotiopsis hadrolaeliae sp. nov., a new endophytic species from the roots of the endangered orchid Hadrolaelia jongheana in Brazil. Phytotaxa 416:211–220. https://doi.org/10.11646/phytotaxa.416.3.2
- Geng K, Zhang B, Song Y, Hyde KD, Kang JC, Wang Y (2013) A new species of Pestalotiopsis from leaf spots of Licuala grandis from Hainan, China. Phytotaxa 88:49–54. https://doi.org/10.11646/phytotaxa.88.3.2
- Ghuffar S, Irshad G, Naz F, Zhang XY, Bashir A, Yang HL, Zhai FY, Gleason ML (2018) First report of postharvest rot caused by Pestalotiopsis sp. on grapes in Punjab, Pakistan. Plant Disease 102:1175. https://doi.org/10.1094/PDIS-08-17-1281-PDN
- Gilardi G, Bergeretti F, Gullino ML, Garibaldi A (2019) First report of Neopestalotiopsis clavispora causing root and crown rot on strawberry in Italy. Plant Disease 103:2959. https://doi.org/10.1094/PDIS-03-19-0673-PDN
- Glass NL, Donaldson GC (1995) Development of primer sets designed for use with the PCR to amplify conserved genes from filamentous ascomycetes. Applied and Environmental Microbiology 61:1323−1330.
- Glässnerová K, Sklenář F, Jurjević Ž, Houbraken J, Yaguchi T, Visagie CM, Gené J, Siqueira JPZ, Kubátová A, Kolařík M, Hubka V (2022) A monograph of Aspergillus section Candidi. Studies in Mycology 102:1–51. https://doi.org/10.3114/sim.2022.102.01
- Godinho VM, de Paula MTR, Silva DAS, Paresque K, Martins AP, Colepicolo P, Rosa CA, Rosa LH (2019) Diversity and distribution of hidden cultivable fungi associated with marine animals of Antarctica. Fungal Biology 123:507–516. https://doi.org/10.1016/j.funbio.2019.05.001
- Goshima G (2022) Growth and division mode plasticity is dependent on cell density in marine‐derived black yeasts. Genes to Cells 27:124–137. https://doi.org/10.1111/gtc.12916
- Gotting K, May DS, Sosa-Calvo J, Khadempour L, Francoeur CB, Berasategui A, Thairu MW, Sandstrom S, Carlson CM, Chevrette MG, Pupo MT, Bugni TS, Schultz TR, Johnston JS, Gerardo NM, Currie CR (2022) Genomic diversification of the specialized parasite of the fungus-growing ant symbiosis. Proceedings of the National Academy of Sciences 119:e2213096119. https://doi.org/10.1073/pnas.2213096119
- Gou Y, Sun S, Gao S, Xue C, Liu S, Tian T, Wen S, Meng Q (2023) First report of Pestalotiopsis clavispora causing leaf spot on Pandanus amaryllifolius in China. Plant Disease 107:2853. https://doi.org/10.1094/PDIS-02-23-0302-PDN
- Gu M, Hu DW, Han B, Jiang N, Tian CM (2021) Pestalotiopsis abietis sp. nov. from Abies fargesii in China. Phytotaxa 509:93–105. https://doi.org/10.11646/phytotaxa.509.1.4
- Gu R, Bao DF, Shen HW, Su XJ, Li YX, Luo ZL (2022) Endophytic Pestalotiopsis species associated with Rhododendron in Cangshan Mountain, Yunnan Province, China. Frontiers in Microbiology 13:1016782. https://doi.org/10.3389/fmicb.2022.1016782
- Gualberto GF, Catarino AdM, Sousa TF, Cruz JC, Hanada RE, Caniato FF, Silva GF (2021) Pseudopestalotiopsis gilvanii sp. nov. and Neopestalotiopsis formicarum leaves spot pathogens from guarana plant: a new threat to global tropical hosts. Phytotaxa 489:121–139. https://doi.org/10.11646/phytotaxa.489.2.2
- Guba EF (1961) Monograph of Pestalotia and Monochaetia. Harvard University Press, Cambridge.
- Guterres DC, Silva MA, Martins MD, Azevedo DMQ, Lisboa DO, Pinho DB, Furtado GQ (2023) Leaf spot caused by Neopestalotiopsis species on Arecaceae in Brazil. Australasian Plant Pathology 52:47–62. https://doi.org/10.1007/s13313-022-00893-6
- Gutiérrez EE, Garbino GST (2018) Species delimitation based on diagnosis and monophyly, and its importance for advancing mammalian taxonomy. Zoological Research 39:301–308. https://doi.org/10.24272/j.issn.2095-8137.2018.037
- Haas BJ, Salzberg SL, Zhu W, Pertea M, Allen JE, Orvis J, White O, Buell CR, Wortman JR (2008) Automated eukaryotic gene structure annotation using EVidenceModeler and the Program to Assemble Spliced Alignments. Genome Biology 9:R7. https://doi.org/10.1186/gb-2008-9-1-r7
- Hamzah TNT, Lee SY, Hidayat A, Terhem R, Faridah-Hanum I, Mohamed R (2018) Diversity and characterization of endophytic fungi isolated from the tropical mangrove species, Rhizophora mucronata, and identification of potential antagonists against the soil-borne fungus, Fusarium solani. Frontiers in Microbiology 9:01707. https://doi.org/10.3389/fmicb.2018.01707
- Han Y, Wessler SR (2010) MITE-Hunter: a program for discovering miniature inverted-repeat transposable elements from genomic sequences. Nucleic Acids Research 38:e199. https://doi.org/10.1093/nar/gkq862
- Haq IU, Ijaz S, Khan NA (2021) Genealogical concordance of phylogenetic species recognition-based delimitation of Neopestalotiopsis species associated with leaf spots and fruit canker disease affected guava plants. Pakistan Journal of Agricultural Sciences 58:1301–1313. https://doi.org/10.21162/PAKJAS/21.1045
- Hassan SHA, Koutb M, Nafady NA, Hassan EA (2018) Potentiality of Neopestalotiopsis clavispora ASU1 in biosorption of cadmium and zinc. Chemosphere 202:750–756. https://doi.org/10.1016/j.chemosphere.2018.03.114
- He YK, Yang Q, Sun YR, Zeng XY, Jayawardena RS, Hyde KD, Wang Y (2022) Additions to Neopestalotiopsis (Amphisphaeriales, Sporocadaceae) fungi: two new species and one new host record from China. Biodiversity Data Journal 10:e90709. https://doi.org/10.3897/BDJ.10.e90709
- Hermawan R, Safitri RR, Sidiq MR (2021) Neopestalotiopsis zimbabwana isolated from Xylaria stromata. Journal of Tropical Biology 9:203–209. https://doi.org/10.21776/ub.biotropika.2021.009.03.04
- Hidayat A, Turjaman M, Faulina SA, Ridwan F, Aryanto, Najmulah, Irawadi TT, Iswanto AH (2019) Antioxidant and antifungal activity of endophytic fungi associated with agarwood trees. Journal of the Korean Wood Science and Technology 47:459–471. https://doi.org/10.5658/WOOD.2019.47.4.459
- Hidrobo-Chavez J, Ramírez-Villacís DX, Barriga-Medina N, Herrera K, León-Reyes A (2022) First report of Neopestalotiopsis mesopotamica causing root and crown rot on strawberry in Ecuador. Plant Disease 106:1066. https://doi.org/10.1094/PDIS-06-21-1278-PDN
- Hlaiem S, Yangui I, Della Rocca G, Barberini S, Danti R, Ben Jamâa ML (2022) First report of Pestalotiopsis biciliata causing dieback on Quercus coccifera and Pistacia lentiscus in Tunisia. Canadian Journal of Plant Pathology 44:534–541. https://doi.org/10.1080/07060661.2022.2032831
- Hlaiem S, Zouaoui-Boutiti M, Ben Jemâa ML, Della Rocca G, Barberini S, Danti R (2018) Identification and pathogenicity of Pestalotiopsis chamaeropis, causal agent of white heather (Erica arborea) dieback, and in vitro biocontrol with the antagonist Trichoderma sp. Tunisian Journal of Plant Protection 13:49–60.
- Hoede C, Arnoux S, Moisset M, Chaumier T, Inizan O, Jamilloux V, Quesneville H (2014) PASTEC: an automatic transposable element classification tool. PLoS ONE 9:e91929. https://doi.org/10.1371/journal.pone.0091929
- Hofstetter V, Miadlikowska J, Kauff F, Lutzoni F (2007) Phylogenetic comparison of protein-coding versus ribosomal RNA-coding sequence data: a case study of the Lecanoromycetes (Ascomycota). Molecular Phylogenetics and Evolution 44:412–426. https://doi.org/10.1016/j.ympev.2006.10.016
- Hsu SY, Lin YC, Xu YC, Chang HX, Chung PC, Ariyawansa HA (2022) High-quality genome assembly of Neopestalotiopsis rosae ML1664, the pathogen causing strawberry leaf blight and crown rot. Molecular Plant-Microbe Interactions 35:949–953. https://doi.org/10.1094/MPMI-04-22-0077-A
- Hsu SY, Xu YC, Lin YC, Chuang WY, Lin SR, Stadler M, Tangthirasunun N, Cheewangkoon R, Al-Shwaiman HA, Elgorban AM, Ariyawansa HA (2024) Hidden diversity of Pestalotiopsis and Neopestalotiopsis (Amphisphaeriales, Sporocadaceae) species allied with the stromata of entomopathogenic fungi in Taiwan. MycoKeys 101:275–312. https://doi.org/10.3897/mycokeys.101.113090
- Hu H, Jeewon R, Zhou D, Zhou T, Hyde KD (2007) Phylogenetic diversity of endophytic Pestalotiopsis species in Pinus armandii and Ribes spp.: evidence from rDNA and β-tubulin gene phylogenies. Fungal Diversity 24:1–22.
- Huanaluek N, Jayawardena RS, Maharachchikumbura SSN, Harishchandra DL (2021) Additions to pestalotioid fungi in Thailand: Neopestalotiopsis hydeana sp. nov. and Pestalotiopsis hydei sp. nov. Phytotaxa 479:23–43. https://doi.org/10.11646/phytotaxa.479.1.2
- Huang Y, Wang H, Huo S, Lu J, Norvienyeku J, Miao W, Qin C, Liu W (2024) Comparative mitogenomics analysis revealed evolutionary divergence among Neopestalotiopsis species complex (fungi: Xylariales). International Journal of Molecular Sciences 25:3093. https://doi.org/10.3390/ijms25063093
- Huelsenbeck JP, Ronquist F (2001) MRBAYES: bayesian inference of phylogenetic trees. Bioinformatics 17:754–755.
- Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Molecular Biology and Evolution 23:254–267.
- Hyde KD, Dong Y, Phookamsak R, Jeewon R, Bhat DJ, Jones EBG, Liu N-G, Abeywickrama PD, Mapook A, Wei D, Perera RH, Manawasinghe IS, Pem D, Bundhun D, Karunarathna A et al. (2020a) Fungal diversity notes 1151–1276: taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Diversity 100:5–277. https://doi.org/10.1007/s13225-020-00439-5
- Hyde KD, Hongsanan S, Jeewon R, Bhat DJ, McKenzie EHC, Jones EBG, Phookamsak R, Ariyawansa HA, Boonmee S, Zhao Q, Abdel-Aziz FA, Abdel-Wahab MA, Banmai S, Chomnunti P, Cui BK et al. (2016) Fungal diversity notes 367–490: taxonomic and phylogenetic contributions to fungal taxa. Fungal Diversity 80:1–270. https://doi.org/10.1007/s13225-016-0373-x
- Hyde KD, Jeewon R, Chen YJ, Bhunjun CS, Calabon MS, Jiang HB, Lin CG, Norphanphoun C, Sysouphanthong P, Pem D, Tibpromma S, Zhang Q, Doilom M, Jayawardena RS, Liu JK et al. (2020b) The numbers of fungi: is the descriptive curve flattening? Fungal Diversity 103:219–271. https://doi.org/10.1007/s13225-020-00458-2
- Hyde KD, Norphanphoun C, Ma J, Yang HD, Zhang JY, Du TY, Gao Y, Gomes de Farias AR, Gui H, He SC, He YK, Li CJY, Liu XF, Lu L, Su HL et al. (2023) Mycosphere notes 387–412 – novel species of fungal taxa from around the world. Mycosphere 14:663–744. https://doi.org/10.5943/mycosphere/14/1/8
- Hyde KD, Norphanphoun C, Maharachchikumbura SSN, Bhat DJ, Jones EBG, Bundhun D, Chen YJ, Bao DF, Boonmee S, Calabon MS, Chaiwan N, Chethana KWT, Dai DQ, Dayarathne MC, Devadatha B et al. (2020c) Refined families of Sordariomycetes. Mycosphere 11:305–1059. https://doi.org/10.5943/mycosphere/11/1/7
- Hyde KD, Wijesinghe SN, Afshari N, Aumentado HD, Bhunjun CS, Boonmee S, Camporesi E, Chethana KWT, Doilom M, Dong W, Du TY, Farias ARG, Gao Y, Jayawardena RS et al. (2024) Mycosphere Notes 469–520. Mycosphere 15:1294–1454. https://doi.org/10.5943/mycosphere/15/1/11
- Ismail SI, Zulperi D, Jamian S, Mohd Hata E (2022) First report of Neopestalotiopsis saprophytica causing leaf blight on royal palm in Malaysia. Journal of Plant Pathology 104:869–870. https://doi.org/10.1007/s42161-022-01071-9
- Ismail SI, Zulperi D, Norddin S, Ahmad-Hamdani S (2017) First report of Neopestalotiopsis saprophytica causing leaf spot of oil palm (Elaeis guineensis) in Malaysia. Plant Disease 101:1821. https://doi.org/10.1094/PDIS-02-17-0271-PDN
- Jain C, Rodriguez-R LM, Phillippy AM, Konstantinidis KT, Aluru S (2018) High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nature Communications 9:5114. https://doi.org/10.1038/s41467-018-07641-9
- Jayawardena RS, Hyde KD, Chethana KWT, Daranagama DA, Dissanayake AJ, Goonasekara ID, Manawasinghe IS, Mapook A, Jayasiri SC, Karunarathna A, Li CG, Phukhamsakda C, Senanayake IC, Wanasinghe DN, Camporesi E, Bulgakov TS, Li XH, Liu M, Zhang W, Yan JY (2018) Mycosphere notes 102–168: saprotrophic fungi on Vitis in China, Italy, Russia and Thailand. Mycosphere 9:1–114. https://doi.org/10.5943/mycosphere/9/1/1
- Jayawardena RS, Hyde KD, McKenzie EHC, Jeewon R, Phillips AJL, Perera RH, de Silva NI, Maharachchikumburua SSN, Samarakoon MC, Ekanayake AH, Tennakoon DS, Dissanayake AJ, Norphanphoun C, Lin C, Manawasinghe IS et al. (2019) One stop shop III: taxonomic update with molecular phylogeny for important phytopathogenic genera: 51–75 (2019). Fungal Diversity 98:77–160. https://doi.org/10.1007/s13225-019-00433-6
- Jayawardena RS, Hyde KD, Wang S, Sun YR, Suwannarach N, Sysouphanthong P, Abdel-Wahab MA, Abdel-Aziz FA, Abeywickrama PD, Abreu VP, Armand A, Aptroot A, Bao DF, Begerow D, Bellanger JM et al. (2022) Fungal diversity notes 1512–1610: taxonomic and phylogenetic contributions on genera and species of fungal taxa. Fungal Diversity 117:1–272. https://doi.org/10.1007/s13225-022-00513-0
- Jayawardena RS, Liu M, Maharachchikumbura SSN, Zhang W, Xing Q, Hyde KD, Nilthong S, Li X, Yan J (2016) Neopestalotiopsis vitis sp. nov. causing grapevine leaf spot in China. Phytotaxa 258:63–74. https://doi.org/10.11646/phytotaxa.258.1.4
- Jayawardena RS, Zhang W, Liu M, Maharachchikumbura SSN, Zhou Y, Huang J, Nilthong S, Wang Z, Li X, Yan J, Hyde KD (2015) Identification and characterization of pestalotiopsis-like fungi related to grapevine diseases in China. Fungal Biology 119:348–361. https://doi.org/10.1016/j.funbio.2014.11.001
- Jevremović D, Vasić T, Živković S, Vasilijević B, Marić M, Vojvodić M, Bulajić A (2022) Neopestalotiopsis clavispora: a causal agent of twig dieback on highbush blueberries in Serbia. Journal of Plant Diseases and Protection 129:1277–1283. https://doi.org/10.1007/s41348-022-00610-x
- Jiang N, Bonthond G, Fan XL, Tian CM (2018) Neopestalotiopsis rosicola sp. nov. causing stem canker of Rosa chinensis in China. Mycotaxon 133:271–283. https://doi.org/10.5248/133.271
- Jiang N, Fan X, Tian C (2021) Identification and characterization of leaf-inhabiting fungi from Castanea plantations in China. Journal of Fungi 7:64. https://doi.org/10.3390/jof7010064
- Jiang N, Tian LY, Xue H, Piao CG, Li Y (2023) Pestalotiopsis sonneratiae sp. nov. from China. Mycotaxon 137:725–735. https://doi.org/10.5248/137.725
- Jiang N, Voglmayr H, Xue H, Piao CG, Li Y (2022) Morphology and phylogeny of Pestalotiopsis (Sporocadaceae, Amphisphaeriales) from Fagaceae leaves in China. Microbiology Spectrum 10:e03272-22. https://doi.org/10.1128/spectrum.03272-22
- Jorna J, Linde JB, Searle PC, Jackson AC, Nielsen ME, Nate MS, Saxton NA, Grewe F, Herrera‐Campos MdlA, Spjut RW, Wu H, Ho B, Lumbsch HT, Leavitt SD (2021) Species boundaries in the messy middle—A genome‐scale validation of species delimitation in a recently diverged lineage of coastal fog desert lichen fungi. Ecology and Evolution 11:18615–18632. https://doi.org/10.1002/ece3.8467
- Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nature Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
- Kamil D, Devi TP, Mathur N, Singh OP, Pandey P, Prabhakaran N, Patil V (2012) Addition of two new species of Pestalotiopsis to the fungal diversity in India. Journal of Mycopathological Research 50:185–191.
- Kanehisa M, Furumichi M, Tanabe M, Sato Y, Morishima K (2017) KEGG: new perspectives on genomes, pathways, diseases and drugs. Nucleic Acids Research 45:D353−D361. https://doi.org/10.1093/nar/gkw1092
- Kapli P, Lutteropp S, Zhang J, Kobert K, Pavlidis P, Stamatakis A, Flouri T (2017) Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo. Bioinformatics 33: 1630–1638. https://doi.org/10.1093/bioinformatics/btx025
- Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Briefings in Bioinformatics 20:1160−1166. https://doi.org/10.1093/bib/bbx108
- Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Molecular Biology and Evolution 30:772−780. https://doi.org/10.1093/molbev/mst010
- Kaur T, Vashisht A, Prakash NT, Reddy MS (2022) Role of selenium-tolerant fungi on plant growth promotion and selenium accumulation of maize plants grown in seleniferous soils. Water, Air, & Soil Pollution 233:17. https://doi.org/10.1007/s11270-021-05490-9
- Keilwagen J, Wenk M, Erickson JL, Schattat MH, Grau J, Hartung F (2016) Using intron position conservation for homology-based gene prediction. Nucleic Acids Research 44:e89. https://doi.org/10.1093/nar/gkw092
- Keith LM, Velasquez ME, Zee FT (2006) Identification and characterization of Pestalotiopsis spp. causing scab disease of guava, Psidium guajava, in Hawaii. Plant Disease 90:16–23. https://doi.org/10.1094/PD-90-0016
- Kenfaoui J, Amiri S, Goura K, Radouane N, Mennani M, Belabess Z, Tahiri A, Fontaine F, Barka EA, Ghadraoui LE, Lahlali R (2024) Uncovering the hidden diversity of fungi associated with grapevine trunk diseases in the Moroccan vineyards. Tropical Plant Pathology 49:662–688. https://doi.org/10.1007/s40858-024-00656-2
- Khalmuratova I, Kim H, Nam YJ, Oh Y, Jeong MJ, Choi HR, You YH, Choo YS, Lee IJ, Shin JH, Yoon H, Kim JG (2015) Diversity and plant growth promoting capacity of endophytic fungi associated with halophytic plants from the west coast of Korea. Mycobiology 43:373–383. https://doi.org/10.5941/MYCO.2015.43.4.373
- Kim D, Bauer BH, Near TJ (2022a) Introgression and species delimitation in the longear sunfish Lepomis megalotis (Teleostei: Percomorpha: Centrarchidae). Systematic Biology 71:273–285. https://doi.org/10.1093/sysbio/syab029
- Kim SH, Lee JW, Kim JS, Lee W, Park MS, Lim YW (2022b) Plastic-inhabiting fungi in marine environments and PCL degradation activity. Antonie van Leeuwenhoek 115:1379–1392. https://doi.org/10.1007/s10482-022-01782-0
- Kohout P, Tedersoo L (2017) Effect of soil moisture on root-associated fungal communities of Erica dominans in Drakensberg mountains in South Africa. Mycorrhiza 27:397–406. https://doi.org/10.1007/s00572-017-0760-5
- Konta S, Tibpromma S, Karunarathna SC, Samarakoon MC, Steven LS, Mapook A, Boonmee S, Senwanna C, Balasuriya A, Eungwanichayapant PD, Hyde KD (2023) Morphology and multigene phylogeny reveal ten novel taxa in Ascomycota from terrestrial palm substrates (Arecaceae) in Thailand. Mycosphere 14:107–152. https://doi.org/10.5943/mycosphere/14/1/2
- Koodalugodaarachchi V, Chethana KWT, Jayawardena RS, Bundhun D, Aluthmuhandiram JVS, Suwannarach N, Manawasinghe I, Lumyong S (2024) New records of pestalotioid species associated with leaf spot disease on Camellia sinensis from northern Thailand. Phytotaxa 647:91–114. https://doi.org/10.11646/phytotaxa.647.1.5
- Korf I (2004) Gene finding in novel genomes. BMC Bioinformatics 5:59. https://doi.org/10.1186/1471-2105-5-59
- Kozlov AM, Darriba D, Flouri T, Morel B, Stamatakis A (2019) RAxML-NG: a fast, scalable and user-friendly tool for maximum likelihood phylogenetic inference. Bioinformatics 35:4453–4455. https://doi.org/10.1093/bioinformatics/btz305
- Krogh A, Larsson B, von Heijne G, Sonnhammer ELL (2001) Predicting transmembrane protein topology with a hidden markov model: application to complete genomes. Journal of Molecular Biology 305:567–580. https://doi.org/10.1006/jmbi.2000.4315
- Kumar V, Cheewangkoon R, Gentekaki E, Maharachchikumbura SSN, Brahmanage RS, Hyde KD (2019) Neopestalotiopsis alpapicalis sp. nov. a new endophyte from tropical mangrove trees in Krabi Province (Thailand). Phytotaxa 393:251–262. https://doi.org/10.11646/phytotaxa.393.3.2
- Kurbetli İ, Aydoğdu M, Sülü G, Woodward S, Demirci F (2020) A potential role for Botryosphaeria parva (Anamorph Neofusicoccum parvum) in plane tree (Platanus orientalis) decline in İstanbul, Turkey. Forest Pathology 50:e12653. https://doi.org/10.1111/efp.12653
- Kwon SL, Cho M, Kim C, Jae-Jin K (2024) First report of seven unrecorded bambusicolous fungi in Korea. Journal of Species Research 13:111–126. https://doi.org/10.12651/JSR.2024.13.2.111
- Lamderos Gálvez EC, Hernández Pérez A, Cerna Chávez E, Ochoa Fuentes YM (2024) Fungi diversity associated with blueberry cultivation in Michoacan, Mexico. Scientia Fungorum 55:e1449
- Lan CZ, Lin X, Dai YL, Gan L, Liu XF, Ruan HC, Yang XJ (2023) First report of leaf blight on Panax notoginseng caused by Pestalotiopsis trachycarpicola in China. Journal of Plant Pathology 105:321. https://doi.org/10.1007/s42161-022-01236-6
- Langer GJ, Peters S, Bußkamp J, Bien S (2024) Cryptostroma corticale and fungal endophytes associated with Fraxinus excelsior affected by ash dieback. Journal of Plant Diseases and Protection 131:1289–1299. https://doi.org/10.1007/s41348-023-00750-8
- Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large datasets. Bioinformatics 30:3276–3278. https://doi.org/10.1093/bioinformatics/btu531
- Lateef AA, Sepiah M, Bolhassan MH (2015) Description of Pseudopestalotiopsis kubahensis sp. nov., a new species of microfungi from Kubah National Park, Sarawak, Malaysia. Current Research in Environmental & Applied Mycology 5:376–381. https://doi.org/10.5943/cream/5/4/8
- Lateef AA, Sepiah M, Bolhassan MH (2018) Molecular identification and diversity of Pestalotiopsis, Neopestalotiopsis and Pseudopestalotiopsis species from four host plants in Sarawak, Borneo Island (Malaysia). Journal of Science and Technology 10:33–43. https://doi.org/10.30880/jst.2018.10.01.006
- Lawrence DP, Brittain GD, Aglave B, Sances FV (2023) First report of Neopestalotiopsis rosae causing crown and root rot of strawberry in California. Plant Disease 107:566. https://doi.org/10.1094/PDIS-04-22-0871-PDN
- Lazarotto M, Bovolini MP, Muniz MFB, Harakawa R, Reiniger LRS, Santos ÁFd (2014) Identification and characterization of pathogenic Pestalotiopsis species to pecan tree in Brazil. Pesquisa Agropecuária Brasileira 49:440–448. https://doi.org/10.1590/S0100-204X2014000600005
- Lazarotto M, Muniz MFB, Poletto T, Dutra CB, Blume E, Harakawa R, Poletto I (2012) First report of Pestalotiopsis clavispora causing leaf spot of Carya illinoensis in Brazil. Plant Disease 96:1826. https://doi.org/10.1094/PDIS-07-12-0615-PDN
- Lee S, Crous PW, Wingfield MJ (2006) Pestalotioid fungi from Restionaceae in the Cape Floral Kingdom. Studies in Mycology 55:175–187. https://doi.org/10.3114/sim.55.1.175
- Lee Y, Kim GH, Kim Y, Park SY, Koh YJ (2019) First report of twig dieback caused by Neopestalotiopsis clavispora on blueberry in Korea. Plant Disease 103:1022. https://doi.org/10.1094/PDIS-10-18-1734-PDN
- Li B, Liu X, Cai J, Feng Y, Huang G (2021a) First report on Neopestalotiopsis aotearoa of rubber tree in China. Plant Disease 105:1223. https://doi.org/10.1094/PDIS-09-20-1930-PDN
- Li H, Manawasinghe IS, Zhang Y, Senanayake IC (2023a) Taxonomic and phylogenic appraisal of Pestalotiopsis linguae sp. nov., and a new record of P. nanjingensis from Pyrrosia lingua (Polypodiaceae) in Southern China. Phytotaxa 587:229–250. https://doi.org/10.11646/phytotaxa.587.3.3
- Li H, Peng BY, Xie JY, Bai YQ, Li DW, Zhu LH (2024a) Pestalotiopsis jiangsuensis sp. nov. causing needle blight on Pinus massoniana in China. Journal of Fungi 10:230. https://doi.org/10.3390/jof10030230
- Li JL, Sun X, Chen L, Guo LD (2016) Community structure of endophytic fungi of four mangrove species in southern China. Mycology 7:180–190. https://doi.org/10.1080/21501203.2016.1258439
- Li K, Zhang C, Wang W, Chen C, Liu Q, Yin H (2023b) First report of Neopestalotiopsis clavispora causing postharvest fruit rot on Actinidia arguta in Liaoning Province, China. Plant Disease 107:217. https://doi.org/10.1094/PDIS-01-22-0240-PDN
- Li L, Yang Q, Li H (2021b) Morphology, phylogeny, and pathogenicity of pestalotioid species on Camellia oleifera in China. Journal of Fungi 7:1080. https://doi.org/10.3390/jof7121080
- Li WL, Dissanayake AJ, Zhang T, Maharachchikumbura SSN, Liu JK (2022) Identification and pathogenicity of pestalotiod fungi associated with woody oil plants in Sichuan province, China. Journal of Fungi 8:1175. https://doi.org/10.3390/jof8111175
- Li X, Liu L, Li H, Lin J, Zuo Y, Peng L, Ding H (2024b) Pestalotiopsis kenyana causes leaf spot disease on Rhododendron agastum in China. Crop Protection 184:106859. https://doi.org/10.1016/j.cropro.2024.106859
- Li Y, Steenwyk JL, Chang Y, Wang Y, James TY, Stajich JE, Spatafora JW, Groenewald M, Dunn CW, Hittinger CT, Shen XX, Rokas A (2021c) A genome-scale phylogeny of the kingdom fungi. Current Biology 31:1653–1665. https://doi.org/10.1016/j.cub.2021.01.074
- Lim SK, Ten LN, Avalos-Ruiz D, Ryu JJ, Kang IK, Lee SY, Jung HY (2022) Isolation and identification of two unreported Sordariomycetes fungi in Korea: Pestalotiopsis clavata and Botryotrichum iranicum. The Korean Journal of Mycology 50:183–194. https://doi.org/10.4489/KJM.20220019
- Lin L, Pan M, Gao H, Tian C, Fan X (2023) The potential fungal pathogens of Euonymus japonicus in Beijing, China. Journal of Fungi 9:271. https://doi.org/10.3390/jof9020271
- Lin Q (2023) First report of Neopestalotiopsis clavispora causing leaf blight on Aegiceras corniculatum in China. Plant Disease 107:3304. https://doi.org/10.1094/PDIS-05-23-0916-PDN
- Lin QL, Li HY, Liu YL (2024) First report of Neopestalotiopsis clavispora causing guava scab in China. Plant Disease 108:3180. https://doi.org/10.1094/PDIS-11-23-2357-PDN
- Lin YZ, Chang TD, Wen CJ, Tsai SH, Lin YH (2022) First report of leaf brown blight caused by Neopestalotiopsis formicarum on jabuticaba in Taiwan. Plant Disease 106:2527. https://doi.org/10.1094/PDIS-07-21-1414-PDN
- Linnakoski R, Puhakka-tarvainen H, Pappinen A (2012) Endophytic fungi isolated from Khaya anthotheca in Ghana. Fungal Ecology 5:298–308. https://doi.org/10.1016/j.funeco.2011.08.006
- Liu AR, Chen SC, Wu SY, Xu T, Guo LD, Jeewon R, Wei JG (2010) Cultural studies coupled with DNA based sequence analyses and its implication on pigmentation as a phylogenetic marker in Pestalotiopsis taxonomy. Molecular Phylogenetics and Evolution 57:528–535. https://doi.org/10.1016/j.ympev.2010.07.017
- Liu AR, Xu T, Guo LD (2007) Molecular and morphological description of Pestalotiopsis hainanensis sp. nov., a new endophyte from a tropical region of China. Fungal Diversity 24:23–36.
- Liu F, Bonthond G, Groenewald JZ, Cai L, Crous PW (2019) Sporocadaceae, a family of coelomycetous fungi with appendage-bearing conidia. Studies in Mycology 92:287–415. https://doi.org/10.1016/j.simyco.2018.11.001
- Liu F, Hou L, Raza M, Cai L (2017) Pestalotiopsis and allied genera from Camellia, with description of 11 new species from China. Scientific Reports 7:866. https://doi.org/10.1038/s41598-017-00972-5
- Liu F, Hu ZD, Yurkov A, Chen XH, Bao WJ, Ma Q, Zhao WN, Pan S, Zhao XM, Liu JH, Wang QM, Boekhout T (2024) Saccharomycetaceae: delineation of fungal genera based on phylogenomic analyses, genomic relatedness indices and genomics-based synapomorphies. Persoonia 52:1–21. https://doi.org/10.3767/persoonia.2024.52.01
- Liu JK, Hyde KD, Jones EBG, Ariyawansa HA, Bhat DJ, Boonmee S, Maharachchikumbura SSN, McKenzie EHC, Phookamsak R, Phukhamsakda C, Shenoy BD, Abdel-Wahab MA, Buyck B, Chen J, Chethana KWT et al. (2015) Fungal diversity notes 1–110: taxonomic and phylogenetic contributions to fungal species. Fungal Diversity 72:1–197. https://doi.org/10.1007/s13225-015-0324-y
- Liu X, Tibpromma S, Zhang F, Xu J, Chethana KWT, Karunarathna SC, Mortimer PE (2021) Neopestalotiopsis cavernicola sp. nov. from gem cave in Yunnan Province, China. Phytotaxa 512:1–27. https://doi.org/10.11646/PHYTOTAXA.512.1.1
- Liu XF, Tibpromma S, Hughes AC, Chethana KWT, Wijayawardene NN, Dai DQ, Du TY, Elgorban AM, Stephenson SL, Suwannarach N, Xu JC, Lu L, Xu RF, Maharachchikumbura SSN, Zhao CL, Bhat DJ, Sun YM, Karunarathna SC, Mortimer PE (2023) Culturable mycota on bats in central and southern Yunnan Province, China. Mycosphere 14:497–662. https://doi.org/10.5943/mycosphere/14/1/7
- Liyanapathiranage P, Avin FA, Swiggart E, Lopez EFP, Parajuli M, Oksel C, Gao Y, Baysal-Gurel F (2023) First report of leaf spot of Panax quinquefolius caused by Pestalotiopsis nanjingensis in Tennessee and the United States. Plant Disease 107:2518. https://doi.org/10.1094/PDIS-01-23-0078-PDN
- Lokesh S, Raghavendra VB, Nagesh KS, Govindappa M (2017) First report of leaf blight of bakul (Mimusops elengi Linn) caused by Pestalotiopsis clavispora (G.F. Atk.) Steyaert in India. Journal of Plant Physiology & Pathology 5:1. https://doi.org/10.4172/2329-955X.1000160
- Lorenzini M, Zapparoli G (2018) Identification of Pestalotiopsis bicilita, Diplodia seriata and Diaporthe eres causing fruit rot in withered grapes in Italy. European Journal of Plant Pathology 151:1089–1093. https://doi.org/10.1007/s10658-017-1416-1
- Lu LM, Chen GQ, Hu XR, Du DC, Pu ZX, Peng AT, Cheng BP (2015) Identification of Pestalotiopsis clavispora causing brown leaf spot on Chinese bayberry in China. Canadian Journal of Plant Pathology 37:397–402. https://doi.org/10.1080/07060661.2015.1065001
- Lücking R, Aime MC, Robbertse B, Miller AN, Ariyawansa HA, Aoki T, Cardinali G, Crous PW, Druzhinina IS, Geiser DM, Hawksworth DL, Hyde KD, Irinyi L, Jeewon R, Johnston PR et al. (2020) Unambiguous identification of fungi: where do we stand and how accurate and precise is fungal DNA barcoding? IMA Fungus 11:14. https://doi.org/10.1186/s43008-020-00033-z
- Ma W, Liu M, Liu L, Tang Z, Dan Y, Cui X, Yin F (2023a) First report of Pseudopestalotiopsis theae causing leaf spot on Euonymus japonicus in China. Plant Disease 107:1238. https://doi.org/10.1094/PDIS-07-22-1722-PDN
- Ma X, Qin Y, Xiang Y, He L, Song F, Wang Z, Jiang Y, Wu L (2023b) First report of Neopestalotiopsis rosae causing leaf blight on shatangju in southern China. Plant Disease 107:2535. https://doi.org/10.1094/PDIS-09-22-2066-PDN
- Ma XY, Maharachchikumbura SSN, Chen BW, Hyde KD, McKenzie EHC, Chomnunti P, Kang JC (2019) Endophytic pestalotiod taxa in Dendrobium orchids. Phytotaxa 419:268–286. https://doi.org/10.11646/phytotaxa.419.3.2
- Machín A, González P, Vicente E, Sánchez M, Estelda C, Ghelfi J, Silvera-Pérez E (2019) First report of root and crown rot caused by Neopestalotiopsis clavispora on strawberry in Uruguay. Plant Disease 103:2946. https://doi.org/10.1094/PDIS-05-19-0948-PDN
- Maharachchikumbura SSN, Chen Y, Ariyawansa HA, Hyde KD, Haelewaters D, Perera RH, Samarakoon MC, Wanasinghe DN, Bustamante DE, Liu JK, Lawrence DP, Cheewangkoon R, Stadler M (2021) Integrative approaches for species delimitation in Ascomycota. Fungal Diversity 109:155–179. https://doi.org/10.1007/s13225-021-00486-6
- Maharachchikumbura SSN, Chukeatirote E, Guo LD, Crous PW, Mckenzie EHC, Hyde KD (2013a) Pestalotiopsis species associated with Camellia sinensis (tea). Mycotaxon 123:47–61. https://doi.org/10.5248/123.47
- Maharachchikumbura SSN, Guo LD, Cai L, Chukeatirote E, Wu WP, Sun X, Crous PW, Bhat DJ, McKenzie EHC, Bahkali AH, Hyde KD (2012) A multi-locus backbone tree for Pestalotiopsis, with a polyphasic characterization of 14 new species. Fungal Diversity 56:95–129. https://doi.org/10.1007/s13225-012-0198-1
- Maharachchikumbura SSN, Guo LD, Chukeatirote E, Hyde KD (2014a) Improving the backbone tree for the genus Pestalotiopsis; addition of P. steyaertii and P. magna sp. nov. Mycological Progress 13:617–624. https://doi.org/10.1007/s11557-013-0944-0
- Maharachchikumbura SSN, Guo LD, Chukeatirote E, McKenzie EHC, Hyde KD (2013b) A destructive new disease of Syzygium samarangense in Thailand caused by the new species Pestalotiopsis samarangensis. Tropical Plant Pathology 38:227–235.
- Maharachchikumbura SSN, Guo LD, Liu ZY, Hyde KD (2016c) Pseudopestalotiopsis ignota and Ps. camelliae spp. nov. associated with grey blight disease of tea in China. Mycological Progress 15:22. https://doi.org/10.1007/s11557-016-1162-3
- Maharachchikumbura SSN, Hyde KD, Groenewald JZ, Xu J, Crous PW (2014b) Pestalotiopsis revisited. Studies in Mycology 79:121–186.
- Maharachchikumbura SSN, Larignon P, Hyde KD, Al-Sadi AM, Liu ZY (2016b) Characterization of Neopestalotiopsis, Pestalotiopsis and Truncatella species associated with grapevine trunk diseases in France. Phytopathologia Mediterranea 55:380–390. https://doi.org/10.14601/Phytopathol_Mediterr-18298
- Maharachchikumbura SSN, Wu SP, Hyde KD, Al-Sadi AM, Liu ZY (2016a) First report of sweet potato leaf spot caused by Neopestalotiopsis ellipsospora in Guizhou Province, China. Journal of Plant Pathology 98:686
- Maharachchikumbura SSN, Zhang Y, Wang Y, Hyde KD (2013c) Pestalotiopsis anacardiacearum sp. nov.(Amphisphaeriaceae) has an intricate relationship with Penicillaria jocosatrix the mango tip borer. Phytotaxa 99: 49–57. https://doi.org/10.11646/phytotaxa.99.2.1
- Majoros WH, Pertea M, Salzberg SL (2004) TigrScan and GlimmerHMM: two open source ab initio eukaryotic gene-finders. Bioinformatics 20:2878−2879. https://doi.org/10.1093/bioinformatics/bth315
- Manawasinghe IS, Jayawardena RS, Li HL, Zhou YY, Zhang W, Phillips AJL, Wanasinghe DN, Dissanayake AJ, Li XH, Li YH, Hyde KD, Yan JY (2021) Microfungi associated with Camellia sinensis: a case study of leaf and shoot necrosis on Tea in Fujian, China. Mycosphere 12:430–518. https://doi.org/10.5943/mycosphere/12/1/6
- Manni M, Berkeley MR, Seppey M, Simão FA, Zdobnov EM (2021a) BUSCO Update: novel and streamlined workflows along with broader and deeper phylogenetic coverage for scoring of eukaryotic, prokaryotic, and viral genomes. Molecular Biology and Evolution 38:4647–4654. https://doi.org/10.1093/molbev/msab199
- Manni M, Berkeley MR, Seppey M, Zdobnov EM (2021b) BUSCO: assessing genomic data quality and beyond. Current Protocols 1:e323. https://doi.org/10.1002/cpz1.323
- McCarthy CGP, Fitzpatrick DA (2019) Pan-genome analyses of model fungal species. Microbial genomics 5:e000243. https://doi.org/10.1099/mgen.0.000243
- McClymont M, Nessia H, Waipara N, Blanchon DJ (2013) First report of Pestalotiopsis clavispora from Selaginella kraussiana (African club moss): an invasive plant species in New Zealand. Australasian Plant Disease Notes 8:79–80. https://doi.org/10.1007/s13314-013-0100-3
- Mertin AA, Laurence MH, van der Merwe M, French K, Liew ECY (2022) The culturable seed mycobiome of two Banksia species is dominated by latent saprotrophic and multi-trophic fungi. Fungal Biology 126:738–745. https://doi.org/10.1016/j.funbio.2022.09.002
- Miles LA, Lopera CA, González S, de García MCC, Franco AE, Restrepo S (2012) Exploring the biocontrol potential of fungal endophytes from an Andean Colombian Paramo ecosystem. BioControl 57:697–710. https://doi.org/10.1007/s10526-012-9442-6
- Minh BQ, Schmidt HA, Chernomor O, Schrempf D, Woodhams MD, von Haeseler A, Lanfear R (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Molecular Biology and Evolution 37:1530−1534. https://doi.org/10.1093/molbev/msaa015
- Mohamed-Azni INA, Sritharan K, Ho SH, Roslan ND, Arulandoo X, Sundram S (2022) Isolation, identification and pathogenicity of fungi associated with leaf blotches in Tenera x Tenera (TxT) variety of oil palm in Malaysia. Journal of Plant Pathology 104:167–177. https://doi.org/10.1007/s42161-021-00953-8
- Monteiro P, Gonçalves MFM, Pinto G, Silva B, Martín-García J, Diez JJ, Alves A (2022) Three novel species of fungi associated with pine species showing needle blight-like disease symptoms. European Journal of Plant Pathology 162:183–202. https://doi.org/10.1007/s10658-021-02395-5
- Morales‐Rodríguez C, Dalla Valle M, Aleandri M, Vannini A (2019) Pestalotiopsis biciliata, a new leaf pathogen of Eucalyptus spp. recorded in Italy. Forest Pathology 49:e12492. https://doi.org/10.1111/efp.12492
- Moreno-Velázquez M, Hernández-Pablo S, QUEZADA-SALINAS A, Alvarado-Rosales D, Rodríguez-Mendoza J, Tovar-Pedraza JM, Márquez-Licona G, de Lourdes Saavedra-Romero L (2022) Neopestalotiopsis australis causing scab disease on Byrsonima crassifolia in Mexico. Research Square. https://doi.org/10.21203/rs.3.rs-1789926/v1
- Moslemi A, Taylor PWJ (2015) Pestalotiopsis chamaeropis causing leaf spot disease of round leaf mint-bush (Prostanthera rotundifolia) in Australia. Australasian Plant Disease Notes 10:29. https://doi.org/10.1007/s13314-015-0179-9
- Mukhtar I, Li H, Quan X, Chou T, Jiang S, Chen B, Wen Z, Xie B (2019) First report of Pestalotiopsis theae causing leaf spot of Ixora chinensis in China. Plant Disease 103:370. https://doi.org/10.1094/PDIS-06-18-1030-PDN
- Nandhini M, Rajini SB, Udayashankar AC, Niranjana SR, Lund OS, Shetty HS, Prakash HS (2018) Diversity, plant growth promoting and downy mildew disease suppression potential of cultivable endophytic fungal communities associated with pearl millet. Biological Control 127:127–138. https://doi.org/10.1016/j.biocontrol.2018.08.019
- Nelson A, Vandegrift R, Carroll GC, Roy BA (2020) Double lives: transfer of fungal endophytes from leaves to woody substrates. PeerJ 8:e9341. https://doi.org/10.7717/peerj.9341
- Norphanphoun C, Jayawardena RS, Chen Y, Wen TC, Meepol W, Hyde KD (2019) Morphological and phylogenetic characterization of novel pestalotioid species associated with mangroves in Thailand. Mycosphere 10:531–578. https://doi.org/10.5943/mycosphere/10/1/9
- Nozawa S, Ando K, Phay N, Watanabe K (2018) Pseudopestalotiopsis dawaina sp. nov. and Ps. kawthaungina sp. nov.: two new species from Myanmar. Mycological Progress 17:865–870. https://doi.org/10.1007/s11557-018-1398-1
- Nozawa S, Seto Y, Watanabe K (2019) First report of leaf blight caused by Pestalotiopsis chamaeropis and Neopestalotiopsis sp. in Japanese andromeda. Journal of General Plant Pathology 85:449–452. https://doi.org/10.1007/s10327-019-00868-4
- Nozawa S, Togawa M, Watanabe K (2022) Reidentification of Pestalotiopsis sensu lato causing gray blight of tea in Japan. Journal of General Plant Pathology 88:293–299. https://doi.org/10.1007/s10327-022-01074-5
- Nozawa S, Uchikawa K, Suga Y, Watanabe K (2020) Infection sources of Pestalotiopsis sensu lato related to loquat fruit rot in Nagasaki Prefecture, Japan. Journal of General Plant Pathology 86:173–179. https://doi.org/10.1007/s10327-020-00908-4
- Nozawa S, Yamaguchi K, Hoang Yen LT, Van Hop D, Nyunt P, Ando K, Watanabe K (2017) Identification of two new species and a sexual morph from the genus Pseudopestalotiopsis. Mycoscience 58:328–337. https://doi.org/10.1016/j.myc.2017.02.008
- Nuthan BR, Meghavarshinigowda BR, Maharachchikumbura SSN, Mahadevakumar S, Marulasiddaswamy KM, Sunilkumar CR, Amruthesh KN, Satish S (2021) Morphological and molecular characterization of Neopestalotiopsis vitis associated with leaf blight disease of Manilkara zapota—a new record from India. Letters in Applied Microbiology 73:352–362. https://doi.org/10.1111/lam.13521
- Obiol JF, Herranz JM, Paris JR, Whiting JR, Rozas J, Riutort M, González-Solís J (2023) Species delimitation using genomic data to resolve taxonomic uncertainties in a speciation continuum of pelagic seabirds. Molecular Phylogenetics and Evolution 179:107671. https://doi.org/10.1016/j.ympev.2022.107671
- Obregón VG, Meneguzzi NG, Ibañez JM, Lattar TE, Kirschbaum DS (2018) First report of Neopestalotiopsis clavispora causing root and crown rot on strawberry plants in Argentina. Plant Disease 102:1856. https://doi.org/10.1094/PDIS-02-18-0330-PDN
- O'Donnell K, Cigelnik E (1997) Two divergent intragenomic rDNA ITS2 types within a monophyletic lineage of the fungus Fusarium are nonorthologous. Molecular Phylogenetics and Evolution 7:103−116.
- Oh SY, Yang JH, Woo JJ, Oh SO, Hur JS (2020) Diversity and distribution patterns of endolichenic fungi in Jeju Island, South Korea. Sustainability 12:3769. https://doi.org/10.3390/su12093769
- Okpalanozie OE, Adebusoye SA, Troiano F, Cattò C, Ilori MO, Cappitelli F (2018) Assessment of indoor air environment of a Nigerian museum library and its biodeteriorated books using culture-dependent and independent techniques. International Biodeterioration & Biodegradation 132:139–149. https://doi.org/10.1016/j.ibiod.2018.03.003
- Oliveira‐Silva L, Batalha‐Filho H, Camelier P, Zanata AM (2023) Past riverine connectivity effects in population structure and distribution of an endemic freshwater fish from northeastern Brazilian rivers: Phylogeographic, taxonomic, and conservation implications. Freshwater Biology 68:1685–1702. https://doi.org/10.1111/fwb.14150
- Ortiz B, Enríquez L, Mejía K, Yanez Y, Sorto Y, Guzman S, Aguilar K, Fontecha G (2022) Molecular characterization of endophytic fungi from pine (Pinus oocarpa) in Honduras. Revis Bionatura 7:13. https://doi.org/10.21931/RB/2022.07.03.13
- Ou S, Jiang N (2019) LTRFINDERparallel: parallelization of LTR_FINDER enabling rapid identification of long terminal repeat retrotransposons. Mobile DNA 10:48. https://doi.org/10.1186/s13100-019-0193-0
- Pak D, You MP, Lanoiselet V, Barbetti MJ (2017) Reservoir of cultivated rice pathogens in wild rice in Australia. European Journal of Plant Pathology 147:295–311. https://doi.org/10.1007/s10658-016-1002-y
- Pandey AK, Hubbali M, Vandana, Dutta P, Babu A (2023) Characterization and identification of fungicide insensitive Pestalotiopsis-like species pathogenic to tea crop in India. World Journal of Microbiology and Biotechnology 39:34. https://doi.org/10.1007/s11274-022-03474-3
- Pandey AK, Sinniah GD, Yadav S, Maharachchikumbura SSN (2024) Pestalotiopsis-like species: host network and lifestyle on tea crop. Fungal Biology Reviews 47:100340. https://doi.org/10.1016/j.fbr.2023.100340
- Park MS, Oh SY, Lee S, Eimes JA, Lim YW (2018) Fungal diversity and enzyme activity associated with sailfin sandfish egg masses in Korea. Fungal Ecology 34:1–9. https://doi.org/10.1016/j.funeco.2018.03.004
- Park MS, Yoo S, Cho Y, Park KH, Kim NK, Lee HS, Lim YW (2021) Investigation of the fungal diversity of the federated states of micronesia and the construction of an updated fungal inventory. Mycobiology 49:551–558. https://doi.org/10.1080/12298093.2021.2012327
- Park S, Lee SY, Lee JJ, Back CG, Ten L, Lee HB, Jung HY (2016) First report of Neopestalotiopsis australis isolated from soil in Korea. The Korean Journal of Mycology 44:360–364. https://doi.org/10.4489/KJM.2016.44.4.360
- Paulino GVB, Félix CR, Landell MF (2019) Diversity of filamentous fungi associated with coral and sponges in coastal reefs of northeast Brazil. Journal of Basic Microbiology 60:103–111. https://doi.org/10.1002/jobm.201900394
- Peng C, Crous PW, Jiang N, Fan XL, Liang YM, Tian CM (2022) Diversity of Sporocadaceae (pestalotioid fungi) from Rosa in China. Persoonia 49:201–260. https://doi.org/10.3767/persoonia.2022.49.07
- Pereira DS, Phillips AJL (2024) Diaporthe species on palms–integrative taxonomic approach for species boundaries delimitation in the genus Diaporthe, with the description of D. pygmaeae sp. nov. Studies in Mycology 109:487–594. https://doi.org/10.3114/sim.2024.109.08
- Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nature Methods 8:785–786. https://doi.org/10.1038/nmeth.1701
- Phukhamsakda C, McKenzie EHC, Phillips AJL, Gareth Jones EB, Jayarama Bhat D, Stadler M, Bhunjun CS, Wanasinghe DN, Thongbai B, Camporesi E, Ertz D, Jayawardena RS, Perera RH, Ekanayake AH, Tibpromma S, Doilom M, Xu J, Hyde KD (2020) Microfungi associated with Clematis (Ranunculaceae) with an integrated approach to delimiting species boundaries. Fungal Diversity 102:1–203. https://doi.org/10.1007/s13225-020-00448-4
- Pornsuriya C, Chairin T, Thaochan N, Sunpapao A (2020) Identification and characterization of Neopestalotiopsis fungi associated with a novel leaf fall disease of rubber trees (Hevea brasiliensis) in Thailand. Journal of Phytopathology 168:416–427. https://doi.org/10.1111/jph.12906
- Prasannath K, Galea VJ, Akinsanmi OA (2020) Characterisation of leaf spots caused by Neopestalotiopsis clavispora and Colletotrichum siamense in macadamia in Australia. European Journal of Plant Pathology 156:1219–1225. https://doi.org/10.1007/s10658-020-01962-6
- Prasannath K, Galea VJ, Akinsanmi OA (2023) Diversity and pathogenicity of species of Botrytis, Cladosporium, Neopestalotiopsis and Pestalotiopsis causing flower diseases of macadamia in Australia. Plant Pathology 72:881–899. https://doi.org/10.1111/ppa.13707
- Prasannath K, Shivas RG, Galea VJ, Akinsanmi OA (2021) Neopestalotiopsis species associated with flower diseases of Macadamia integrifolia in Australia. Journal of Fungi 7:771. https://doi.org/10.3390/jof7090771
- Prematunga CJ, You LQ, Gomdola D, Balasuriya A, Yang YH, Jayawardena RS, Luo M (2022) An addition to pestalotioid fungi in China: Neopestalotiopsis fragariae sp. nov. causing leaf spots on Fragaria × ananassa. Asian Journal of Mycology 5:220–238. https://doi.org/10.5943/ajom/5/2/10
- Price AL, Jones NC, Pevzner PA (2005) De novo identification of repeat families in large genomes. Bioinformatics 21:i351–i358. https://doi.org/10.1093/bioinformatics/bti1018
- Puillandre N, Brouillet S, Achaz G (2021) ASAP: assemble species by automatic partitioning. Molecular Ecology Resources 21:609–620. https://doi.org/10.1111/1755-0998.13281
- Puillandre N, Lambert A, Brouillet S, Achaz G (2012) ABGD, Automatic Barcode Gap Discovery for primary species delimitation. Molecular Ecology 21:1864–1877. https://doi.org/10.1111/j.1365-294X.2011.05239.x
- Qi M, Xie CX, Chen QW, Yu ZD (2021) Pestalotiopsis trachicarpicola, a novel pathogen causes twig blight of Pinus bungeana (Pinaceae: Pinoideae) in China. Antonie van Leeuwenhoek 114:1–9. https://doi.org/10.1007/s10482-020-01500-8
- Qi XL, He J, Ju YW, Huang L (2023a) First report of leaf spot on Elaeagnus pungens caused by Neopestalotiopsis clavispora in China. Plant Disease 107:2251. https://doi.org/10.1094/PDIS-10-22-2457-PDN
- Qi YX, Zhang H, Peng J, Zeng FY, Xie YX, Yu QF, Zhang X (2023b) First report of Neopestalotiopsis clavispora causing leaf spot disease on banana (Musa acuminata) in China. Plant Disease 107:220. https://doi.org/10.1094/PDIS-03-22-0455-PDN
- Qian Y, Kang J, Geng K, Wang L, Lei B (2014) Endophytic fungi from Artemisia argyi Levl. et Vant. and their bioactivity. Chiang Mai Journal of Science 41:910–921.
- Qin Q, Lu Z, Lu Z, Ding L, Chi Z, Shan B (2020) First report of leaf spot on Paphiopedilum micranthum caused by Neopestalotiopsis saprophytica in China. Plant Disease 104:2738. https://doi.org/10.1094/PDIS-02-20-0275-PDN
- Qin R, Li Q, Huang S, Chen X, Mo J, Guo T, Huang H, Tang L, Yu Z (2023) Fruit rot on persimmon caused by Neopestalotiopsis saprophytica and Neopestalotiopsis ellipsospora in Guangxi, China. Plant Disease 107:2531. https://doi.org/10.1094/PDIS-05-22-1168-PDN
- Qin R, Tang L, Li Q, Huang S, Chen X, Mo J, Guo T, Yu Z (2022) Leaf spot on persimmon caused by Neopestalotiopsis ellipsospora in China. New Disease Reports 46:e12127. https://doi.org/10.1002/ndr2.12127
- Qiu L, Liu J, Kuang W, Zhang K, Ma J (2022) First report of Pestalotiopsis chamaeropis causing leaf spot on Eurya nitida in China. Plant Disease 106:329. https://doi.org/10.1094/PDIS-06-21-1170-PDN
- R Core Team (2024). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. <https://www.R-project.org/>.
- Rajashekara H, Pandian RTP, Mahadevakumar S, Raviprasad TN, Vanitha K, Siddanna S, Thube SH, Khandelwal V, Chandranayaka S (2023) First report of Neopestalotiopsis clavispora causing cashew leaf blight disease in India. Plant Disease 107:2864. https://doi.org/10.1094/PDIS-03-23-0545-PDN
- Ran SF, Maharachchikumbura SSN, Ren YL, Liu H, Chen KR, Wang YX, Wang Y (2017) Two new records in Pestalotiopsidaceae associated with Orchidaceae disease in Guangxi Province, China. Mycosphere 8:121–130. https://doi.org/10.5943/mycosphere/8/1/11
- Razaghi P, Raza M, Han SL, Ma ZY, Cai L, Zhao P, Chen Q, Phurbu D, Liu F (2024) Sporocadaceae revisited. Studies in Mycology 109:155–272. https://doi.org/10.3114/sim.2024.109.03
- Rebollar-Alviter A, Silva-Rojas HV, Fuentes-Aragón D, Acosta-González U, Martínez-Ruiz M, Parra-Robles BE (2020) An emerging strawberry fungal disease associated with root rot, crown rot and leaf spot caused by Neopestalotiopsis rosae in Mexico. Plant Disease 104:2054–2059. https://doi.org/10.1094/PDIS-11-19-2493-SC
- Reddy MS, Murali TS, Suryanarayanan TS, Govinda Rajulu MB, Thirunavukkarasu N (2016) Pestalotiopsis species occur as generalist endophytes in trees of Western Ghats forests of southern India. Fungal Ecology 24:70–75. https://doi.org/10.1016/j.funeco.2016.09.002
- Rodríguez-Gálvez E, Hilário S, Lopes A, Alves A (2020) Diversity and pathogenicity of Lasiodiplodia and Neopestalotiopsis species associated with stem blight and dieback of blueberry plants in Peru. European Journal of Plant Pathology 157:89–102. https://doi.org/10.1007/s10658-020-01983-1
- Rosado AWC, Machado AR, Pereira OL (2015) Postharvest stem-end rot on immature coconut caused by Pestalotiopsis adusta in Brazil. Plant Disease 99:1036. https://doi.org/10.1094/PDIS-01-15-0058-PDN
- Sahibu A, Abdullah MM, Farizah AN, Nordahliawate MSS (2024) First report of pestalotioid species causing foliar diseases on mangrove trees in Terengganu, Peninsular Malaysia. New Disease Reports 49:e12287. https://doi.org/10.1002/ndr2.12287
- Samaradiwakara NP, de Farias ARG, Tennakoon DS, Aluthmuhandiram JVS, Bhunjun CS, Chethana KWT, Kumla J, Lumyong S (2023) Appendage-bearing Sordariomycetes from Dipterocarpus alatus leaf litter in Thailand. Journal of Fungi 9:625. https://doi.org/10.3390/jof9060625
- Sandoval-Sánchez M, Nieto-Ángel D, Sandoval-Islas JS, Téliz-Ortiz D, Orozco-Santos M, Silva-Rojas HV (2013) Fungi associated to stem-end rot and dieback of mango (Mangifera indica L.). Agrociencia 47:61–73.
- Sane S, Sharma S, Konduri R, Fernandes M (2019) Emerging corneal pathogens first report of Pseudopestalotiopsis theae keratitis. Indian Journal of Ophthalmology 67:150–152. https://doi.org/10.4103/ijo.IJO79118
- Santos CC, Domingues JL, Santos RF, Spósito MB, Santos A, Novaes QS (2019) First report of Neopestalotiopsis clavispora causing leaf spot on macadamia in Brazil. Plant Disease 103:1790. https://doi.org/10.1094/PDIS-01-19-0108-PDN
- Santos GS, Mafia RG, Aguiar AM, Zarpelon TG, Damacena MB, Barros AF, Ferreira MA (2020) Stem rot of eucalyptus cuttings caused by Neopestalotiopsis spp. in Brazil. Journal of Phytopathology 168:311–321. https://doi.org/10.1111/jph.12894
- Santos J, Hilário S, Pinto G, Alves A (2022) Diversity and pathogenicity of pestalotioid fungi associated with blueberry plants in Portugal, with description of three novel species of Neopestalotiopsis. European Journal of Plant Pathology 162:539–555. https://doi.org/10.1007/s10658-021-02419-0
- Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding Consortium (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proceedings of the National Academy of Sciences of the United States of America 109: 6241–6246. https://doi.org/10.1073/pnas.1117018109
- Seifollahi E, de Farias ARG, Jayawardena RS, Hyde KD (2023) Taxonomic advances from fungal flora associated with ferns and fern-like hosts in Northern Thailand. Plants 12:683. https://doi.org/10.3390/plants12030683
- Senanayake IC, Lian TT, Mai XM, Jeewon R, Maharachchikumbura SSN, Hyde KD, Zeng YJ, Tian SL, Xie N (2020) New geographical records of Neopestalotiopsis and Pestalotiopsis species in Guangdong Province, China. Asian Journal of Mycology 3:510–530. https://doi.org/10.5943/ajom/3/1/19
- Shara M, Basyuni M, Hasanuddin (2023) Potential of phylloplane fungi from mangrove plant (Rhizophora apiculata Blume) as biological control agents against Fusarium oxysporum f. sp. cubense in banana plant (Musa acuminata L.). Forests 14:167. https://doi.org/10.3390/f14020167
- Shetty KG, Rivadeneira DV, Jayachandran K, Walker DM (2016) Isolation and molecular characterization of the fungal endophytic microbiome from conventionally and organically grown avocado trees in South Florida. Mycological Progress 15:977–986. https://doi.org/10.1007/s11557-016-1219-3
- Shi J, Li B, Wang S, Zhang W, Shang M, Wang Y, Liu B (2024) Occurrence of Neopestalotiopsis clavispora causing apple leaf spot in China. Agronomy 14:1658. https://doi.org/10.3390/agronomy14081658
- Shi J, Zhang X, Liu Y, Zhang Z, Wang Z, Xue C, Ma Y, Wang F (2022) First report of Neopestalotiopsis clavispora causing calyx and receptacle blight on strawberry in China. Plant Disease 106:1307. https://doi.org/10.1094/PDIS-07-21-1376-PDN
- Shu J, Yu Z, Sun W, Zhao J, Li Q, Tang L, Guo T, Huang S, Mo J, Hsiang T, Shuming L (2020) Identification and characterization of pestalotioid fungi causing leaf spots on mango in southern China. Plant Disease 104:1207–1213. https://doi.org/10.1094/PDIS-03-19-0438-RE
- Silva AC, Diogo E, Henriques J, Ramos AP, Sandoval-Denis M, Crous PW, Bragança H (2020) Pestalotiopsis pini sp. nov., an emerging pathogen on stone pine (Pinus pinea L.). Forests 11:805. https://doi.org/10.3390/f11080805
- Silvério ML, de Queiroz Calvacanti MA, da Silva GA, de Oliveira RJV, Bezerra JL (2016) A new epifoliar species of Neopestalotiopsis from Brazil. Agrotrópica 28:151–158. https://doi.org/10.21757/0103-3816.2016v28n2p151-158
- Simão FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM (2015) BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31:3210–3212. https://doi.org/10.1093/bioinformatics/btv351
- Singha R, Sharma D, Saha AK, Das P (2024) Foliar phenols and flavonoids level in pteridophytes: an insight to culturable fungal endophyte colonisation. Archives of Microbiology 206:170. https://doi.org/10.1007/s00203-024-03880-1
- Sklenář F, Glässnerová K, Jurjević Ž, Houbraken J, Samson RA, Visagie CM, Yilmaz N, Gené J, Cano J, Chen AJ, Nováková A, Yaguchi T, Kolařík M, Hubka V (2022) Taxonomy of Aspergillus series Versicolores: species reduction and lessons learned about intraspecific variability. Studies in Mycology 102:53–93. https://doi.org/10.3114/sim.2022.102.02
- Solpot TC, Villanueva JD, Abello MP, Zuñiga JBM (2024) First report of Neopestalotiopsis clavispora causing foliar disease of rubber (Hevea brasiliensis) in the Philippines. Journal of Phytopathology 172:e13238. https://doi.org/10.1111/jph.13238
- Song X, Li Y, Cao Z, Wang D (2022) Diversity assessment of endophytic fungi isolated from Euonymus japonicus. Journal of Plant Diseases and Protection 129:261–269. https://doi.org/10.1007/s41348-022-00588-6
- Song Y, Geng K, Zhang B, Hyde KD, Zhao WS, Wei JG, Kang JC, Wang Y (2013) Two new species of Pestalotiopsis from Southern China. Phytotaxa 126:22–30. https://doi.org/10.11646/phytotaxa.126.1.2
- Song Y, Maharachchikumbura SSN, Jiang YL, Hyde KD, Wang Y (2014a) Pestalotiopsis keteleeria sp. nov., isolated from Keteleeria pubescens in China. Chiang Mai Journal of Science 41:885–893
- Song Y, Tangthirasunun N, Maharachchikumbura SSN, Jiang Y, Xu J, Hyde KD, Wang Y (2014b) Novel Pestalotiopsis species from Thailand point to the rich undiscovered diversity of this chemically creative genus. Cryptogamie, Mycologie 35:139–149. https://doi.org/10.7872/crym.v35.iss2.2014.139
- Sperschneider J, Gardiner DM, Dodds PN, Tini F, Covarelli L, Singh KB, Manners JM, Taylor JM (2016) EffectorP: predicting fungal effector proteins from secretomes using machine learning. New Phytologist 210:743–761. https://doi.org/10.1111/nph.13794
- Spjut R, Simon A, Guissard M, Magain N, Sérusiaux E (2020) The fruticose genera in the Ramalinaceae (Ascomycota, Lecanoromycetes): their diversity and evolutionary history. MycoKeys 73:1–68. https://doi.org/10.3897/mycokeys.73.47287
- Stanke M, Keller O, Gunduz I, Hayes A, Waack S, Morgenstern B (2006) AUGUSTUS: ab initio prediction of alternative transcripts. Nucleic Acids Research 34:W435−W439. https://doi.org/10.1093/nar/gkl200
- Steenwyk JL, Balamurugan C, Raja HA, Gonçalves C, Li N, Martin F, Berman J, Oberlies NH, Gibbons JG, Goldman GH, Geiser DM, Houbraken J, Hibbett DS, Rokas A (2024) Phylogenomics reveals extensive misidentification of fungal strains from the genus Aspergillus. Microbiology Spectrum 12:e03980-23. https://doi.org/10.1128/spectrum.03980-23
- Steinrucken TV, Raghavendra AKH, Powell JR, Bissett A, van Klinken RD (2017) Triggering dieback in an invasive plant: endophyte diversity and pathogenicity. Australasian Plant Pathology 46:157–170. https://doi.org/10.1007/s13313-017-0472-5
- Steinrucken TV, Vitelli JS, Holdom DG, Tan YP (2022) The diversity of microfungi associated with grasses in the Sporobolus indicus complex in Queensland, Australia. Frontiers in Fungal Biology 3:956837. https://doi.org/10.3389/ffunb.2022.956837
- Stengel A, Stanke KM, Quattrone AC, Herr JR (2022) Improving taxonomic delimitation of fungal species in the age of genomics and phenomics. Frontiers in Microbiology 13:847067. https://doi.org/10.3389/fmicb.2022.847067
- Steyaert RL (1949) Contribution à l'étude monographique de Pestalotia de Not. et Monochaetia Sacc. (Truncatella gen. nov. et Pestalotiopsis gen. nov.). Bulletin du Jardin botanique de l'Etat, Bruxelles 19:285–347.
- Su H, Kang JC, Cao JJ, Mo L, Hyde KD (2014) Medicinal plant endophytes produce analogous bioactive compounds. Chiang Mai Journal of Science 41:1–13.
- Sumaya NPDN, Caluban PIM, Borja BT (2023) First report of Pseudopestalotiopsis theae causing leaf spot of robusta coffee in the Philippines. New Disease Reports 48:e12218. https://doi.org/10.1002/ndr2.12218
- Sun Q, Harishchandra D, Jia J, Zuo Q, Zhang G, Wang Q, Yan J, Zhang W, Li X (2021) Role of Neopestalotiopsis rosae in causing root rot of strawberry in Beijing, China. Crop Protection 147:105710. https://doi.org/10.1016/j.cropro.2021.105710
- Sun YR, Jayawardena RS, Sun JE, Wang Y (2023) Pestalotioid species associated with medicinal plants in southwest China and Thailand. Microbiology Spectrum 11:e03987-22. https://doi.org/10.1128/spectrum.03987-22
- Suwannarach N, Sujarit K, Kumla J, Bussaban B, Lumyong S (2013) First report of leaf spot disease on oil palm caused by Pestalotiopsis theae in Thailand. Journal of General Plant Pathology 79:277–279. https://doi.org/10.1007/s10327-013-0453-7
- Tan XM, Zhou YQ, Zhou XL, Xia XH, Wei Y, He LL, Tang HZ, Yu LY (2018) Diversity and bioactive potential of culturable fungal endophytes of Dysosma versipellis; a rare medicinal plant endemic to China. Scientific Reports 8:5929. https://doi.org/10.1038/s41598-018-24313-2
- Tang J, Lai LX, Du YL, Yang Q (2021) First report of Neopestalotiopsis protearum causing seed rot on Camellia oleifera in China. Plant Disease 105:4152. https://doi.org/10.1094/PDIS-12-20-2717-PDN
- Tao Y, Quan X, Khokhar I, Anjum T, Song H, Mukhtar I (2021) First report of Pseudopestalotiopsis theae causing leaf spot of date palm (Phoenix dactylifera) in China. Plant Disease 105:508. https://doi.org/10.1094/PDIS-06-20-1356-PDN
- Taylor JW, Jacobson DJ, Kroken S, Kasuga T, Geiser DM, Hibbett DS, Fisher MC (2000) Phylogenetic species recognition and species concepts in fungi. Fungal Genetics and Biology 31:21−32. https://doi.org/10.1006/fgbi.2000.1228
- Tejesvi MV, Kini KR, Prakash HS, Subbiah V, Shetty HS (2008) Antioxidant, antihypertensive, and antibacterial properties of endophytic Pestalotiopsis species from medicinal plants. Canadian Journal of Microbiology 54:769–780. https://doi.org/10.1139/W08-070
- Tennakoon DS, Kuo CH, Maharachchikumbura SSN, Thambugala KM, Gentekaki E, Phillips AJL, Bhat DJ, Wanasinghe DN, de Silva NI, Promputtha I, Hyde KD (2021) Taxonomic and phylogenetic contributions to Celtis formosana, Ficus ampelas, F. septica, Macaranga tanarius and Morus australis leaf litter inhabiting microfungi. Fungal Diversity 108:1–215. https://doi.org/10.1007/s13225-021-00474-w
- Tian S, Xu R, Bhunjun CS, Su W, Hyde KD, Li Y, Fu Y, Phukhamsakda C (2022) Combination of morphological and molecular data support Pestalotiopsis eleutherococci (Sporocadaceae) as a new species. Phytotaxa 566:105–120. https://doi.org/10.11646/PHYTOTAXA.566.1.6
- Tian XG, Bao DF, Karunarathna SC, Jayawardena RS, Hyde KD, Bhat DJ, Luo ZL, Elgorban AM, Hongsanan S, Rajeshkumar KC, Maharachchikumbura SSN, Suwannarach N, Dawoud TM, Lu YZ, Han JJ et al. (2024) Taxonomy and phylogeny of ascomycetes associated with selected economically important monocotyledons in China and Thailand. Mycosphere 15:1–274. https://doi.org/10.5943/mycosphere/15/1/1
- Tibpromma S, Hyde KD, Bhat JD, Mortimer PE, Xu J, Promputtha I, Doilom M, Yang JB, Tang AMC, Karunarathna SC (2018a) Identification of endophytic fungi from leaves of Pandanaceae based on their morphotypes and DNA sequence data from southern Thailand. MycoKeys 33:25–67. https://doi.org/10.3897/mycokeys.33.23670
- Tibpromma S, Hyde KD, McKenzie EHC, Bhat DJ, Phillips AJL, Wanasinghe DN, Samarakoon MC, Jayawardena RS, Dissanayake AJ, Tennakoon DS, Doilom M, Phookamsak R, Tang AMC, Xu J, Mortimer PE, Promputtha I, Maharachchikumbura SSN, Khan S, Karunarathna SC (2018b) Fungal diversity notes 840–928: micro-fungi associated with Pandanaceae. Fungal Diversity 93:1–160. https://doi.org/10.1007/s13225-018-0408-6
- Tibpromma S, Mortimer PE, Karunarathna SC, Zhan F, Xu J, Promputtha I, Yan K (2019) Morphology and multi-gene phylogeny reveal Pestalotiopsis pinicola sp. nov. and a new host record of Cladosporium anthropophilum from edible pine (Pinus armandii) seeds in Yunnan Province, China. Pathogens 8:285. https://doi.org/10.3390/pathogens8040285
- Tsai I, Chung CL, Lin SR, Hung TH, Shen TL, Hu CY, Hozzein WN, Ariyawansa HA (2021) Cryptic diversity, molecular systematics, and pathogenicity of genus Pestalotiopsis and allied genera causing gray blight disease of tea in Taiwan, with a description of a new Pseudopestalotiopsis species. Plant Disease 105:425–443. https://doi.org/10.1094/pdis-05-20-1134-re
- Tsai I, Maharachchikumbura SSN, Hyde KD, Ariyawansa HA (2018) Molecular phylogeny, morphology and pathogenicity of Pseudopestalotiopsis species on Ixora in Taiwan. Mycological Progress 17:941–952. https://doi.org/10.1007/s11557-018-1404-7
- Tsvetkova YV, Kuznetsova AA (2022) Detection of anthracnose in strawberry and methods of etiological diagnosis. Doklady Biological Sciences 507:473–484. https://doi.org/10.1134/S0012496622060229
- Turland NJ, Wiersema JH, Barrie FR, Greuter W, Hawksworth DL, Herendeen PS, Knapp S, Kusber WH, Li DZ, Marhold K, May TW, McNeill J, Monro AM, Prado J, Price MJ, Smith GF (2018) International Code of Nomenclature for algae, fungi, and plants (Shenzhen Code) adopted by the Nineteenth International Botanical Congress Shenzhen, China. Koeltz Botanical Books. https://doi.org/10.12705/Code.2018
- Undugoda L, Thambugala K, Kannangara S, Munasinghe J, Premarathna N, Dharmasiri N (2023) Phylloremediation of pyrene and anthracene by endophytic fungi inhabiting tea leaves (Camellia sinensis (L.) Kuntze) in Sri Lanka. New Zealand Journal of Botany 63: 1922–1935. https://doi.org/10.1080/0028825X.2023.2258829
- Valencia AL, Torres R, Latorre BA (2011) First report of Pestalotiopsis clavispora and Pestalotiopsis spp. causing postharvest stem end rot of avocado in Chile. Plant Disease 95:492. https://doi.org/10.1094/PDIS-11-10-0844
- Varghese NJ, Mukherjee S, Ivanova N, Konstantinidis KT, Mavrommatis K, Kyrpides NC, Pati A (2015) Microbial species delineation using whole genome sequences. Nucleic Acids Research 43:6761–6771. https://doi.org/10.1093/nar/gkv657
- Vega Gutierrez SM, Illescas Guevara JF, Andersen CC, von Stein JK, Robinson SC (2020) Exploratory sampling of spalting fungi in the Southern Peruvian Amazon forest. Challenges 11:32. https://doi.org/10.3390/challe11020032
- Vences M, Miralles A, Brouillet S, Ducasse J, Fedosov A, Kharchev V, Kostadinov I, Kumari S, Patmanidis S, Scherz MD , Puillandre N, Renner SS (2021) iTaxoTools 0.1: Kickstarting a specimen-based software toolkit for taxonomists. BioRxiv. https://doi.org/10.1101/2021.03.26.435825
- Vu D, Groenewald M, de Vries M, Gehrmann T, Stielow B, Eberhardt U, Al-Hatmi A, Groenewald JZ, Cardinali G, Houbraken J, Boekhout T, Crous PW, Robert V, Verkley GJM (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Studies in Mycology 92:135–154. https://doi.org/10.1016/j.simyco.2018.05.001
- Vu THN, Pham NS, Quach NT, Le PC, Pham QA, Ngo CC, Nguyen VT, Anh DH, Quang TH, Chu HH, Phi QT (2024) Fusarium foetens AQF6 isolated from Amentotaxus ynnanensis H.L.Li as a prolific source of antioxidant compounds. Applied Sciences 14:2048. https://doi.org/10.3390/app14052048
- Wang QC, Zhan ZJ, Sattar A, Wang HN, Zhou LF, Eckhardt L, Li GQ, Liu FF, Xu HC, Zhou XD (2025) Pestalotiopsis (Amphisphaeriales, Sporocadaceae) species including six new taxa inhabiting pines from different climate zones in China. IMA Fungus 16:e151614. https://doi.org/10.3897/imafungus.16.151614
- Wang QT, Cheng YH, Zhuo L, Wang Y, Zhou H, Hou CL (2022) Neopestalotiopsis longiappendiculata as the agent of grey blight disease of Camellia spp. Journal of Phytopathology 170:770–777. https://doi.org/10.1111/jph.13139
- Wang R, Chen S, Zheng B, Liu P, Li B, Weng Q, Chen Q (2019a) Occurrence of leaf spot disease caused by Neopestalotiopsis clavispora on Taxus chinensis in China. Forest Pathology 49:e12540. https://doi.org/10.1111/efp.12540
- Wang S, Mi X, Wu Z, Zhang L, Wei C (2019b) Characterization and pathogenicity of Pestalotiopsis-like species associated with gray blight disease on Camellia sinensis in Anhui Province, China. Plant Disease 103:2786–2797. https://doi.org/10.1094/PDIS-02-19-0412-RE
- Wang Y, Tsui KM, Chen S, You C (2024) Diversity, pathogenicity and two new species of pestalotioid fungi (Amphisphaeriales) associated with Chinese Yew in Guangxi, China. MycoKeys 102:201–224. https://doi.org/10.3897/mycokeys.102.113696
- Wang Y, Xiong F, Lu Q, Hao X, Zheng M, Wang L, Li N, Ding C, Wang X, Yang Y (2019c) Diversity of Pestalotiopsis-like species causing gray blight disease of tea plants (Camellia sinensis) in China, including two novel Pestalotiopsis species, and analysis of their pathogenicity. Plant Disease 103:2548–2558. https://doi.org/10.1094/PDIS-02-19-0264-RE
- Wang ZH, Zhao ZX, Hong N, Ni D, Cai L, Xu WX, Xiao YN (2017) Characterization of causal agents of a novel disease inducing brown-black spots on tender tea leaves in China. Plant Disease 101:1802–1811. https://doi.org/10.1094/PDIS-04-17-0495-RE
- Watanabe K, Motohashi K, Ono Y (2010) Description of Pestalotiopsis pallidotheae: a new species from Japan. Mycoscience 51:182–188. https://doi.org/10.1007/s10267-009-0025-z
- Watanabe K, Nakazono T, Ono Y (2012) Morphology evolution and molecular phylogeny of Pestalotiopsis (Coelomycetes) based on ITS2 secondary structure. Mycoscience 53:227–237. https://doi.org/10.1007/s10267-011-0157-9
- Watanabe K, Nozawa S, Hsiang T, Callan B (2018) The cup fungus Pestalopezia brunneopruinosa is Pestalotiopsis gibbosa and belongs to Sordariomycetes. PLoS ONE 13:e0197025. https://doi.org/10.1371/journal.pone.0197025
- Wei JG, Phan CK, Wang L, Xu T, Luo JT, Sun X, Guo LD (2013) Pestalotiopsis yunnanensis sp. nov., an endophyte from Podocarpus macrophyllus (Podocarpaceae) based on morphology and ITS sequence data. Mycological Progress 12:563–568. https://doi.org/10.1007/s11557-012-0863-5
- Wei JG, Xu T, Guo LD, Liu AR, Zhang Y, Pan XH (2007) Endophytic Pestalotiopsis species associated with plants of Podocarpaceae, Theaceae and Taxaceae in southern China. Fungal Diversity 24:55–74.
- Wei T, Luo M, Zhang H, Jia W, Zeng Y, Jiang Y (2023) First report of leaf spot disease associated with faba bean (Vicia faba) caused by Pestalotiopsis rosea in China. Plant Disease 107:2885. https://doi.org/10.1094/PDIS-12-22-2920-PDN
- White TJ, Bruns T, Lee S, Taylor J (1990) Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. PCR Protocols: A Guide to Methods and Applications 18:315−322.
- Wibberg D, Stadler M, Lambert C, Bunk B, Spröer C, Rückert C, Kalinowski J, Cox RJ, Kuhnert E (2021) High quality genome sequences of thirteen Hypoxylaceae (Ascomycota) strengthen the phylogenetic family backbone and enable the discovery of new taxa. Fungal Diversity 106:7–28. https://doi.org/10.1007/s13225-020-00447-5
- Wijayawardene NN, Hyde KD, Dai DQ, Sánchez-García M, Goto BT, Saxena RK, Erdoğdu M, Selçuk F, Rajeshkumar KC, Aptroot A, Błaszkowski J, Boonyuen N, da Silva GA, de Souza FA, Dong W et al. (2022) Outline of Fungi and fungus-like taxa – 2021. Mycosphere 13:53–453. https://doi.org/10.5943/mycosphere/13/1/2
- Win PM, Matsumura E, Fukuda K (2018) Diversity of tea endophytic fungi: cultivar-and tissue preferences. Applied Ecology & Environmental Research 16:677–695. https://doi.org/10.15666/aeer/1601_677695
- Win PM, Matsumura E, Fukuda K (2021) Effects of pesticides on the diversity of endophytic fungi in tea plants. Microbial Ecology 82:62–72. https://doi.org/10.1007/s00248-020-01675-7
- Wu C, Wang Y, Yang Y (2022) Pestalotiopsis diversity: species, dispositions, secondary metabolites, and bioactivities. Molecules 27:8088. https://doi.org/10.3390/molecules27228088
- Xavier KV, Yu X, Vallad GE (2021) First report of Neopestalotiopsis rosae causing foliar and fruit spots on pomegranate in Florida. Plant Disease 105:504. https://doi.org/10.1094/PDIS-06-20-1282-PDN
- Xia Z, Yin Q, Qiu C, Yang Y, Guo D, Huang H, Tang Q, Jiang S, Zhao X, Chen Z (2022) Transcriptome profiling of the leaf spot pathogen, Pestalotiopsis trachicarpicola, and its host, tea (Camellia sinensis), during infection. Plant Disease 106:2247–2252. https://doi.org/10.1094/PDIS-12-21-2698-A
- Xiao J, Lin L, Hu J, Jiao F, Duan D, Zhang Q, Tang H, Gao J, Wang L, Wang X (2017) Highly oxygenated caryophyllene-type and drimane-type sesquiterpenes from Pestalotiopsis adusta, an endophytic fungus of Sinopodophyllum hexandrum. RSC advances 7:29071-29079. https://doi.org/10.1039/C7RA04267A
- Xie J, Wei JG, Huang RS, Wei JF, Luo JT, Yang XH, Yang XB (2018) First report of ring spot on Kadsura coccinea caused by Neopestalotiopsis clavispora in China. Plant Disease 102:2032. https://doi.org/10.1094/PDIS-01-18-0078-PDN
- Xing X, Guo S (2011) Fungal endophyte communities in four Rhizophoraceae mangrove species on the south coast of China. Ecological Research 26:403–409. https://doi.org/10.1007/s11284-010-0795-y
- Xing YM, Chen J, Cui JL, Chen XM, Guo SX (2011) Antimicrobial activity and biodiversity of endophytic fungi in Dendrobium devonianum and Dendrobium thyrsiflorum from Vietman. Current Microbiology 62:1218–1224. https://doi.org/10.1007/s00284-010-9848-2
- Xiong YR, Manawasinghe IS, Maharachchikumbura SSN, Lu L, Dong ZY, Xiang MM, Xu B (2022) Pestalotioid species associated with palm species from Sothern China. Current Research in Environmental & Applied Mycology 12:285–321. https://doi.org/10.5943/cream/12/1/18
- Xu M, Fang Y, Cui X, Zeng YC, Zhang Y, Su S (2024) First report of leaf spot on Daphniphyllum macropodum caused by Neopestalotiopsis clavispora in China. Plant Disease 108:2235. https://doi.org/10.1094/PDIS-05-23-0893-PDN
- Xu M, Ye R, Cui X, Zeng Y, Zhang Y (2023a) First report of Neopestalotiopsis clavispora causing leaf spot on Photinia bodinieri in China. Plant Disease 107:3310. https://doi.org/10.1094/PDIS-06-23-1108-PDN
- Xu M, Ying Q, Zhang Y (2023b) First report of Neopestalotiopsis clavispora causing leaf spot on Phoebe bournei in China. Plant Disease 107:3633. https://doi.org/10.1094/PDIS-05-23-0884-PDN
- Xu MF, Jia OY, Wang SJ, Zhu Q (2016) A new bioactive diterpenoid from Pestalotiopsis adusta, an endophytic fungus from Clerodendrum canescens. Natural Product Research 30:2642–2647. https://doi.org/10.1080/14786419.2016.1138297
- Xu XL, Liu SY, Lv YC, Zeng Q, Liu YG, Yang CL (2022) Leaf blight on Photinia× fraseri caused by Pestalotiopsis trachicarpicola in China. Plant Disease 106:1520. https://doi.org/10.1094/PDIS-06-21-1351-PDN
- Xu XL, Liu SY, Xiao QG, Lv YC, Liu YG, Yang CL (2023c) First report of Pestalotiopsis neolitseae causing leaf blight on Photinia × fraseri in China. Journal of Plant Pathology 105:1689–1690. https://doi.org/10.1007/s42161-023-01441-x
- Yan X, Meng T, Qi Y, Fei N, Liu C, Dai H (2019) First report of Pestalotiopsis adusta causing leaf spot on raspberry in China. Plant Disease 103:2688. https://doi.org/10.1094/PDIS-07-18-1135-PDN
- Yang LN, Miao XY, Bai QR, Wang MQ, Gu ML, Zhao TC (2017) Neopestalotiopsis clavispora causing leaf spot on Phedimus aizoon var. latifolius, a new disease in China. Plant Disease 101:1952. https://doi.org/10.1094/PDIS-03-17-0422-PDN
- Yang Q, He YK, Yuan J, Wang Y (2022) Two new Pseudopestalotiopsis species isolated from Celtis sinensis and Indocalamus tessellatus plants in southern China. Phytotaxa 543:274–282. https://doi.org/10.11646/phytotaxa.543.5.2
- Yang Q, Zeng XY, Yuan J, Zhang Q, He YK, Wang Y (2021) Two new species of Neopestalotiopsis from southern China. Biodiversity data journal 9:e70446. https://doi.org/10.3897/BDJ.9.e70446
- Yang QQ, Li F, Guo QQ, Liang WX (2018) First report that Pestalotiopsis clavispora causes leaf spots on Photinia serratifolia in China. Plant Disease 102:1457. https://doi.org/10.1094/PDIS-12-17-1941-PDN
- Yin C, Zhang Z, Wang S, Ma L, Zhang X (2024) Three new species of Pestalotiopsis (Amphisphaeriales, Sporocadaceae) were identified by morphology and multigene phylogeny from Hainan and Yunnan, China. Mycokeys 107:51–74. https://doi.org/10.3897/mycokeys.107.122026
- Yu X, Müller WEG, Meier D, Kalscheuer R, Guo Z, Zou K, Umeokoli BO, Liu Z, Proksch P (2020) Polyketide derivatives from mangrove derived endophytic fungus Pseudopestalotiopsis theae. Marine Drugs 18:129. https://doi.org/10.3390/md18020129
- Yun YH, Ahn GR, Kim SH (2015) First report and characterization of Pestalotiopsis ellipsospora causing canker on Acanthopanax divaricatus. Mycobiology 43:366–370. https://doi.org/10.5941/MYCO.2015.43.3.366
- Zakaria K, Teet SE, Hamzah NH, Aznan AS, Manaf MTA, Ibrahim WNW, Leong LK, Iberahim NA, Musa N, Abdulrazzak L, Daud HM, Taib M, Hatai K, Akbar JB, Jalal KCA, Sheikh HI, Musa N (2021) Isolation and identification of fungi associated with diseased freshwater fishes in Terengganu, Malaysia. Songklanakarin Journal of Science & Technology 43:1131–1139. https://doi.org/10.14456/sjst-psu.2021.148
- Zakaria L, Aziz WNW (2018) Molecular identification of endophytic fungi from banana leaves (Musa spp.). Tropical Life Sciences Research 29:201–211. https://doi.org/10.21315/tlsr2018.29.2.14
- Zhang GJ, Cheng XR, Ding HX, Liang S, Li Z (2021a) First report of Pestalotiopsis lushanensis causing brown leaf spot on Sarcandra glabra in China. Plant Disease 105:1219. https://doi.org/10.1094/PDIS-08-20-1855-PDN
- Zhang GQ, Wijayawardene NN, Han LH, Kumla J, Suwannarach N, Li Q, Elgorban AM, Moussa IM, Coleine C, Dai DQ (2024a) Three novel woody litter inhabiting fungi in Didymosphaeriaceae, Phaeoseptaceae and Synnemasporellaceae from Zhujiangyuan Nature Reserve, Yunnan Province, P.R. China. Mycokeys 106:173–200. https://doi.org/10.3897/mycokeys.106.123105
- Zhang H, Mao YT, Ma MX, Tao GC, Wei TP, Jiang YL (2024b) Culturable endophytic Sordariomycetes from Rosa roxburghii: new species and lifestyles. Journal of Systematics and Evolution 62:637–676. https://doi.org/10.1111/jse.13035
- Zhang H, Wei TP, Li LZ, Luo MY, Jia WY, Zeng Y, Jiang YL, Tao GC (2021b) Multigene phylogeny, diversity and antimicrobial potential of endophytic Sordariomycetes from Rosa roxburghii. Frontiers in Microbiology 12:755919. https://doi.org/10.3389/fmicb.2021.755919
- Zhang J, Kapli P, Pavlidis P, Stamatakis A (2013a) A general species delimitation method with applications to phylogenetic placements. Bioinformatics 29:2869–2876. https://doi.org/10.1093/bioinformatics/btt499
- Zhang SN, Hyde KD, Jones EBG, Yu XD, Cheewangkoon R, Liu JK (2024c) Current insights into palm fungi with emphasis on taxonomy and phylogeny. Fungal Diversity 127: 55–301. https://doi.org/10.1007/s13225-024-00536-9
- Zhang W, Li Y, Lin L, Jia A, Fan X (2024d) Updating the species diversity of pestalotioid fungi: four new species of Neopestalotiopsis and Pestalotiopsis. Journal of Fungi 10:475. https://doi.org/10.3390/jof10070475
- Zhang Y, Maharachchikumbura SSN, Mckenzie EHC, Hyde KD (2012a) A novel species of Pestalotiopsis causing leaf spots of Trachycarpus fortunei. Cryptogamie, Mycologie 33:311–318. https://doi.org/10.7872/crym.v33.iss3.2012.311
- Zhang YM, Maharachchikumbura SSN, Tian Q, Hyde KD (2013b) Pestalotiopsis species on ornamental plants in Yunnan Province, China. Sydowia 65:113–128.
- Zhang YM, Maharachchikumbura SSN, Wei JG, McKenzie EHC, Hyde KD (2012b) Pestalotiopsis camelliae, a new species associated with grey blight of Camellia japonica in China. Sydowia 64:335–344.
- Zhang Z, Liu R, Liu S, Mu T, Zhang X, Xia J (2022) Morphological and phylogenetic analyses reveal two new species of Sporocadaceae from Hainan, China. MycoKeys 88:171–192. https://doi.org/10.3897/mycokeys.88.82229
- Zhang Z, Zhang J, Li D, Xia J, Zhang X (2023) Morphological and phylogenetic analyses reveal three new species of Pestalotiopsis (Sporocadaceae, Amphisphaeriales) from Hainan, China. Microorganisms 11:1627. https://doi.org/10.3390/microorganisms11071627
- Zhao H, Shu Y, Doilom M, Zeng X, Zhang H, Dong W (2024) Pestalotiopsis phyllostachydis sp. nov. from Guangdong, China. Phytotaxa 633:68−85. https://doi.org/10.11646/phytotaxa.633.1.8
- Zhao J, Zhang DP, Liu T, Liu WC, Pan LQ, Liao NY, Liu DW (2020) First report of Pseudopestalotiopsis and Neopestalotiopsis species causing leaf spot of Camellia chrysantha in China. Plant Disease 104:3071. https://doi.org/10.1094/PDIS-03-20-0598-PDN
- Zheng J, Ge Q, Yan Y, Zhang X, Huang L, Yin Y (2023a) dbCAN3: automated carbohydrate-active enzyme and substrate annotation. Nucleic Acids Research 51:W115–W121. https://doi.org/10.1093/nar/gkad328
- Zheng X, Liu X, Li X, Quan C, Li P, Chang X, Gu J, Khaskheli MI, Gong G (2023b) Pestalotiopsis species associated with blueberry leaf spots and stem cankers in Sichuan Province of China. Plant Disease 107:149–156. https://doi.org/10.1094/PDIS-07-21-1550-RE
- Zheng XR, Zhang MJ, Chen FM (2022) Occurrence of Pestalotiopsis lushanensis causing leaf blight on Buddhist pine in China. European Journal of Plant Pathology 162:655–665. https://doi.org/10.1007/s10658-021-02429-y
- Zhonghou L, Jingna L, Xiang L, Ping L, Hua Y (2023) Isolation and identification of pathogens causing dragon stripe disease on Kadsura coccinea. Journal of King Saud University Science 35:102516. https://doi.org/10.1016/j.jksus.2022.102516
- Zhou YK, Li FP, Hou CL (2018) Pestalotiopsis lijiangensis sp. nov., a new endophytic fungus from Yunnan, China. Mycotaxon 133:513–522. https://doi.org/10.5248/133.513
- Zhu H, Niu XQ, Yu FY, Liu L, Yan W, Qin WQ (2015) First report of leaf blight of Dictyosperma album caused by Pestalotiopsis adusta in China. Plant Disease 99:1040. https://doi.org/10.1094/PDIS-10-14-1093-PDN
- Zhu RJ, Liu YL, Li HY, Liu JB, Han XY (2023a) First report of Neopestalotiopsis clavispora causing leaf spots on Garcinia mangostana in China. Plant Disease 107:1946. https://doi.org/10.1094/PDIS-05-22-1120-PDN
- Zhu W, Hu S, Zhou Z, Tang X, Wu X, Li Z, Ding H (2023b) First report of leaf spot on Rhododendron delavayi caused by Pestalotiopsis scoparia in China. Plant Disease 107:1626. https://doi.org/10.1094/PDIS-08-22-1896-PDN
- Copyright: The Author(s) 2026. Published by BioAcademic Press on behalf of Kunming Institute of Botany, Chinese Academy of Sciences (CAS) and Mushroom Research Foundation. This is an open access article under the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0), which permits use, distribution and reproduction in any medium, provided the original work is properly cited.